Abstract
To respond to the external environmental changes for survival, bacteria regulates expression of a number of genes including transcription factors (TFs). To characterize complex biological phenomena, a biological system-level approach is necessary. Here we utilized six computational biology methods to infer regulatory network and to characterize underlying biologically mechanisms relevant to radiation-resistance. In particular, we inferred gene regulatory network (GRN) and operons of radiation-resistance bacterium Spirosoma montaniterrae DY10\(^T\) and identified the major regulators for radiation-resistance. Our results showed that DNA repair and reactive oxygen species (ROS) scavenging mechanisms are key processes and Crp/Fnr family transcriptional regulator works as a master regulatory TF in early response to radiation.
Similar content being viewed by others
Introduction
Exposure to high levels radiation adversely affects cells, damaging the nucleic acids, proteins, lipids and cellular components of cells, which results in cell death. Ultraviolet (UV) radiation directly damages DNA and causes genetic disruption by inducing pyrimidine dimerization, single-stranded breaks (SSBs) and double-stranded breaks (DSBs)1,2,3. In addition, radiation induces indirect damage through reactive oxygen species (ROS), such as hydroxyl radicals, superoxide anions, and hydrogen peroxide within cells by radiolysis of water. Interestingly, there are radiation-resistant organisms which are able to withstand high levels of radiation exposure. The radiation-resistant organisms are known to protect cells from genome-wide disruption and homeostasis disruption by activating underlying defense systems such as DNA repair and ROS detoxification.
Deinococcus radiodurans, is the most well characterized radiation-resistant organism. Deinococcus radiodurans shows remarkable radiation-resistance characteristics for high levels of ionizing radiation (IR), X-rays and gamma rays4, compared to the yeast Saccharomyces cerevisiae, the bacterium Escherichia coli, and the human cells. When Deinococcus radiodurans radiated, cell recovers the radiation induced damage through efficient regulation of DNA repair systems, including extended synthesis-dependent strand annealing (ESDSA) process and homologous recombination by the RecF pathway5. This organism deals with radiation induced ROS through enzymatic systems and non-enzymatic systems, including the following molecules: superoxide dismutase, catalase, peroxidase, pyrroloquinoline-quinone, deinoxanthin, and bacillithiol6. In particular, the unusual high intracellular Mn/Fe ratio, has been correlated with radiation-resistance through the low-molecular-weight metabolite (LMWM) systems that forms ROS-scavenging Mn\(^{2+}\)-metabolite complexes7. In addition, it is reported that organisms in three domains of life (Archaea, Bacteria, and Eukaryota) such as Rubrobacter radiotolerans, Kineococcus radiotolerans, Halobacterium salinarum, Thermococcus gammatolerans and Pyrococcus furiosus have radiation-resistance through different mechanisms8,9,10,11.
Organism-level biological properties are driven by interactions among genes rather than the effect of individual genes. Genes rarely work individually and generally cooperate to perform specific biochemical functions. Also worth noting is transcription of genes that is mainly regulated by transcription factors (TFs) in a cell-specific manner12. Thus, we view that radiation-resistance property is the result of association of multiple intracellular elements including TFs and target genes. Existing studies on radiation-resistance in various strains, such as Deinococcus radiodurans, also identified that radiation-resistance is driven through a complex pathways. However, in order to understand the complex underlying mechanism of radiation-resistance both at the transcriptome-level and at the system-level, powerful network-based approaches are needed.
Innovative advances in bioinformatics have accelerated biological knowledge discovery and allowed mathematical modeling of biological systems with transcriptional repressors/activators and post-transcriptional RNA regulators. Ahn et al., applied Gene Regulatory Network (GRN) and successfully identified regulatory modules that respond to cold and heat stress in Arabipodsis13. GRN is a collection of genetic molecules that represent systematic understanding of the molecular interactions underlying biological processes includes how gene expressions are regulated and carry out cellular functions14,15,16,17. GRN based network analysis aids in the identification of regulatory mechanisms to understand complex biological phenomena12. For decades, a number of computational methods have attempted to develop GRN inference algorithms18,19,20,21. One of these methods relies on TF binding motifs or genomic accessibility22.
In our previous work, we isolated a remarkable radiation-resistant bacterium Spirosoma montaniterrae DY10\(^{T}\) from the soil of south Korea and identified their taxon and radiation-resistance property23. In addition, several radiation-resistant species have been reported in the Spirosoma genus24,25. Besides this, the specific mechanism of radiation-resistance of Spirosoma genus has not been investigated. Understanding the complex underlying mechanisms of Spirosoma genus involves two major issues. First, for understanding complex underlying mechanism, gene-level approach could not bridge the gap between transcriptome-level perturbations and organism-level phenomenon. Second, to apply a system-level approach, knowledge between genes and regulators is limited, and is only available for a few well-known model strains. In this study, we generated time-series transcriptome data of radiation-resistant strain DY10\(^T\) under UVC radiation. Subsequently, we show that gene regulatory network for uncharacterized strain can be inferred using six in silico methods without labor-intensive and time-consuming in vitro and in vivo experiments (Fig. 1). Dataflow of utilized computational tools were summarized in Supplementary Table S1. In summary, our biological system-level approach revealed that the radiation-resistance phenotype of strain DY10\(^{T}\) stems from DNA repair and ROS scavenging mechanisms, and we identified Crp/Fnr family transcriptional regulator as a master regulatory TF in early response to UVC radiation.
Schematic overview of our approach. In order to investigate radiation-resistance mechanism, RNA from UVC treated and untreated samples was extracted and quantified. In order to understand radiation-resistance mechanism in system-level, regulatory network based analysis and operon analysis were conducted. In the regulatory network based analysis, GRN of Spirosoma montaniterrae DY10\(^{T}\) was constructed using four computational methods (DeepTFactor, BLAST, HOMER, and Bedtools) and Key regulatory relationships over time on the GRN identified using PropaNet. In the operon analysis, operons in Spirosoma montaniterrae DY10\(^{T}\) were detected using operonSEQer.
Results
Biological implications of Spirosoma montaniterrae DY10\(^T\) in response to UVC radiation
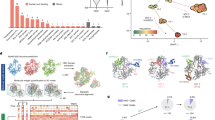
To investigate transcriptome-level responses to UVC radiation, we analyzed differentially expressed genes (DEGs) between the UVC untreated and UVC treated samples over time (|\(\log _{2} \text {(fold change)}\)| \(\ge\) 1; Fig. 2A,B).Number of DEGs at 5 min, 1 h, 3 h, and 12 h after UVC radiation for Spirosoma montaniterrae was 97, 65, 34, and 54 for up-regulated and 83, 61, 14, and 11 for down-regulated, for a total of 264 unique genes. In the case of E. coli, the number of DEGs at 5 min, 10 min, 20 min, 40 min and 60 min after UVC radiation was 355, 241, 247, 238 and 137 for up-regulated and 261, 220, 284, 152 and 77 for down-regulated, respectively for a total of 1039 unique genes. The gene expression patterns of strain DY10\(^T\) and E. coli were visualized through a hierarchically-clustered heatmap (Fig. 2C,D). We identified two clusters for strain DY10\(^T\) and three clusters for E. coli with hierarchically-clustered heatmaps by Scipy package26 (Linkage threshold was set to 3). Cluster1 in strain DY10\(^T\) contained 105 genes and was the set of genes that were up-regulated in early response, and cluster2 in strain DY10\(^T\) contained 162 genes and was the set of genes that were down-regulated in early response (Fig. 2C). Cluster1 in E. coli contained 410 genes and was the set of genes that were up-regulated in early response, and cluster2 in E. coli contained 309 genes and was the set of genes that were down-regulated in early response (Fig. 2D). Cluster1 and cluster2 of each strain showed similar expression patterns. However, gene comparison based on sequence homology, 18 genes were shared in cluster1 and 33 genes were shared in cluster2. From these results, we confirmed that radiation-resistant strain and radiation-sensitive strain respond differently to UV radiation. Furthermore, we performed pathway enrichment analysis with KEGG pathway and DEGs to capture the biological implications of DEGs in strain DY10\(^T\) and E. coli. The results showed that Homologous recombination, Mismatch repair (MMR) and Nitrogen metabolism were significantly enriched terms (p<0.05) among up-regulated genes in early response to UVC radiation in strain DY10\(^{T}\) (Fig. 3A). In addition, we investigated the perturbations over time of the genes belonging to significantly enriched terms.
In Homologous recombination and Mismatch repair pathways, ssb, recA, recR, ruvB and mutS were remarkably up-regulated genes at an early response after UVC radiation and gradually decreased over time (Fig. 3C). ssb (AWR27_RS23025) is a gene that involved in both Homologous recombination and Mismatch repair process. Expression of ssb is a part of the SOS regulon and elevation of ssb gene in UVC radiated cells in an SOS-dependent manner was reported27. recA (AWR27_RS17265) and recR (AWR27_RS05955) play a central role in DNA maintenance and its induction is considered as a dominant marker for the onset of homologous recombination. When DNA damage occurs, bacterial RecR forms complexes with RecO and RecF that enhances loading RecA on to damaged DNA and catalyzes homologous recombination process28,29,30. We found it interesting that the strain DY10\(^T\) also has RecR, RecF and RecO encoded genes but only recR was perturbed significantly. ruvB (AWR27_RS21175) is a gene encodes Holliday junction DNA helicase RuvB which is also key molecule for DNA maintenance especially in homologous recombination31,32. mutS (AWR27_RS05750) encodes a bacterial MMR protein that responsible for the repair of mispaired bases33. In Nitrogen metabolism, nitrite reductase large subunit (AWR27_RS22355) and glutamine synthetase beta-grasp domain-containing protein (AWR27_RS10835) are products of remarkably up-regulated genes at an early response after UVC radiation (Fig. 3D). Nitrite reductase (NiR) is an enzyme that catalyzes the reduction of nitrite (NO2\(^{-}\)) to nitric oxide (NO), thereby carrying out ionizing radiation-induced nitric oxide detoxification34,35. Glutamine synthetase (GS) is an ATP-dependent enzyme that catalyzes the assimilation of glutamate and ammonia to glutamine and the up-regulation of GS in bacteria is an effective means to survive when assaulted by ROS36.
Unlike strain DY10\(^T\), among the up-regulated DEGs of E. coli in response to UVC radiation, the significantly enriched terms (Ribosome, Fatty acid biosynthesis, Polyketide sugar unit biosynthesis, Pyrimidine metabolism, and Biotin metabolism) were not related with DNA repair or ROS detoxification processes (Fig. 3B). This difference explains partially why E. coli is less resistant to radiation stress for survival.
Differentially expressed genes (DEGs) analysis of Spirosoma montaniterrae DY10\(^{T}\) and Escherichia coli. (A,B) The number of up-regulated and down-regulated DEGs with |\(\log _{2} \text {(fold change)}\)| \(\ge\) 1 in varying time points. (C,D) Heatmap of 264 DEGs of Spirosoma montaniterrae and 1039 DEGs of E. coli in varying time points. The row represents the gene, and the columns represents each sample with varying time points. The redder the color, the higher the gene expression value. The bluer the color, the lower the gene expression value. Each bounding box identifies a gene cluster.
Pathway enrichment analysis of Spirosoma montaniterrae DY10\(^{T}\) and Escherichia coli. (A) The significantly enriched terms of KEGG pathways of Spirosoma montaniterrae in varying time points. (B) The significantly enriched terms of KEGG pathways of Escherichia coli in varying time points. (C,D) Variation over time of significantly changed genes among genes related to homologous recombination, mismatch repair and Nitrogen metabolism processes in Spirosoma montaniterrae.
Regulatory mechanism in response to UVC radiation in Spirosoma montaniterrae DY10\(^{T}\)
In order to understand the regulatory mechanism of strain DY10\(^{T}\) in response to UVC radiation, we investigated time-varying transcriptional network module using PropaNet (“Materials and methods” section: “Gene regulatory network based analysis”) with GRN and time-series transcriptome data; untreated sample and UVC radiated samples collected at 5 min, 1 h, 3 h and 12 h after radiation. Transcriptional network module is a regulatory network of TFs and target genes that can describe state of organism at each time point. The results showed the visualization of identified time-varying transcriptional network modules in response to UVC radiation (Fig. 4A). The nodes and edges shown in the figure represent the condition-specific sub-network modules containing TFs and target genes that respond to UVC stimulation among the template GRN. Changes of the sub-network modules showed that the effects of UVC radiation gradually recovered over time. In order to further understand the biological implications of time-varying transcriptional network module, we performed pathway enrichment analysis with genes in each module and KEGG pathways. Homologous recombination, and Nitrogen metabolism was identified as significantly enriched and up-regulated terms at an early response (Fig. 4B). Homologous recombination involved genes in the early response module (5 min) were recA (AWR27_RS17265), ruvB (AWR27_RS21175), and ssb (AWR27_RS23025). Nitrogen metabolism involved genes in the early response module (5 min) were nitrite reductase large subunit (AWR27_RS22355), and glutamine synthetase beta-grasp domain containing protein (AWR27_RS10835). Additionally, Two-component system and Quorum sensing pathway were identified as significantly enriched and down-regulated terms at the early response module. Biological implications for all time points were shown in Supplementary Fig. S2. When analyzing the regulator of each network module, 49 TFs were identified as major TFs in early response. Over time, different target genes in the gene regulatory network were regulated by different sets of TFs, leading the cell to different states (Fig. 4C).
The key to regulatory network based analysis is regulatory mechanism with identified major transcription factors and their target genes by time. In the DEGs analysis, we identified 180 genes that were significantly affected by UVC radiation. Through the regulatory network-based analysis, we prioritized 120 target genes in early response to UVC stimulation, and additionally identified regulatory relationships with 49 regulatory factors. In particular, significantly up-regulated pathways (Homologous recombination and Nitrogen metabolism) in the 5-min module were analyzed in detail and visualized (Fig. 5). The result showed the Crp/Fnr family transcriptional regulator (AWR27_RS09520) is a master regulatory TF that initiates expression changes of recA, ruvB, ssb, nitrite reductase large subunit, and glutamine synthetase beta-grasp domain containing protein directly or indirectly. Crp/Fnr family transcription regulators typically function as transcriptional activators in several bacterial species and regulate fumarate reductase, nitrate, and nitrite reductase to respond to intracellular and extracellular signals such as temperature, oxidative and nitrative stress, and nitric oxide37.
As a result, both the DEGs based analysis and the regulatory network based analysis identified that Homologous recombination and Nitrogen metabolism pathways were the most significant processes for radiation-resistance in strain DY10\(^T\). However, the DEG based analysis showed only quantitatively meaningful genes. Meanwhile, the regulatory network based approach identified the master regulatory TF of radiation-resistance mechanism. In addition, our approach identified the other 17 radiation-resistance mechanism related genes and their regulatory interactions that could not be captured by the gene-level quantitative approach.
Transcriptional network module analysis. (A) Perturbation of transcriptional network module in varying time points. Major transcription factors are shown in red color and target genes are shown in green color. (B) Pathway enrichment analysis of genes in early response (5 min) module with KEGG pathways. Red colored bar represents up-regulated terms, blue colored bar represents down-regulated terms, and statistically insignificant terms are colored in light blue. (C) Venndiagrams show major transcription factors and major target genes over time identified by our analysis.
Homologous recombination and Nitrogen metabolism-related transcriptional network module in early response of Spirosoma montaniterrae DY10\(^T\) to UVC radiation. Network represents regulatory relation between genes. In the network, diamond shaped nodes are transcription factors and circled nodes are target genes. Grey colored nodes are Homologous recombination-related genes and orange colored genes are Nitrogen metabolism-related genes.
Operons involved in early response to UVC radiation
Operon is an important genetic regulatory system found in bacteria. The genes in operon are co-transcribed under the control of a single promoter and are often functionally related38. By analyzing the operon, we can understand how the expression of genes in organism is efficiently regulated in response to external stimuli. It is an important part of understanding the complex regulatory system of bacteria. operonSEQer39 is a statistics and machine learning based algorithm that predicts relevant operon pair with signals from RNA-seq data across two genes. Unlike other existing tools, operonSEQer is a flexible tool that does not use functional relationship information and showed remarkable operon predictive performance to bacterial strains that have not been studied.
In this study, we identified 1113 operons in Spirosoma montaniterrae and classified them into six clusters with similar operon value perturbation patterns (Fig. 6A). To understand the function of operons involved in initial response to UVC radiation, we conducted pathway enrichment analysis with genes in operon cluster1 and operon cluster2 that initially up-regulated and gradually decline over time. The results showed that the most significant term of genes in operon cluster1 is a Base excision repair (Fig. 6B) and the most significant term of genes in operon cluster2 is an Oxidative phosphorylation and nitrogen metabolism was also included in significant terms (Fig. 6C). Base excision repair is also involved in DNA repair process induced by UVC radiation like Homologous recombination and Oxidative phosphorylation is also biological process that involved in ROS scavenging40,41,42. Among the operons involved in the initial reaction, three operons contain Base excision repair-related genes and eight operons contain Oxidative phosphorylation-related genes (Fig. S1). We combined GRN based regulatory information with predicted operons, two of the eight Oxidative phosphorylation-related operons have the binding site of the master regulatory TF identified in the previous analysis in front of the operon. Thereby those two operons have potential to be an another target regulatory gene set of master regulatory TF in early response. As a result, some of DNA repair and ROS scavenging processes-related genes in strain DY10\(^T\) are located close to each other on the genome and are regulated efficiently by forming operons.
Operon prediction and analysis. (A) Six operon clusters were identified with k-means clustering method. Black colored line is the average of the operon values that belonging to the cluster at the each time points. (B,C) Pathway enrichment analysis of genes in operon cluster 1 and 2. Red colored bar is most enriched term in cluster1 and orange colored bar is most enriched term in cluster2. The red dotted line indicates statistical significance of p-value.
Discussion
In radiation-resistance species, such as Deinococcus sp., DNA repair and ROS scavenging processes have been experimentally demonstrated as major mechanisms for survival under UVC radiation. However, due to the genetic diversity between species or genera43 of bacteria, biological mechanisms of DNA repair and ROS scavenging processes can be significantly differ in diverse bacteria species. Thus, straightforward DEG analysis may not be sufficient for identifying DNA repair and ROS scavenging processes in uncharacterized genomes such as strain DY10\(^T\) in our study. Furthermore, little is known about the regulatory mechanisms for survival under UVC radiation. Therefore, we sought to understand the radiation-resistance characteristic in terms of biological regulatory system by constructing homology based GRN and identifying operons with six computational biology techniques.
Gene expression level comparison identified that 180 genes (97 up-regulated and 83 down-regulated) are involved in the early response of strain DY10\(^T\) to UVC radiation and that the gene set changes over time. With gene-level analysis, we identified condition-specific expression perturbed gene sets. However, gene-level approach cannot capture global patterns of gene expression that are important for understanding complex biological processes. To complement this limitation, pathway-level analysis was conducted. Pathway-level approaches capture a more comprehensive view of biological processes by integrating information from multiple genetic and molecular pathways. We observed that Homologous recombination, Mismatch repair and Nitrogen metabolism were significantly altered processes in the early response of strain DY10\(^T\) to UVC radiation. Homologous recombination is an important DNA repair process that repairs damaged DNA sequences using identical or very similar undamaged DNA sequences as templates. Recombinational DNA repair is both the most complex and the least understood of bacterial DNA repair processes44. MMR also contributes to genome stability by mediating DNA damage signaling in response to DNA damage45. MMR proteins have a role in the imposition of cell cycle check-points and apoptotic signaling in response to UVC radiation. One mechanism of the MMR-induced DNA damage response is that MMR proteins recognize sites of damage and recruit components of the DNA damage response pathway either directly or via interactions with other entities46. Nitrogen metabolism is a basic biological process of nitrogen cycle and plays a crucial role in removing ROS in the cell. When cells are exposed to UVC radiation, nitric oxide and ROS are formed and react with hydroxyl radical-induced adenine radicals. This reaction forms common DNA damage bases (hypoxanthine and 8-azaadenine) and induces DNA double-strand breaks47. Our results showed that strain DY10\(^T\) repair damaged DNA and remove ROS together to increase viability in early response to UVC radiation.
Gene-level and pathway-level analysis are highly annotation-biased approach, whereas network-based approaches focus on associations between genes rather than relying on annotations. By constructing time-varying transcriptional network modules, we identified which genes were regulated and how those genes were regulated during UVC damage recovery. In particular, we identified DNA repair and ROS scavenging processes are key mechanisms for UVC induced damage recovery in strain DY10\(^{T}\) as in other radiation-resistant bacterium. Furthermore, we first revealed that Crp/Fnr family transcription regulator (AWR27_RS09520) is a master regulator TF that regulates the initial activation of UVC induced damage recovery processes of strain DY10\(^T\). The function of Crp/Fnr family transcription regulator as a regulator of nitrite reductase with ROS scavenging function has also been found in other species37. However, its role as a regulator of the DNA repair system has not been well studied. In addition, two genes (AWR27_RS09820 and AWR27_RS11185) in the regulatory module did not show significant expression changes in response to UVC radiation and their functions are unknown. However, they were captured as intermediate genes in regulatory module and it is possible that it has functions involved in DNA repair and ROS scavenging processes through regulatory network based analysis. Operon is a major regulatory system in bacteria48, and operon analysis also showed the genetic response to UVC in terms of systems. Our results showed that genes involved in DNA repair and ROS scavenging processes are located close to each other on the genome and are efficiently regulated. Similar to network-based approach, operon analysis provides intuition about the function of genes belonging to an operon but whose function is unknown.
Conclusion
In this study, we characterized the underlying mechanisms for radiation-resistance by constructing gene regulatory network of an uncharacterized bacterium strain DY10\(^T\). In particular, we identified major regulatory TFs for radiation-resistance in a computational framework. It is interesting that a large number of genes regulated by the TFs identified are not well characterized, thus we believe that identifying functions of these genes can be valuable new knowledge for the mechanisms for UVC resistance. Consequently, our biological system-level approach can help bridge the gap between the transcriptome-level and the organism-level and enable interpretsation of complex biological processes under diverse conditions.
On the other hand, there are two limitations in our approach. First, PropaNet depends on reliable TF-target interactions. GRN of the strain DY10\(^T\), is inferred based on homology with genomic sequence of bacteria in UniProt database and it has potential to contain false positive TF-target interactions. To overcome this limitation, validation through additional complicate experiments is required. The most widely used method for identifying TF-target interaction is TF ChIP-seq. With TF ChIP-seq, binding sites and DNA sequence motif for TFs can be identified precisely. Second, our framework construct GRN that do not consider operon system in bacteria. Operon is a very important gene regulatory system in bacterial gene network. In previous studies, it was found that more than 50% of all genes in E. coli are organized into operons49. In this study, 66% of all genes in strain DY10\(^T\) were predicted to be regulated by the operon system. To capture more meaningful underlying mechanisms in bacteria, developing operon system considered bacterial GRN inference method can be future work.
Materials and methods
Cell growth, radiation and RNA extraction
The bacterial strain DY10\(^{T}\) (KCTC 23999\(^{T}\)) was grown at \(25\,^{\circ }\textrm{C}\) in liquid nutrient-rich medium tryptone glucose yeast extract (TGY; 1% tryptone, 0.1% glucose and 0.5% yeast extract) or TGY agar plates23. For the UVC treatment, the early stationary phase cells on TGY agar plates were exposed to 600 JM\(^{-2}\) of ultraviolet using an ultraviolet crosslinker (UVP, CX-2000, CA, USA) at 254 nm50,51. The post-radiation cells of UVC were collected by centrifugation (17,500×g for 5 min) at 5 min, 1 h, 3 h and 12 h with washing and stored with RNAlater solution at \(-80\,^{\circ }\textrm{C}\). Total RNA was isolated using the Ambion Ribopure-Yeast RNA kit, according to manufacturer’s instructions.
Library preparation and gene expression analysis
Strand-specific RNA-seq library preparation and sequencing was carried out as a paid service by Somagenics Inc., Santa Cruz, California, USA. Paired-end reads (Illumina NextSeq 500 v2, 2 \(\times\) 150 bp, 3 Gb average output per sample) were obtained from strain DY10\(^{T}\). Each sample was obtained from three independent RNA isolations, for each strain. Reads of each samples’ adaptor trimmed by Trim galore (version 0.6.7)52 for quality control. Clean reads of the samples were aligned against the strain DY10\(^{T}\) reference genome (Build: ASM198895v1) using Hisat2 (version 2.1.0)53. HTSeq (version 0.12.4)54 was used to quantify the mapped reads per ORF. Differentially expressed genes were identified with a \(\log _{2} \text {(fold change)}\) threshold of − 1 and 1 (|\(\log _{2} \text {(fold change)}\)| \(\ge\) 1). For radiation-sensitive (negative) strain, UVC radiated E. coli with multiple time points gene expression data (GSE9) downloaded from GEO (Gene Expression Omnibus).
Homology based reconstruction of Gene Regulatory Network (GRN)
To determine the regulatory interactions between genes in strain DY10\(^T\) with genome and protein sequence information, We used four computational methods. GRN construction operates in five steps as below. In the entire process, homology to known prokaryotic sequences is an important concept to reflect the regulatory mechanisms present in prokaryotic organisms and has been used for TF screening and TF binding site prediction55.
-
Step 1. Transcription factor prediction. DeepTFactor56 is a state-of-the-art deep learning based tool that learns DNA-binding domains of TFs in several bacteria species and predicts whether a query protein is a TF. In this study, pre-trained DeepTFactor56 was used for identifying potential TFs using protein sequences from strain DY10\(^{T}\). Protein sequences from strain DY10\(^{T}\) were used as input queries of model and model returns whether a query protein is a TF.
-
Step 2. Homology based transcription factor screening. BLAST57 is a widely used local alignment tool. Using the program blastp with the default parameters, among the potential TFs predicted by DeepTFactor, potential TFs that are statistically similar (e-value < 0.05) to known prokaryotic TFs included in the prokaryotic TF database were selected. Sequences of known prokaryotic TFs were collected from UniProt58 database based on strains in PRODORIC59 database.
-
Step 3. Prediction of accessible genome regions for each transcription factors. HOMER60 is a motif discovery algorithm that searches accessible genome regions for TFs using the position weight matrix (PWM) of TFs. We computed the genome binding sites of screened TFs of strain DY10\(^T\) using HOMER. PWM of each TF of strain DY10\(^T\) is derived from PWM of the most similar TF among the TFs present in the PRODORIC database.
-
Step 4. Identification of target gene of transcription factors. Bedtools61 is a genomic analysis tool. We identified the target genes of the selected TFs by analyzing the computed accessible genomic regions and its down-stream genes with bedtools. Through the bedtools closest command of bedtools v2.30.0, the gene closest to the binding site of each TF was identified and assigned as the target gene of TF.
-
Step 5. Reconstruction of Gene Regulatory Network With identified TFs and their target genes, we reconstructed a GRN of strain DY10\(^{T}\). GRN is a bipartite graph where source nodes are TFs and target nodes are target genes of TFs.
Gene regulatory network based analysis
Ahn et al. developed a PropaNet13, a state-of-the-art time-series trancriptome analysis method. PropaNet calculates the impact of each TF on the target genes at each time point using influence maximization and network propagation62,63 techniques with template GRN. The calculated impact of each TF is used to select the major TFs for each time points by prioritizing them. A regulatory network was constructed for each time point using the selected major TFs and their target genes. The constructed time-varying regulatory networks allow interpretation of the regulation of key mechanisms for UVC radiation response. In this study, a constructed GRN of strain DY10\(^T\), time-series expression profile, DEGs list in each time point and TF list of strain DY10\(^T\) were used as inputs of PropaNet and time-varying networks were obtained and additional analysis were conducted.
Operon prediction and clustering
Krishnakumar et al. developed a operonSEQer39, a state-of-the-art statistic and machine learning based algorithm that predicts bacterial genome operon. operonSEQer shows remarkable predictive power using only RNA-seq data and genomic features. In this study, we identified operons in Spirosoma montaniterrae using operonSEQer with follow criteria. Among the six machine-learning models included in operonSEQer, 3 or more models predicted gene pairs as operon were selected as being in the same operons.
Predicted operons were classified into six clusters according to the pattern of operon value change over time. i-th operon (\(O_{i}\)) is represented as a set of operon values listed in time order and the operon value at time t (\(O_{i,t}\)) is the average of z-score values of n genes({\(g_{i,t,1}\), \(\cdot\) \(\cdot\) \(\cdot\), \(g_{i,t,n}\)}) at time t belonging to the i-th operon. For clustering, we used the scikit-learn module of python3 to implement the K-means clustering algorithm64. In this study, we divided a set of operons (\(O_{i}\)) into six disjoint clusters.
where \(O_{i}\) is a set of operon values listed in time order of i-th operon, \(G_{i,t}\) is a set of expression level of genes included in i-th operon at time t, \(g_{i,t,j}\) is a expression level (z-score) of gene included in \(G_{i,t}\).
Pathway database
For pathway enrichment analysis, we used biological pathway information obtained from the Kyoto Encyclopedia of Genes and Genomes (KEGG) database65. We used 117 pathways and 1051 genes for Spirosoma montaniterrae DY10\(^{T}\) and 121 pathways and 1705 genes for E. coli from the KEGG database.
Data availability
The datasets generated and analysed during the current study are deposited in the Gene Expression Omnibus (GEO) repository. Data accession link (GSE223604): https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE223604.
References
Kripke, M. L., Cox, P. A., Alas, L. G. & Yarosh, D. B. Pyrimidine dimers in DNA initiate systemic immunosuppression in UV-irradiated mice. Proc. Natl. Acad. Sci. 89, 7516–7520 (1992).
Buatti, J. M., Rivero, L. R. & Jorgensen, T. J. Radiation-induced DNA single-strand breaks in freshly isolated human leukocytes. Radiat. Res. 132, 200–206 (1992).
Vignard, J., Mirey, G. & Salles, B. Ionizing-radiation induced DNA double-strand breaks: A direct and indirect lighting up. Radiother. Oncol. 108, 362–369 (2013).
Makarova, K. S. et al. Genome of the extremely radiation-resistant bacterium Deinococcus radiodurans viewed from the perspective of comparative genomics. Microbiol. Mol. Biol. Rev. 65, 44–79 (2001).
Bentchikou, E., Servant, P., Coste, G. & Sommer, S. A major role of the RecFOR pathway in DNA double-strand-break repair through ESDSA in Deinococcus radiodurans. PLoS Genet. 6, e1000774 (2010).
Lim, S., Jung, J.-H., Blanchard, L. & de Groot, A. Conservation and diversity of radiation and oxidative stress resistance mechanisms in Deinococcus species. FEMS Microbiol. Rev. 43, 19–52 (2019).
Slade, D. & Radman, M. Oxidative stress resistance in Deinococcus radiodurans. Microbiol. Mol. Biol. Rev. 75, 133–191 (2011).
Ferreira, A. C. et al. Characterization and radiation resistance of new isolates of Rubrobacter radiotolerans and Rubrobacter xylanophilus. Extremophiles 3, 235–238 (1999).
McCready, S. & Marcello, L. Repair of UV damage in Halobacterium salinarum. Biochem. Soc. Trans. 31, 694–698 (2003).
Bagwell, C. E. et al. Survival in nuclear waste, extreme resistance, and potential applications gleaned from the genome sequence of Kineococcus radiotolerans SRS30216. PLoS ONE 3, e3878 (2008).
Strahl, H. & Greie, J.-C. The extremely halophilic archaeon Halobacterium salinarum R1 responds to potassium limitation by expression of the K+-transporting KdpFABC P-type ATPase and by a decrease in intracellular K+. Extremophiles 12, 741–752 (2008).
Das, D., Banerjee, N. & Zhang, M. Q. Interacting models of cooperative gene regulation. Proc. Natl. Acad. Sci. 101, 16234–16239 (2004).
Ahn, H. et al. Propanet: Time-varying condition-specific transcriptional network construction by network propagation. Front. Plant Sci. 10, 698 (2019).
Friedman, N. Inferring cellular networks using probabilistic graphical models. Science 303, 799–805 (2004).
Ihmels, J. et al. Revealing modular organization in the yeast transcriptional network. Nat. Genet. 31, 370–377 (2002).
Segal, E. et al. Module networks: Identifying regulatory modules and their condition-specific regulators from gene expression data. Nat. Genet. 34, 166–176 (2003).
Sachs, K., Perez, O., Pe’er, D., Lauffenburger, D. A. & Nolan, G. P. Causal protein-signaling networks derived from multiparameter single-cell data. Science 308, 523–529 (2005).
Huynh-Thu, V. A., Irrthum, A., Wehenkel, L. & Geurts, P. Inferring regulatory networks from expression data using tree-based methods. PLoS ONE 5, e12776 (2010).
Chan, T. E., Stumpf, M. P. & Babtie, A. C. Gene regulatory network inference from single-cell data using multivariate information measures. Cell Syst. 5, 251–267 (2017).
Papili Gao, N., Ud-Dean, S. M., Gandrillon, O. & Gunawan, R. Sincerities: Inferring gene regulatory networks from time-stamped single cell transcriptional expression profiles. Bioinformatics 34, 258–266 (2018).
Moerman, T. et al. GRNBoost2 and Arboreto: Efficient and scalable inference of gene regulatory networks. Bioinformatics 35, 2159–2161 (2019).
Kamimoto, K., et al. Celloracle: Dissecting cell identity via network inference and in silico gene perturbation. Nature 1–10 (2023).
Lee, J.-J. et al. Spirosoma montaniterrae sp. nov., an ultraviolet and gamma radiation-resistant bacterium isolated from mountain soil. J. Microbiol. 53, 429–434 (2015).
Lee, J.-J. et al. Spirosoma radiotolerans sp. nov., a gamma-radiation-resistant bacterium isolated from gamma ray-irradiated soil. Curr. Microbiol. 69, 286–291 (2014).
Lee, J. H. et al. Spirosoma taeanense sp. nov., a radiation resistant bacterium isolated from a coastal sand dune. Antonie van Leeuwenhoek 114, 151–159 (2021).
Virtanen, P. et al. SciPy 1.0: Fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272. https://doi.org/10.1038/s41592-019-0686-2 (2020).
Feliciello, I. et al. Regulation of SSB gene expression in Escherichia coli. Int. J. Mol. Sci. 23, 10917 (2022).
Umezu, K. & Kolodner, R. D. Protein interactions in genetic recombination in Escherichia coli. Interactions involving RecO and RecR overcome the inhibition of RecA by single-stranded DNA-binding protein. J. Biol. Chem. 269, 30005–30013 (1994).
Seitz, E. M., Brockman, J. P., Sandler, S. J., Clark, A. J. & Kowalczykowski, S. C. RadA protein is an archaeal RecA protein homolog that catalyzes DNA strand exchange. Genes Dev. 12, 1248–1253 (1998).
Morimatsu, K. & Kowalczykowski, S. C. RecFOR proteins load RecA protein onto gapped DNA to accelerate DNA strand exchange: A universal step of recombinational repair. Mol. Cell 11, 1337–1347 (2003).
West, S. C. & Connolly, B. Biological roles of the Escherichia coli RuvA, RuvB and RuvC proteins revealed. Mol. Microbiol. 6, 2755–2759 (1992).
Iwasa, T. et al. Synergistic effect of ATP for RuvA–RuvB–Holliday junction DNA complex formation. Sci. Rep. 5, 1–13 (2015).
Oliver, A., Baquero, F. & Blázquez, J. The mismatch repair system (mutS, mutL and uvrD genes) in Pseudomonas aeruginosa: Molecular characterization of naturally occurring mutants. Mol. Microbiol. 43, 1641–1650 (2002).
Vučetić, M., Cormerais, Y., Parks, S. K. & Pouysségur, J. The central role of amino acids in cancer redox homeostasis: Vulnerability points of the cancer redox code. Front. Oncol. 7, 319 (2017).
Bowman, L. A., McLean, S., Poole, R. K. & Fukuto, J. M. The diversity of microbial responses to nitric oxide and agents of nitrosative stress: Close cousins but not identical twins. Adv. Microb. Physiol. 59, 135–219 (2011).
Aldarini, N., Alhasawi, A. A., Thomas, S. C. & Appanna, V. D. The role of glutamine synthetase in energy production and glutamine metabolism during oxidative stress. Antonie Van Leeuwenhoek 110, 629–639 (2017).
Li, B., Wing, H., Lee, D., Wu, H.-C. & Busby, S. Transcription activation by Escherichia coli FNR protein: Similarities to, and differences from, the CRP paradigm. Nucleic Acids Res. 26, 2075–2081 (1998).
Wolf, Y. I., Rogozin, I. B., Kondrashov, A. S. & Koonin, E. V. Genome alignment, evolution of prokaryotic genome organization, and prediction of gene function using genomic context. Genome Res. 11, 356–372 (2001).
Krishnakumar, R. & Ruffing, A. M. OperonSEQer: A set of machine-learning algorithms with threshold voting for detection of operon pairs using short-read RNA-sequencing data. PLoS Comput. Biol. 18, e1009731 (2022).
Rastogi, R. P. et al. Molecular mechanisms of ultraviolet radiation-induced DNA damage and repair. J. Nucl. Acids 2010 (2010).
Ristow, M. Oxidative metabolism in cancer growth. Curr. Opin. Clin. Nutr. Metab. Care 9, 339–345 (2006).
Cheng, Z. & Ristow, M. Mitochondria and metabolic homeostasis (2013).
Hanage, W. P. Not so simple after all: Bacteria, their population genetics, and recombination. Cold Spring Harb. Perspect. Biol. 8, a018069 (2016).
Cox, M. M. Recombinational DNA repair in bacteria and the RecA protein. Prog. Nucl. Acid Res. Mol. Biol. 63, 311–366 (1999).
Li, Z., Pearlman, A. H. & Hsieh, P. DNA mismatch repair and the DNA damage response. DNA Repair 38, 94–101 (2016).
Zhang, H. et al. Apoptosis induced by overexpression of hMSH2 or hMLH1. Cancer Res. 59, 3021–3027 (1999).
Folkes, L. K. & O’Neill, P. DNA damage induced by nitric oxide during ionizing radiation is enhanced at replication. Nitric Oxide 34, 47–55 (2013).
Price, M. N., Arkin, A. P. & Alm, E. J. The life-cycle of operons. PLoS Genet. 2, e96 (2006).
Okuda, S. et al. Characterization of relationships between transcriptional units and operon structures in Bacillus subtilis and Escherichia coli. BMC Genomics 8, 1–12 (2007).
Im, S. et al. Comparative survival analysis of 12 histidine kinase mutants of Deinococcus radiodurans after exposure to DNA-damaging agents. Bioprocess Biosyst. Eng. 36, 781–789 (2013).
Selvam, K., Duncan, J. R., Tanaka, M. & Battista, J. R. DdrA, DdrD, and PprA: Components of UV and mitomycin C resistance in Deinococcus radiodurans R1. PLoS ONE 8, e69007 (2013).
Krueger, F., James, F., Ewels, P., Afyounian, E. & Schuster-Boeckler, B. Felixkrueger/trimgalore: v0.6.7 - via zenodo, https://doi.org/10.5281/zenodo.5127899 (2021).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Romero, L., Contreras-Riquelme, S., Lira, M., Martin, A. J. & Perez-Rueda, E. Homology-based reconstruction of regulatory networks for bacterial and archaeal genomes. Front. Microbiol. 13, 923105 (2022).
Kim, G. B., Gao, Y., Palsson, B. O. & Lee, S. Y. Deeptfactor: A deep learning-based tool for the prediction of transcription factors. Proc. Natl. Acad. Sci. 118, e2021171118 (2021).
Camacho, C. et al. Blast+: Architecture and applications. BMC Bioinform. 10, 1–9 (2009).
Consortium, U. Uniprot: A hub for protein information. Nucleic Acids Res. 43, D204–D212 (2015).
Münch, R. et al. Prodoric: Prokaryotic database of gene regulation. Nucleic Acids Res. 31, 266–269 (2003).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Quinlan, A. R. Bedtools: The swiss-army tool for genome feature analysis. Curr. Protocols Bioinform. 47, 11–12 (2014).
Cowen, L., Ideker, T., Raphael, B. J. & Sharan, R. Network propagation: A universal amplifier of genetic associations. Nat. Rev. Genet. 18, 551–562 (2017).
Pak, M. et al. Network propagation for the analysis of multi-omics data. In Recent Advances in Biological Network Analysis, 185–217 (Springer, 2021).
Pedregosa, F. et al. Scikit-learn: Machine learning in python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951 (2019).
Acknowledgements
This work was supported by National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (No. 2018R1D1A1B07050323). This work was supported by Institute of Information & communications Technology Planning & Evaluation (IITP) grant funded by the Korea government(MSIT) [No. 2021-0-01343, Artificial Intelligence Graduate School Program (Seoul National University)].
Author information
Authors and Affiliations
Contributions
Conceptualization, S.K., M.K., and S.S.; methodology, C.C., D.L., S.K., and S.S.; software, C.C.; formal analysis, C.C. and D.L.; investigation, C.C. and D.J.; data curation, C.C.; writing-original draft, S.S., S.K. and C.C.; writing-review and editing, C.C., D.L., D.J., S.K. and S.S.; visualization, C.C.; supervision, S.K. and S.S.; project administration, S.K., M.K. and S.S.; All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cho, C., Lee, D., Jeong, D. et al. Characterization of radiation-resistance mechanism in Spirosoma montaniterrae DY10T in terms of transcriptional regulatory system. Sci Rep 13, 4739 (2023). https://doi.org/10.1038/s41598-023-31509-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-31509-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.