Abstract
Salmonella enterica serovar Typhi is the causative agent of typhoid fever restricted to humans and does not replicate in commonly used inbred mice. Genetic variation in humans is far greater and more complex than that in a single inbred strain of mice. The Collaborative Cross (CC) is a large panel of recombinant inbred strains which has a wider range of genetic diversity than laboratory inbred mouse strains. We found that the CC003/Unc and CC053/Unc strains are permissive to intraperitoneal but not oral route of S. Typhi infection and show histopathological changes characteristic of human typhoid. These CC strains are immunocompetent, and immunization induces antigen-specific responses that can kill S. Typhi in vitro and control S. Typhi in vivo. Our results indicate that CC003/Unc and CC053/Unc strains can help identify the genetic basis for typhoid susceptibility, S. Typhi virulence mechanism(s) in vivo, and serve as a preclinical mammalian model system to identify effective vaccines and therapeutics strategies.
Similar content being viewed by others
Introduction
Salmonella enterica serovar Typhi (S. Typhi) is the causative agent of typhoid fever in humans. Global estimates reported by the CDC indicate that 21.6 million cases of typhoid fever occur each year resulting in 226,000 deaths1. The rapid emergence of multiple drug-resistant strains of S. Typhi now complicates the treatment of typhoid2. Typhoid is a vaccine-preventable disease and vaccination of high-risk populations, such as infants and young children, is considered the most promising strategy for control3. S. Typhi expresses Vi polysaccharide (ViPS), a well-known virulence factor4,5 and is a target for protective immune responses3,6. Three types of vaccines are currently available: (1) live attenuated vaccine, (2) subunit vaccines composed of plain ViPS7,8 and, (3) ViPS conjugated to a variety of carrier proteins such as rEPA, a recombinant Exoprotein A from P. aeruginosa9, CRM197, a non-toxic mutant of diphtheria toxin10, tetanus toxoid11, or diphtheria toxoid12.
Because S. Typhi is a human-restricted pathogen, vaccine efficacy data originates from clinical trials9,13. The live attenuated vaccine (Vivotif®) has an efficacy of ~ 60%7,8, does not express ViPS and is not recommended for children < 6 years of age due to safety concerns7. Subunit vaccines are safe for all ages, but the immune responses induced by plain ViPS vaccines, such as Typhim Vi® or Typherix®, are short-lived and their efficacy is ~ 55% in older children and adults7. Importantly, plain ViPS vaccines do not induce optimal antibody responses in children under 2 years of age. Fortunately, ViPS conjugated to rEPA, and tetanus toxoid can induce anti-ViPS responses in infants and young children with 80–90% efficacy9,13. Although antibodies are considered as the correlate of protection, the lack of a preclinical animal model makes it difficult to understand the variation in efficacy and the mechanism of protection conferred by these vaccines.
Ethical considerations make it difficult to fully characterize histopathology and immunology relevant to typhoid fever in humans8,14. A human S. Typhi challenge model was carried out in the UK, a non-disease endemic country. In this controlled challenge model, 4 of 40 participants fulfilled predetermined criteria for typhoid infection15. Nevertheless, this model enabled the accelerated testing of the newly WHO-prequalified ViPS-tetanus toxoid conjugate vaccine16. However, this human model will not provide an in-depth understanding of the factors involved in host susceptibility/resistance or the in vivo mechanism(s) of protective immunity to the extent that experimental animal models could provide. Commonly used inbred mouse strains e.g., C57BL/6J or BALB/cJ are not susceptible to S. Typhi infection. Previous research had demonstrated that Tlr11−/− mice permit S. Typhi infection17; because humans do not have the gene-encoding for Tlr11, it was hypothesized that might account for the resistance of mice to S. Typhi. For reasons that are not clear, several other researchers including the original researchers, failed to reproduce Tlr11−/− mouse permissiveness to S. Typhi18,19. Several research groups demonstrated that CD34+ human hematopoietic stem cell-engrafted NOD/SCID/IL-2Rγcnull or NOD/SCID/IL-2Rγcnull mice, referred to as “Human Immune System” or “humanized” mice are susceptible to S. Typhi infection20,21,22. Although humanized mice have the potential to understand typhoid pathogenesis, the humoral immunity of these mice is immature and such mice do not respond to ViPS immunization23,24. Therefore, the “humanized” mouse model in its current form is not appropriate for understanding immune responses to S. Typhi. Thus, there is a strong need to explore a fully immunocompetent and S. Typhi-permissive mouse model to decipher the mechanisms of typhoid vaccine-mediated protective immune responses.
Genetic variation in humans is far greater and more complex than that in commonly used inbred laboratory mice such as C57BL/6J, which may in part provide an explanation for differences in the infection susceptibility and progression in humans. The establishment of a murine model for human typhoid requires a system for the generation of mouse strains with great genetic diversity in which characterization can take place. A panel of recombinant inbred (RI) strains has been developed that incorporates a wider range of genetic diversity than inbred mouse strains. This set of RI strains is called the Collaborative Cross or CC25. The CC strains were derived from 8 genetically diverse inbred progenitor strains: A/J, C57BL/6J, 129S1/SvImJ, NOD/ShiLtJ, NZO/H1J, CAST/EiJ, PWK/PhJ and WSB/EiJ (https://csbio.unc.edu/CCstatus/index.py). The CC strains were designed specifically for complex trait analysis25. Each CC strain contains portions of the genomes of the 8 progenitor strains and each CC strain has been bred to homozygosity (> 90%). This new resource is supporting Systems Genetics and Systems Biology research at an unprecedented level. In fact, CC strains have been shown to serve as powerful model to identify host resistance and susceptibility alleles governing infections due to a variety of viral, bacterial and fungal pathogens26, including Influenza A virus27, SARS Corona virus28 Ebola virus29, Mycobacterium tuberculosis30, Salmonella enterica serovar Typhimurium (S. Typhimurium)31,32 and Aspergillus fumigatus33.
In the present study, we report the identification of immunocompetent S. Typhi-susceptible CC strains. The CC003/Unc and CC053/Unc strains are permissive to S. Typhi infection, intraperitoneally, but not through the oral route of infection. The CC mouse model described here will not only provide a much-needed preclinical tool for assessing the efficacy of vaccine candidates against typhoid but will also permit genetic analyses for S. Typhi susceptibility/resistance.
Results
CC mice are permissive to S. Typhi replication in vivo
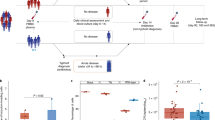
Since S. Typhi is a human-restricted pathogen, researchers often employ S. Typhimurium, a closely related bacterium that causes “typhoid-like” disease in mice. Organs of the mononuclear phagocytic system, such as spleen and liver, are the major sites of replication of S. Typhi in humans and this pattern is well-reflected in the i.p. route of infection of S. Typhimurium in mice34,35. To test whether CC mice are permissive or not, we infected 9 different CC mouse strains, 4 CC progenitors and BALB/cJ of both sexes i.p. with 2 × 104 CFUs of S. Typhi strain Ty2. We assessed bacterial burden in the spleen 6 days post-infection to allow time for a detectable S. Typhi replication or host clearance. BALB/cJ, all 4 CC progenitor strains (C57BL/6J, 129S1/SvImJ, NOD/ShiLtJ, and CAST/EiJ), and several CC strains (CC035/Unc, CC030/GeniUnc, CC022/GeniUnc, and CC052/GeniUnc) showed little-to-no bacteria in the liver (Fig. 1). The bacterial count in the spleens of all these mice was lower than or close to the number of bacteria initially injected (Fig. 1). Some CC017/Unc females, CC038/GeniUnc and CC055/TauUnc mice showed a marginal increase in the bacterial burden in the liver and spleen (Fig. 1). Although an increase in bacterial load was not seen consistently in the livers of all CC003/Unc and CC053/Unc mice, the bacterial load in the spleens of these mice was significantly higher than that observed in the commonly used laboratory strains BALB/cJ, C57BL/6J, and 129S1/SvImJ (Fig. 1), indicating bacterial growth in these two CC strains. Interestingly, we observed a significant difference in the bacterial burden between male and female CC003/Unc mice (Fig. 1), a pattern not observed in the spleens of CC053/Unc mice (Fig. 1). These data suggest that CC053/Unc males and females, and CC003/Unc males but not females are permissive to S. Typhi replication in vivo.
Bacterial burden in the liver and spleen of mice infected with S. Typhi. Inbred mouse strains, CC progenitors and CC strains of both sexes (age 8–12 weeks) were infected i.p. with 2 × 104 CFUs of S. Typhi strain Ty2 in 100 μl of DPBS. Six days post-infection mice were sacrificed and bacterial burden in the liver and spleen was determined by plating serial tenfold dilutions of tissue homogenates followed by colony counting. Each open circle represents an individual mouse, the red bar represents median and interquartile range to identify statistical dispersion and range of susceptibility. Dotted line indicates the limit of detection. Statistics were performed using the Mann Whitney U test. ** = p < 0.01; * = p < 0.05; N.S. = not significant.
Histological features of S. Typhi infection in the livers of CC mice recapitulates that of human and murine typhoid pathology
Since preliminary screening of CC strains suggested the permissiveness to S. Typhi infection in CC003/Unc and CC053/Unc mice (Fig. 1), we examined hematoxylin and eosin-stained liver specimens of these two CC strains to characterize histopathology. We found that the livers of CC003/Unc mice had normal tissue architecture (Fig. 2A,B). However, upon S. Typhi infection the liver tissue showed signs of steatosis, an abnormal retention of lipids within hepatocytes (Fig. 2D), as compared to uninfected mice (Fig. 2C). This indicates a perturbation of liver metabolism in infected mice. Furthermore, S. Typhi-infected CC003/Unc and CC053/Unc mouse livers exhibited lesions that consist of mononuclear cell infiltrates, congestion of sinusoids, and altered staining with little/no steatosis in hepatocytes proximal to the lesion (Fig. 2E–H). While tissue biopsies are rare in typhoid-infected patients, biopsies have been collected in cases where typhoid is not initially considered and are used to aid diagnosis36. These rare cases provide some of the only information known about the histopathology of typhoid. Interestingly, the lesions observed in CC003/Unc and CC053/Unc mice (Fig. 2E–H) appear very similar to those described for human typhoid patients37. As a comparison, we also show that S. Typhimurium infection results in similar liver pathology that is commonly observed in inbred mice, C57BL/6J (Fig. 2I,J) and 129S1/SvImJ (Fig. 2K,L).
Histological features of S. Typhi infection in the livers of CC mice. Mice (age 8–12 weeks) were infected as in Fig. 1. The data presented is representative of five male and five female mice. Hematoxylin and eosin-stained liver sections of 6-day post-infected mice for S. Typhi and 3-day post-infected mice for S. Typhimurium were analyzed. Panel A, B and C show representative livers of the uninfected CC003/Unc mice at 3 different magnifications with an arrowhead pointing to a Kupffer cell. Panel D shows patterns of steatosis (indicated with an arrow) in infected mice, which is absent in uninfected mice (Panel C). Panels E–H depict representative liver sections of S. Typhi-infected male and female CC003/Unc and CC053/Unc mice. Panels I–L are liver sections of S. Typhimurium-infected C57BL/6J and 129S1/SvImJ mice at two different magnifications. Black arrows indicate lesions. Horizontal black bars in the right bottom corner show the size in μm.
CC003/Unc and CC053/Unc mice contain all the major subsets of cells of the innate and adaptive immune system
Susceptibility of mice to an infection can be due to immune deficiency. Since immune competency of CC mice is necessary for studying protective responses to S. Typhi in preclinical models, we performed a comprehensive flow cytometric analysis of cells in the spleen and coelomic cavity of CC003/Unc and CC053/Unc mice, as well as C57BL/6J mice as a control. In mice, the mature B cells can be divided into 4 subsets, namely, Fo, MZ, B1a, and B1b cells. Each of these subsets occupy a distinct functional niche in protective immunity38. For example, antibody responses to ViPS in mice are generated primarily by B1b cells39. We found that all 4 major B cell subsets including B1b are present in CC003/Unc and CC053/Unc mice and their frequency is within the range observed in C57BL/6J mice, except the frequency of MZ B cells, which is higher in the CC mice compared to C57BL/6J mice (Fig. 3A vs. B,C). The ratio of CD4 to CD8 T cells in C57BL/6J mice is typically 2–3:140, and we found a slightly altered ratios in CC003/Unc and CC053/Unc mice (Fig. 3). It has been reported recently that there is a high variablility in the CD4 and CD8 ratios as well as total B cell frequencies in CC strains41. Furthermore, in the spleens of CC003/Unc and CC053/Unc mice, the frequency of natural killer (NK) cells (CD3−, NKp46+) neutrophils (CD11b+, Gr1+ & F4/80−), macrophages (CD11b+ & F4/80+), and dendritic cells (CD11b+/lo, CD11c+ & F4/80−) were 2–4%, 1.5–2.5%, 4–9%, and 5–9%, respectively. These frequencies are within the range reported previously for C57BL/6, BALB/c and 129/Sv mice40. These data indicate that all the major innate and adaptive immune cell compartments are intact in CC003/Unc and CC053/Unc mice, and therefore enabling the study of protective immune responses against S. Typhi in vitro and in vivo.
Flow cytometric analysis of mature lymphocyte subsets in CC mice. Spleen cells of mice (age 8–12 weeks) were stained with a cocktail of antibodies specific for CD19, B220, CD21, and CD23 or a cocktail of antibodies specific for CD3, CD4 and CD8, and analyzed by flow cytometry. Splenic B cells were first identified as CD19+ and B220+ cells (plots not shown) and were resolved further as MZ (CD23lo and CD21hi) and Fo (CD23hi and CD21lo) B cells, and their frequencies among CD19+ & B220+ cells are indicated within the plots. Splenic T cells were first identified as CD3+ (plots not shown) and resolved them into CD4+ and CD8+ cells. Since B1b and B1a cells are found in abundance in the coelomic cavity of mice, peritoneal cells were stained with antibodies specific for CD19, CD11b, surface IgM and CD5. All cells were first identified as B1 cells (CD19+ & CD11b+ cells [plots not shown]). The frequency of B1a (IgM+ & CD5+) and B1b (IgM+ & CD5−) subsets were shown. All data were generated by analyzing 50,000–100,000 cells and are representative of 3–5 mice from each mouse strain. Five percent contour plots are shown.
CC003/Unc mice are immunocompetent
Among the 9 CC strains of both sexes screened, CC003/Unc male mice showed relatively high susceptibility to S. Typhi replication as determined by the bacterial load in the liver and spleen in more than 6 independent experiments comprising of a total of 43 mice (Fig. 1). To test the proof of principle that CC mice can generate protective immunity against S. Typhi, we chose the CC003/Unc strain to test vaccination efficacy. We immunized male CC003/Unc mice with either plain ViPS or heat-killed S. Typhi. We found that both types of immunizations induced a robust anti-ViPS IgM response that peaked at 7 days post-infection (Fig. 4IA,IIA) as in immunocompetent C57BL/6J mice35 and the serum obtained from the CC003/Unc mice at this time point was capable of killing S. Typhi in vitro in a complement-dependent serum bactericidal assay (Fig. 4IC,IIC). IgM response typically declines after 7 days post-infection, which is concurrent with the induction of IgG response due to antibody isotype switching35. Both immunizations generated an anti-ViPS IgG response by 21 days post-infection (Fig. 4IB,IIB), and serum obtained at this time point also killed S. Typhi in vitro (Fig. 4IC,IIC). These data suggest that CC003/Unc mice undergo isotype switching normally and the antibodies generated by these immunizations can provide protection. To test whether anti-ViPS antibodies confer protection in vivo, we challenged male CC003/Unc mice with S. Typhi strain Ty2 and measured bacterial load 6 days later as in Fig. 1. Although ViPS-immunized mice exhibited a reduced bacterial burden in the liver and spleen compared to unimmunized mice, the difference was not statistically significant (Fig. 4ID,E), consistent with the low efficacy of the plain ViPS vaccine7. Immunization with whole bacteria induces a qualitatively different antibody response42. In a S. Typhimurium “surrogate” challenge model, we have previously shown that immunization with heat-killed S. Typhi controls bacterial burden better than ViPS immunization35. Here we found that immunization of CC003/Unc mice with heat-killed S. Typhi confers more efficient protection compared to ViPS, as determined by a significant decrease in bacterial burden in the liver and spleen (Fig. 4IID,E).
CC mice are immunocompetent. Eight to twelve-week-old CC003/Unc male mice were immunized i.p. either with (I) 2.5 μg ViPS or (II) 3 × 108 CFUs of heat-killed S. Typhi strain Ty2 in 100 μl of DPBS. ViPS-specific IgM (A) and IgG (B) on indicated time points were measured by ELISA. Each open circle represents an individual mouse. The solid bar represents mean values. (C) Serum bactericidal antibody titers against live S. Typhi strain Ty2 were determined at 7- and 21-days post-immunization. (D,E) Unimmunized and immunized mice (21 days post-immunization) were infected i.p. with 2 × 104 CFUs of S. Typhi strain Ty2 in 100 μl of DPBS. Six days after challenge, bacterial burden in liver and spleen were determined as in Fig. 1. The dotted line in panels D&E indicates detection limits. Data for all figures is a pool of minimum of two independent experiments. Statistical differences were determined by the Mann–Whitney U test.
Unlike adults, young children and infants do not respond to polysaccharide antigens such as ViPS. We have previously shown that 3-week-old mice (like young children) do not respond to ViPS efficiently due to a restricted antibody repertoire43. Therefore, we compared antibody responses to ViPS in young (3-week-old) and adult (8–12-week-old) CC003/Unc mice. We found that compared to adult CC003/Unc mice, the 3-week-old CC003/Unc mice are not capable of mounting an efficient IgM response to ViPS (Fig. 5A) and their serum was inefficient in killing S. Typhi in vitro (Fig. 5B). In summary, this data demonstrate that CC mice are not only immunocompetent but can control S. Typhi burden upon immunization and suggest that both young and adult CC003/Unc mice can serve as an experimental system to identify novel typhoid vaccines and therapeutics.
Young CC003/Unc mice do not respond as efficiently as adult CC003/Unc to ViPS immunization. (A) Three-week-old (young) or 8- to 12-week-old (adult) CC003/Unc mice were immunized with ViPS as in Fig. 4, and the levels of ViPS-specific IgM and IgG were measured by ELISA. Statistical differences were determined using two-way ANOVA. Each open circle represents a young mouse, and each filled circle represents an adult mouse. The bar represents the mean. (B) Serum bactericidal titers against S. Typhi strain Ty2 were determined at 7 days post-immunization. Statistical differences were determined by the Mann–Whitney U test. The bar represents the geometric mean. The data are pooled from two independent experiments. ***p < 0.001, ****p < 0.0001. N.S., statistically not significant.
Discussion
One of the barriers to advancing the treatment and prevention of typhoid is the lack of a suitable animal model to study S. Typhi infection8. Since S. Typhi is strictly human adapted, S. Typhimurium, a natural pathogen of mice became the most widely used bacterium to understand pathogenesis and immunity in several inbred strains (e.g., C57BL6/J and 129/Sv). S. Typhimurium causes a “Typhoid-like” systemic disease in mice and shares 90% of genes with S. Typhi44. However, ~ 600 S. Typhi genes, including those encoding for ViPS biogenesis and typhoid toxin, are not found in S. Typhimurium44. Furthermore, several genes found in S. Typhimurium are pseudogenes in S. Typhi45. Therefore, wildtype S. Typhimurium strains cannot serve as a synonymous model system to decipher the role of S. Typhi-specific virulence factors. To understand the function of ViPS in vivo, Baumler and coworkers introduced all the S. Typhi genes required for ViPS biogenesis in S. Typhimurium46. Using this “chimeric” S. Typhimurium strain e.g. TH17046, a role for ViPS in resisting C3 deposition and complement receptor 3-mediated clearance47, in evading of TLR4 recognition48,49, and microbe-guided neutrophil chemotaxis was identified50,51. The length of LPS of S. Typhimurium is very long compared to that of S. Typhi. In S. Typhimurium the length of LPS is controlled by the FepE gene product, which is a pseudogene in S. Typhi. To mimic the surface characteristics of S. Typhimurium to resemble that of S. Typhi, the S. Typhimurium strain TH170 was further engineered by deleting the FepE gene4. This strain of S. Typhimurium, referred to as RC60, was shown to exhibit cell surface and other characteristics of S. Typhi4. Using the S. Typhimurium strain RC60, we were able to identify several aspects of the anti-ViPS antibody repertoire required for protective immunity in vivo35,43,52. Thus, the use of S. Typhimurium as a “surrogate” model to understand certain virulence mechanisms of S. Typhi has been justified. However, the heterologous expression of S. Typhi genes in S. Typhimurium may not always permit identification of the role of certain S. Typhi-specific components. For example, the typhoid toxin of S. Typhi appears to be host-adapted and binds to N-acetylneuraminic acid (Neu5Ac) that is abundantly expressed in humans but not in mice53,54. Therefore, studying the role of typhoid toxin in the pathogenesis requires a permissive mouse model that contains human tissue, such as “humanized” mice. Interestingly, a genome-wide analysis of S. Typhi mutants in “humanized" mice confirmed that ViPS is essential for virulence but surprisingly not the typhoid toxin55. The controlled human infection model also revealed that typhoid toxin is neither required for infection nor for the development of early typhoid fever symptoms56. Since the human challenge model represents early events of typhoid, the role of typhoid toxin in severe disease or the establishment of bacterial carriage remains to be determined.
Host genetics clearly play a role in a variety of infectious diseases, as evidenced by hypothesis-driven analyses of polymorphisms in the Tlr4 gene57. In fact, CC mice were developed as a resource for mammalian systems genetics25. Few susceptibility genes e.g., Slc11a1 and Tlr11 have been implicated for the permissiveness of S. Typhi. Notably, the relative susceptibility or resistance to S. Typhimurium, a close relative of S. Typhi, was shown to depend upon a single nucleotide polymorphism in the Slc11a1 gene that encodes an ion transporter commonly referred to as Natural resistance-associated macrophage protein 1 (Nramp1)58 and neutrophil cytosolic factor 2 (Ncf2)59. In the UNC systems genetics database, (https://csbio.unc.edu/CCstatus/index.py), we found that both CC003/Unc and CC053/Unc mice have the Slc11a1 resistant allele inherited from the progenitor A/J and the ncf2 allele is not mutated60. Although the role of Tlr11 is an ongoing controversy18,19, we have not found any mutation in the Tlr11 gene either in CC003/Unc or CC053/Unc mice. Therefore, a comprehensive genetic approach may help us discover novel genes responsible for susceptibility to S. Typhi in CC003/Unc and CC053/Unc. For example, one approach is to perform classical genetic crosses to generate F1 hybrids and F2 offspring by crossing either CC053/Unc or CC003/Unc with BALB/cJ, a known S. Typhi resistant and a non-CC progenitor strain, respectively. The susceptibility of F1 hybrids and F2 offspring to S. Typhi can be tested as in Fig. 1, and the genotype of those mice can be determined using the Giga Mouse Universal Genotyping Array (GigaMUGA)61 or MiniMUGA62 to find associations between resistance/susceptibility phenotype and genotype. Indeed, using a combination of genotyping and quantitative trait loci mapping, several S. Typhimurium infection susceptibility loci have recently been identified in CC042/GeniUnc mice32. Although CC042/GeniUnc mice inherited the susceptible Slc11a1 locus from the C57BL/6 progenitor and its Tlr4 gene product is functional, the bacterial burden in the CC042/GeniUnc mice was 1000-fold more than that in C57BL/6N mice, suggesting that the hyper-susceptibility is due to other factors. Interestingly, CC042/GeniUnc mice also showed lower spleen weights and decreased B, T, and myeloid cell populations compared to control C57BL/6N mice, suggesting that an abnormality in the architecture of the immune system or a partial immune deficiency may be responsible for the hyper-susceptible phenotype. Indeed, an F2 cross between CC042/GeniUnc and C57BL/6N mice identified a susceptibility locus accompanied by a loss-of-function variant of the integrin alpha L (Itgal) gene31, which encodes for LFA-1, a molecule central to immune cell adhesion and trafficking. This hyper-susceptibility phenotype has been confirmed independently by comparing S. Typhimurium infection in C57BL/6N and LFA-1−/− mice31. Unlike the CC042/GeniUnc mouse system for S. Typhimurium hyper-susceptibility, the reason for CC003/Unc and CC053/Unc mice susceptibility is unlikely to be associated with an altered architecture/composition of the immune system. The Itgal gene of CC003/Unc and CC053/Unc mice is inherited from C57BL/6J and 129S1/SvImJ progenitors, but not from WSB/EiJ progenitor as in CC042/GeniUnc mice. Most importantly, CC003/Unc and CC053/Unc mice are immunocompetent (unlike CC042/GeniUnc or “humanized” mice) and possess all the major populations of innate and adaptive immune cells (Fig. 3) and respond to both T cell-independent immunogen (i.e., isolated plain ViPS) or T cell-dependent immunogen (i.e., heat-killed S. Typhi) (Fig. 4).
Feaco-oral route is a natural mode of S. Typhi transmission in humans, that eventually results in a systemic infection. When we attempted oral infection using a gavage needle (109 CFU of strain Ty2 in 100 μl PBS), we did not observe permissiveness of S. Typhi infection in CC003/Unc mice. Therefore, we chose i.p. infection which is commonly used method for sytemic infection in a variety of infectious disease models in mice. This suggests that the CC003/Unc mice do not capture certain aspects of typhoid disease that occurs in humans, such as intestinal pathology, and bacterial shedding in faeces. Screening more CC strains to S. Typhi susceptibility might help identify specific CC strains that can capture the oral infection characteristics and intestinal pathology seen in humans.
In conclusion, the immunocompetent, S. Typhi permissive CC mouse model using CC003/Unc and CC053/Unc presented here can provide an in vivo experimental system to evaluate novel preventive and therapeutic interventions. Additionally, genomic analyses of the CC mouse model can help us understand why responses to different ViPS conjugate vaccines vary among populations in disease-endemic countries10 and why some individuals become chronic and asymptomatic carriers for spreading typhoid. This CC mouse model system can also enable the S. Typhi research community to identify putative S. Typhi virulence factor(s) and characterize their role in the progression of typhoid.
Methods
Mice
The Thomas Jefferson University (TJU) Institutional Animal Care and Use Committee have approved these studies. All methods were performed in accordance with the guidelines and regulations of TJU Institutional Biosafety Committee. Reporting of the animal experiments followed the recommendations in the ARRIVE guidelines. C57BL/6J, 129S1/SvImJ, BALB/cJ, NOD/ShiLtJ and CAST/EiJ were purchased from The Jackson Laboratory (Bar Harbor, ME). All CC strains used were purchased in 2014–2015 from the Systems Genetics Core Facility, University of North Carolina at Chapel Hill (Chapel Hill, NC). Mice were housed in micro-isolator cages with free access to food and water and were maintained and bred in a specific pathogen-free facility at TJU. Three-week-old mice were considered young and 8–12-week-old mice were considered adult.
Infection
For mouse infections S. Typhi strain Ty2, a well-studied strain (obtained from Dr. Andreas Baumler) was grown to an OD600 of ~ 1.0 in Luria Bertani (LB) broth containing 10 mM NaCl. Bacteria were washed twice in Dulbecco’s phosphate-buffered saline (DPBS); and bacterial density was adjusted to 2 × 105 colony forming units (CFU)/ml. Mice were infected intraperitoneally (i.p.) with 2 × 104 CFU in 100 μl of DPBS. Because organs of the mononuclear phagocytic system, such as spleen and liver, are the major sites of replication of S. Typhi in humans, we assessed bacterial burden in these organs 6 days later to allow time for a detectable S. Typhi replication or host clearance. On day 6 post-infection, liver and spleen were collected and tissues were processed using a Minilys tissue homogenizer (Bertin Technologies, Montigny-le-Bretonneux, France). Bacterial burden in these tissue homogenates was measured by counting CFU on LB agar plates.
Histopathology analysis
Liver tissues obtained on day 6 post-infection were fixed in 10% buffered formalin and 4 μM paraffin-embedded sections were stained with hematoxylin and eosin. The specimen slides were scanned at 20 × magnification on Aperio CS2 Scanscope® (Leica Biosystems Inc.) followed by observer-blind histopathological analysis.
Flow cytometry
Peritoneal cavity cells were harvested to determine the frequency of B1a and B1b cells, and spleen cells were harvested to determine the frequency of follicular (Fo) B, marginal zone (MZ) B, CD4+T, CD8+T, Natural Killer (NK), dendritic cells, neutrophils, and macrophages from individual mice. The cell concentrations were adjusted to 2.5 × 107 cells/ml in staining medium [Minimum Essential Medium Eagle with Earle’s salts and without L-glutamine or phenol red (Corning Cellgro, Manassas, VA) with 3% newborn calf serum (HyClone Laboratories Inc. Logan, UT), 1 mM EDTA]. Cell suspensions were incubated with 2.4G2 antibody for 15 min to block Fc receptors and stained with appropriate antibody cocktail. The antibodies, anti-CD11b-AF700 (clone: M1/70), anti-CD5 Pacific Blue (clone: 53–7.3), anti-CD19-PE-Cy7 (clone: eBio1D3) were purchased from eBioscience (San Diego, CA); anti-B220-PercP-Cy5.5 (clone: RA3-6B2) was purchased from Caltag (Burlingame, CA); anti-IgM-PE (clone: 1B4B1) and CD23-PE (clone: B3B4) was purchased from BD Pharmingen (San Jose, California). Anti-CD3 FITC (clone:17A2), anti-CD4-BV605 (clone: RM4-5), anti-CD8-APC-Cy7 (clone: 53–6.7), anti-NKp46-APC (clone: 29A14), anti-F4/80-FITC (clone: BM8E), anti-GR1-Pacific blue (clone: RB6-8C5), anti-CD11c-PE-Cy7 (clone: N418) and anti-CD21/35-APC (clone: 7E9) were purchased from Biolegend. (San Diego, CA). After staining, cells were washed twice with staining medium and the preparations were analyzed on an LSRII™ flow cytometer (Becton Dickinson, Mountain View, CA) using the FACS Diva™ software (Becton Dickinson). Data were analyzed using the FlowJo™ software (Treestar, San Carlos, CA).
Immunization
Two and a half μg of Vi Polysaccharide (ViPS; Lot 5 PDMI 158,299 obtained from the U.S. Food and Drug Administration, Silver Spring, MD) dissolved in 100 µl DPBS was used to immunize mice i.p. For whole bacterial immunization, mice were injected i.p. with 3 × 108 CFUs of heat-killed S. Typhi strain Ty2 in 100 µl DPBS35. The expression of ViPS is confirmed by serologically by slide agglutination test using a commercial Vi monoclonal antibody reagent (Statens Serum Institut diagnostica A/S, Denmark; Lot 188L-8). We also perform serum bactericidal assay using anti-S. Typhi human IgG (Lot R1, 2011; U.S. Food and Drug Administration, Silver Spring, MD) as described35,52,63. Blood samples were obtained 0-, 7-, 14- or 21-days following immunization and stored at − 20 °C.
Enzyme-linked immunosorbent assay (ELISA)
ViPS-specific IgM and IgG were measured by coating 96-well microtiter plates (Nunc MaxiSorp™; Invitrogen, Carlsbad, CA) with 2 µg/ml of ViPS purified from S. Typhi clinical isolate C652464 in DPBS overnight at room temperature. All plates were washed and blocked with 2% Bovine serum albumin (BSA) in PBS pH 7.2 (blocking buffer) for 2 h at room temperature. Blood from immunized mice was diluted to 1:25 for IgG detection and 1:50 for IgM detection in blocking buffer; ViPS-specific mouse IgM and IgG levels were interpreted as ng/μl “equivalents” using normal mouse serum standards (Bethyl Laboratories, Montgomery, TX), as described previously35.
Serum bactericidal assay (SBA)
SBA was performed as previously described35. In brief, log-phase cultures (OD600 of 0.5 at 37 °C) of S. Typhi strain Ty2 were prepared in LB broth with 10 mM NaCl. Bacterial cells were washed in DPBS, and the bacterial cell density was adjusted to 1–3.5 × 104 CFU/ml in DPBS. Serum samples were heat-inactivated by incubating at 56 °C for 30 min prior to use in the assay. Ten microliters of S. Typhi cells in DPBS (100–350 CFU) were added to each well of a round-bottom polypropylene 96-well plate containing 50 μl of heat-inactivated serum in serial dilutions, 12.5 μl baby rabbit complement (Pel-Freeze, Rogers, AR), and 27.5 μl DPBS. Triplicate samples of each dilution were incubated for 120 min at 37 °C with gentle rocking, and 10 μl of this mixture was plated on LB agar plates for enumerating bacterial CFU. Serum bactericidal antibody titers were defined as the reciprocal of the highest dilution that produced > 50% killing in relation to control wells containing complement, but no mouse serum.
Statistical analysis
Data presented throughout depict pooled data from at least two independent experiments unless otherwise noted. Statistics were performed using the Prism 5 software program (GraphPad Software, Inc., La Jolla, CA).
Data availability
All data generated or analysed during this study are included in this published article.
References
Crump, J. A. & Mintz, E. D. Global trends in typhoid and paratyphoid fever. Clin. Infect. Dis. 50, 241–246 (2010).
Britto, C. D., Wong, V. K., Dougan, G. & Pollard, A. J. A systematic review of antimicrobial resistance in Salmonella enterica serovar Typhi, the etiological agent of typhoid. PLoS Negl. Trop. Dis. 12, e0006779. https://doi.org/10.1371/journal.pntd.0006779 (2018).
Birkhold, M., Mwisongo, A., Pollard, A. J. & Neuzil, K. M. Typhoid conjugate vaccines: Advancing the research and public health agendas. J. Infect. Dis. 224, S781–S787. https://doi.org/10.1093/infdis/jiab449 (2021).
Crawford, R. W. et al. Loss of very-long O-antigen chains optimizes capsule-mediated immune evasion by Salmonella enterica serovar Typhi. Bio 4, e00232-13 (2013).
Raffatellu, M. et al. Capsule-mediated immune evasion: A new hypothesis explaining aspects of typhoid fever pathogenesis. Infect. Immun. 74, 19–27. https://doi.org/10.1128/IAI.74.1.19-27.2006 (2006).
Jin, C. et al. Vi-specific serological correlates of protection for typhoid fever. J. Exp. Med https://doi.org/10.1084/jem.20201116 (2021).
Levine, M. M. Typhoid Fever Vaccine 5th edn. (Saunders Elsevier, 2008).
Yang, Y. A., Chong, A. & Song, J. Why is eradicating typhoid fever so challenging: Implications for vaccine and therapeutic design. Vaccines (Basel) https://doi.org/10.3390/vaccines6030045 (2018).
Lin, F. Y. et al. The efficacy of a Salmonella Typhi Vi conjugate vaccine in two-to-five-year-old children. N. Engl. J. Med. 344, 1263–1269 (2001).
Bhutta, Z. A. et al. Immunogenicity and safety of the Vi-CRM197 conjugate vaccine against typhoid fever in adults, children, and infants in south and Southeast Asia: Results from two randomised, observer-blind, age de-escalation, phase 2 trials. Lancet Infect. Dis. 14, 119–129 (2014).
Mohan, V. K. et al. Safety and immunogenicity of a Vi polysaccharide-tetanus toxoid conjugate vaccine (Typbar-TCV) in healthy infants, children, and adults in typhoid endemic areas: A multicenter, 2-cohort, open-label, double-blind, randomized controlled phase 3 study. Clin. Infect. Dis. 61, 393–402. https://doi.org/10.1093/cid/civ295 (2015).
Medise, B. E. et al. A novel Vi-diphtheria toxoid typhoid conjugate vaccine is safe and can induce immunogenicity in healthy Indonesian children 2–11 years: A phase II preliminary report. BMC Pediatr. 20, 480. https://doi.org/10.1186/s12887-020-02375-4 (2020).
Qadri, F. et al. Protection by vaccination of children against typhoid fever with a Vi-tetanus toxoid conjugate vaccine in urban Bangladesh: A cluster-randomised trial. Lancet 398, 675–684. https://doi.org/10.1016/S0140-6736(21)01124-7 (2021).
Waddington, C. S. et al. Advancing the management and control of typhoid fever: A review of the historical role of human challenge studies. J. Infect. 68, 405–418 (2014).
Waddington, C. S. et al. An outpatient, ambulant-design, controlled human infection model using escalating doses of Salmonella Typhi challenge delivered in sodium bicarbonate solution. Clin. Infect. Dis. 58, 1230–1240 (2014).
Meiring, J. E., Giubilini, A., Savulescu, J., Pitzer, V. E. & Pollard, A. J. Generating the evidence for typhoid vaccine introduction: Considerations for global disease burden estimates and vaccine testing through human challenge. Clin Infect Dis 69, S402–S407. https://doi.org/10.1093/cid/ciz630 (2019).
Mathur, R. et al. A mouse model of Salmonella Typhi infection. Cell 151, 590–602 (2012).
Song, J. et al. Absence of TLR11 in mice does not confer susceptibility to Salmonella Typhi. Cell 164, 827–828 (2016).
Mathur, R., Zeng, W., Hayden, M. S. & Ghosh, S. Mice lacking TLR11 exhibit variable Salmonella Typhi susceptibility. Cell 164, 829–830 (2016).
Libby, S. J. et al. Humanized nonobese diabetic-scid IL2rγnull mice are susceptible to lethal Salmonella Typhi infection. Proc. Natl. Acad. Sci. U.S.A. 107, 15589–15594. https://doi.org/10.1073/pnas.1005566107 (2010).
Song, J. et al. A mouse model for the human pathogen Salmonella Typhi. Cell Host Microbe 8, 369–376. https://doi.org/10.1016/j.chom.2010.09.003 (2010).
FirozMian, M., Pek, E. A., Chenoweth, M. J. & Ashkar, A. A. Humanized mice are susceptible to Salmonella Typhi infection. Cell Mol. Immunol. 8, 83–87 (2011).
Walker, J. A., Vuyyuru, R., Manser, T. & Alugupalli, K. R. Humoral Immunity in mice transplanted with hematopoietic stem cells derived from human umbilical cord blood recapitulates that of human infants. Stem Cells Dev. 26, 1715–1723. https://doi.org/10.1089/scd.2017.0156 (2017).
Vuyyuru, R., Liu, H., Manser, T. & Alugupalli, K. R. Characteristics of Borrelia hermsii infection in human hematopoietic stem cell-engrafted mice mirror those of human relapsing fever. Proc. Natl. Acad. Sci. U.S.A. 108, 20707–20712 (2011).
Collaborative Cross Consortium. The genome architecture of the Collaborative Cross mouse genetic reference population. Genetics 190, 389–401 (2012).
Noll, K. E., Ferris, M. T. & Heise, M. T. The collaborative cross: A systems genetics resource for studying host-pathogen interactions. Cell Host Microbe 25, 484–498. https://doi.org/10.1016/j.chom.2019.03.009 (2019).
Ferris, M. T. et al. Modeling host genetic regulation of influenza pathogenesis in the collaborative cross. PLoS Pathog 9, e1003196 (2013).
Xiong, H. et al. Genomic profiling of collaborative cross founder mice infected with respiratory viruses reveals novel transcripts and infection-related strain-specific gene and isoform expression. G3 (Bethesda) 4, 1429–1444 (2014).
Rasmussen, A. L. et al. Host genetic diversity enables Ebola hemorrhagic fever pathogenesis and resistance. Science 346, 987–991 (2014).
Smith, C. M. et al. Functionally overlapping variants control tuberculosis susceptibility in collaborative cross mice. MBio https://doi.org/10.1128/mBio.02791-19 (2019).
Zhang, J. et al. A loss-of-function mutation in the integrin alpha L (Itgal) gene contributes to susceptibility to Salmonella enterica serovar typhimurium infection in collaborative cross strain CC042. Infect. Immun. https://doi.org/10.1128/IAI.00656-19 (2019).
Zhang, J. et al. Identification of new loci involved in the host susceptibility to Salmonella Typhimurium in collaborative cross mice. BMC Genom. 19, 303. https://doi.org/10.1186/s12864-018-4667-0 (2018).
Durrant, C. et al. Collaborative cross mice and their power to map host susceptibility to Aspergillus fumigatus infection. Genome Res. 21, 1239–1248 (2011).
Loomis, W. P. et al. Temporal and anatomical host resistance to chronic Salmonella infection is quantitatively dictated by Nramp1 and influenced by host genetic background. PLoS ONE 9, e111763. https://doi.org/10.1371/journal.pone.0111763 (2014).
Pandya, K. D. et al. An unmutated IgM response to the Vi polysaccharide of Salmonella Typhi contributes to protective immunity in a murine model of typhoid. J. Immunol. 200, 4078–4084. https://doi.org/10.4049/jimmunol.1701348 (2018).
Khosla, S. N. Typhoid hepatitis. Postgrad. Med. J. 66, 923–925 (1990).
Narechania, S., Duran, M., Karivedu, V. & Gopalakrishna, K. V. A case of typhoid fever with hepatic granulomas and enteritis. Case Rep. Pathol. 2015, 745461 (2015).
Martin, F. & Kearney, J. F. B1 cells: Similarities and differences with other B cell subsets. Curr. Opin. Immunol. 13, 195–201 (2001).
Marshall, J. L. et al. The capsular polysaccharide Vi from Salmonella Typhi is a B1b antigen. J. Immunol. 189, 5527–5532. https://doi.org/10.4049/jimmunol.1103166 (2012).
Hensel, J. A., Khattar, V., Ashton, R. & Ponnazhagan, S. Characterization of immune cell subtypes in three commonly used mouse strains reveals gender and strain-specific variations. Lab. Investig. 99, 93–106. https://doi.org/10.1038/s41374-018-0137-1 (2019).
Martin, M. D., Sompallae, R., Winborn, C. S., Harty, J. T. & Badovinac, V. P. Diverse CD8 T cell responses to viral infection revealed by the collaborative cross. Cell Rep. 31, 107508. https://doi.org/10.1016/j.celrep.2020.03.072 (2020).
Khan, A. Q., Lees, A. & Snapper, C. M. Differential regulation of IgG anti-capsular polysaccharide and antiprotein responses to intact Streptococcus pneumoniae in the presence of cognate CD4+ T cell help. J. Immunol. 172, 532–539 (2004).
Dickinson, G. S., Levenson, E. A., Walker, J. A., Kearney, J. F. & Alugupalli, K. R. IL-7 enables antibody responses to bacterial polysaccharides by promoting B cell receptor diversity. J. Immunol. 201, 1229–1240. https://doi.org/10.4049/jimmunol.1800162 (2018).
Sabbagh, S. C., Forest, C. G., Lepage, C., Leclerc, J. M. & Daigle, F. So similar, yet so different: Uncovering distinctive features in the genomes of Salmonella enterica serovars Typhimurium and Typhi. FEMS Microbiol. Lett. 305, 1–13. https://doi.org/10.1111/j.1574-6968.2010.01904.x (2010).
Nuccio, S. P. & Baumler, A. J. Comparative analysis of Salmonella genomes identifies a metabolic network for escalating growth in the inflamed gut. MBio 5, e00929-00914. https://doi.org/10.1128/mBio.00929-14 (2014).
Haneda, T. et al. The capsule-encoding viaB locus reduces intestinal inflammation by a Salmonella pathogenicity island 1-independent mechanism. Infect. Immun. 77, 2932–2942. https://doi.org/10.1128/IAI.00172-09 (2009).
Wilson, R. P. et al. The Vi capsular polysaccharide prevents complement receptor 3-mediated clearance of Salmonella enterica serotype Typhi. Infect. Immun. 79, 830–837 (2011).
Wilson, R. P. et al. The Vi-capsule prevents Toll-like receptor 4 recognition of Salmonella. Cell Microbiol. 10, 876–890 (2008).
Raffatellu, M. et al. The Vi capsular antigen of Salmonella enterica serotype Typhi reduces Toll-like receptor-dependent interleukin-8 expression in the intestinal mucosa. Infect. Immun. 73, 3367–3374. https://doi.org/10.1128/IAI.73.6.3367-3374.2005 (2005).
Hiyoshi, H. et al. Mechanisms to evade the phagocyte respiratory burst arose by convergent evolution in typhoidal Salmonella Serovars. Cell Rep. 22, 1787–1797. https://doi.org/10.1016/j.celrep.2018.01.016 (2018).
Wangdi, T. et al. The Vi capsular polysaccharide enables Salmonella enterica serovar typhi to evade microbe-guided neutrophil chemotaxis. PLoS Pathog 10, e1004306 (2014).
Belde, V. et al. Terminal deoxynucleotidyl transferase is not required for antibody response to polysaccharide vaccines against Streptococcus pneumoniae and Salmonella enterica Serovar Typhi. Infect. Immun. https://doi.org/10.1128/IAI.00211-18 (2018).
Deng, L. et al. Host adaptation of a bacterial toxin from the human pathogen Salmonella Typhi. Cell 159, 1290–1299. https://doi.org/10.1016/j.cell.2014.10.057 (2014).
Song, J., Gao, X. & Galan, J. E. Structure and function of the Salmonella Typhi chimaeric A2B5 typhoid toxin. Nature 499, 350–354. https://doi.org/10.1038/nature12377 (2013).
Karlinsey, J. E. et al. Genome-wide analysis of Salmonella enterica serovar Typhi in humanized mice reveals key virulence features. Cell Host Microbe 26, 426–434. https://doi.org/10.1016/j.chom.2019.08.001 (2019).
Gibani, M. M. et al. Investigation of the role of typhoid toxin in acute typhoid fever in a human challenge model. Nat. Med. 25, 1082–1088. https://doi.org/10.1038/s41591-019-0505-4 (2019).
Poltorak, A. et al. Defective LPS signaling in C3H/HeJ and C57BL/10ScCr mice: Mutations in Tlr4 gene. Science 282, 2085–2088 (1998).
O’Brien, A. D., Rosenstreich, D. L. & Taylor, B. A. Control of natural resistance to Salmonella typhimurium and Leishmania donovani in mice by closely linked but distinct genetic loci. Nature 287, 440–442 (1980).
Khan, R. T., Chevenon, M., Yuki, K. E. & Malo, D. Genetic dissection of the Ity3 locus identifies a role for Ncf2 co-expression modules and suggests Selp as a candidate gene underlying the Ity3.2 locus. Front. Immunol. 5, 375. https://doi.org/10.3389/fimmu.2014.00375 (2014).
Scoggin, K. et al. Elucidating mechanisms of tolerance to Salmonella typhimurium across long-term infections using the collaborative cross. MBio https://doi.org/10.1128/mbio.01120-22 (2022).
Morgan, A. P. et al. The mouse universal genotyping array: From substrains to subspecies. G3 (Bethesda) 6, 263–279 (2015).
Sigmon, J. S. et al. Content and performance of the MiniMUGA genotyping array: A new tool to improve rigor and reproducibility in mouse research. Genetics 216, 905–930. https://doi.org/10.1534/genetics.120.303596 (2020).
Dickinson, G. S., Levenson, E. A., Walker, J. A., Kearney, J. F. & Alugupalli, K. R. IL-7 enables antibody responses to bacterial polysaccharides by promoting B cell receptor diversity. J. Immunol. https://doi.org/10.4049/jimmunol.1800162 (2018).
Kothari, S. et al. A novel method for purification of Vi capsular polysaccharide produced by Salmonella enterica subspecies enterica serovar Typhi. Vaccine 31, 4714–4719. https://doi.org/10.1016/j.vaccine.2013.08.037 (2013).
Acknowledgements
We thank Dr. John Cipollo for providing Vi Polysaccharide. This work was supported by Sidney Kimmel Cancer Center’s NCI core grant P30 CA056036, TJU Dean’s Innovation Award, NIH grant AI159798 to KRA, and AI044170 to AJB.
Author information
Authors and Affiliations
Contributions
K.R.A. conceived the hypothesis designed and conducted the experiments, analyzed the data, and wrote the manuscript. L.S. conceived the genetic approach and bred the CC mice used in this study. F.P.M.V. and D.R.M. selected the CC strains and provided the expertise for the CC project. A.J.B. and T.W. provided S. Typhi and S. Typhimurium strains, expertise on the Salmonella infection systems, and discussion. L.A.G. performed the histopathological analysis. S.K. purified the ViPS for ELISA. M.P.C., J.A.W., D.T.D., and G.S.D. provided technical assistance in performing the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alugupalli, K.R., Kothari, S., Cravens, M.P. et al. Identification of collaborative cross mouse strains permissive to Salmonella enterica serovar Typhi infection. Sci Rep 13, 393 (2023). https://doi.org/10.1038/s41598-023-27400-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-27400-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.