Abstract
Understanding peripheral nerve micro-anatomy can assist in the development of safe and effective neuromodulation devices. However, current approaches for imaging nerve morphology at the fiber level are either cumbersome, require substantial instrumentation, have a limited volume of view, or are limited in resolution/contrast. We present alternative methods based on MUSE (Microscopy with Ultraviolet Surface Excitation) imaging to investigate peripheral nerve morphology, both in 2D and 3D. For 2D imaging, fixed samples are imaged on a conventional MUSE system either label free (via auto-fluorescence) or after staining with fluorescent dyes. This method provides a simple and rapid technique to visualize myelinated nerve fibers at specific locations along the length of the nerve and perform measurements of fiber morphology (e.g., axon diameter and g-ratio). For 3D imaging, a whole-mount staining and MUSE block-face imaging method is developed that can be used to characterize peripheral nerve micro-anatomy and improve the accuracy of computational models in neuromodulation. Images of rat sciatic and human cadaver tibial nerves are presented, illustrating the applicability of the method in different preclinical models.
Similar content being viewed by others
Introduction
Peripheral nerves act as bidirectional pathways, forming communication networks between the central nervous system and other parts of the body. Owing to their key role in the nervous system and relative ease of accessibility, electrical stimulation of these nerves has shown great promise in treating a wide range of neurological conditions and injuries1,2. However, current understanding of peripheral nerve micro-anatomy is limited, resulting in design challenges of neural interfaces. For instance, anatomical features of the nerve (e.g., fascicle diameter, perineural sheath thickness, fiber arrangement within fascicles, myelin sheath thickness, and proximity to extraneural electrodes) are known to significantly influence a fiber’s response to stimulation3,4,5. As researchers continue to improve neural interface designs, it is also important to map nerve micro-anatomy to achieve the desired stimulation patterns and maximize therapeutic efficacy. This manuscript builds on a novel imaging tool to address this problem.
Currently available imaging methods are limited in their ability to image peripheral nerve micro-anatomy across long lengths. The gold standard method of resin-based histology enables automated analysis of nerve morphology on thin sections 6; however, methods to reconstruct a 3D volume from serial sections have proven to be challenging7. Imaging with micro-CT can provide volumetric datasets across longer distances but resolution and contrast are limited to visualizing fascicle bundles8. Recently, diffusion MRI methods have been applied to characterize fiber tracts within peripheral nerves. Although this method has the benefit of imaging in vivo, imaging at a high resolution results in a low signal-to-noise ratio (SNR) and long imaging times9. In the optical microscopy domain, label-free methods such as Coherent anti-Stokes Raman scattering microscopy utilize physical properties of lipid and myelin sheaths to generate contrast10; however, such systems require expensive laser sources and are limited in imaging depth. Developments in light-sheet microscopy can be utilized to image large samples, but these methods require the use of tissue clearing reagents which can distort the tissue, making it not well suited for morphometric analyses11. In this regard, a comprehensive validation of the light-sheet approach in human cadaver nerve tissues is required. Additionally, the working distance of the objective limits the achievable depth, which can be a concern when imaging large human nerves. Thus, developing an imaging tool that is easy to implement, has sufficient resolution and contrast to visualize nerve micro-anatomy over a large field of view can address a critical need in peripheral nerve imaging.
We pursue a slide-free, technically simple, and cost-effective microscopy technique, Microscopy with Ultraviolet Surface Excitation (MUSE)12,13 to image nerve micro-anatomy. Briefly, MUSE imaging uses ultraviolet (UV) excitation at 280 nm and captures fluorescence images of the tissue surface using a color camera. The MUSE system provides several advantages. First, the limited tissue penetration depth of UV light at 280 nm provides optical sectioning by confining fluorescence excitation to the tissue surface, reducing subsurface fluorescence that can degrade effective resolution and contrast. Second, a wide range of commonly used MUSE dyes are fluorescent in the visible range, allowing image acquisition with a color camera. Third, UV light in the appropriate spectral ranges can be generated with inexpensive LED sources. As UV light below 300 nm is absorbed by glass, the microscope optics intrinsically block stray excitation light, thus eliminating the need for specific excitation and emission filter sets. Finally, the process of creating images of the tissue surface with this technique is almost instantaneous, providing a significant time savings compared to conventional histology. Previous reports have utilized this method for imaging surgical specimens of breast and skin cancer using bench-top systems14,15 and in the implementation of a smartphone-based portable fluorescence microscope16. Depending on which structures are being visualized, tissue auto-fluorescence may provide sufficient contrast with suitable illumination patterns17; in other cases, the addition of fluorescent dyes or labeling agents may be necessary.
In order to extend MUSE to image a 3D volume, we utilize a block-face imaging approach. A number of block-face imaging techniques have been previously reported18,19,20,21,22,23,24,25, but are often limited in resolution due to subsurface fluorescence. Some approaches have applied deconvolution-based techniques18,26 to remove out-of-plane blurring; these methods can have limited success and prove to be computationally intensive when applied to large datasets. Alternatively, few approaches have used confocal19 or multi-photon20 imaging to achieve optical sectioning on the block face, but these systems are complex and costly. To address this issue, we build on earlier reports27 and utilize the advantages of MUSE microscopy described previously for imaging the block face. A resin-based embedding medium is used to create tissue blocks since it is known to preserve morphology better than paraffin28.
In this report, we will present methods to image peripheral nerve samples with MUSE. We determined tissue preparation methods utilizing either fixative auto-fluorescence or stain with suitable dyes. We found that resolution and field of view were sufficient to image micro-anatomical features of interest (e.g., fascicles, connective sheaths, and individual nerve fibers) in rat and human cadaver samples. We utilized both 2D and 3D MUSE approaches to determine their characteristics for nerve imaging. We are specifically interested in the capability to use 3D MUSE over extended lengths of nerve tissue.
Results
2D-MUSE imaging
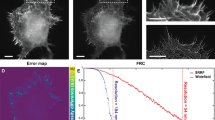
In Fig. 1, we demonstrate 2D-MUSE imaging of a rat sciatic nerve sample, utilizing fixative autofluorescence. Tissue preparation steps included formalin fixation, sectioning with a razor blade, and gentle pressing of the cut tissue onto the quartz slide of the inverted MUSE microscope. When imaging the nerve at this level, we identify three fascicles, separated by thin connective sheaths (perineurium). Image resolution and contrast were found to be sufficient to delineate axons and myelin sheaths of individual nerve fibers. Under the assumption that the space inside the myelin sheath is completely filled with the axon, individual g-ratios can be extracted from these images29. The g-ratio relates to the conduction velocity in nerve fibers and is calculated as the ratio of axon diameter to the total fiber diameter. By performing manual segmentation on a group of randomly selected nerve fibers (Fig. 1d), we quantify both axon diameters (2.78 + /−0.41 µm, n = 27), and individual g-ratios (0.55 + /−0.06, n = 27, Fig. 1e).
2D-MUSE image of a rat sciatic nerve. Fixed nerve tissue was sectioned on a compresstome and laid immediately on a quartz slide for imaging. Individual myelinated fibers are visible (b and c) with contrast originating from tissue autofluorescence under UV illumination. (d) Manual segmentation on a subset of myelinated fibers in (b) allows quantification of axon diameters and corresponding g-ratios, displayed as histograms in (e). Scale bars are: (a) 100 µm, (b and c) 50 µm. Close-up views in (b) and (c) were contrast enhanced to improve visibility of myelin sheaths.
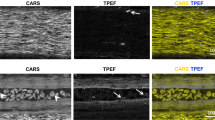
Staining with fluorescent dyes further improved image contrast (Fig. 2). Common MUSE dyes such as Hoechst and rhodamine generated contrast between additional tissue structures, including cell nuclei and vasculature. When imaging the nerve in the longitudinal direction, individual fibers can be clearly delineated from each other across the nerve. Vascular-like structures are evident from Hoechst staining in a close-up view (Fig. 2e). For certain fibers, a gap in myelinated regions is observed, suggestive of Schmidt-Lanterman clefts or nodes of Ranvier. We also tested staining with a fluorescent compound that selectively binds to myelin sheaths (Case Imaging Compound30, a stilbenzene derivative). The compound was found to be fluorescent under MUSE illumination, allowing visualization of myelin sheaths on a human tibial nerve (Fig. 3). Note that even with staining, 2D-MUSE imaging is quite fast, with sample preparation and imaging completed in under five minutes.
2D-MUSE image of a rat sciatic nerve with staining. Sample was sectioned by hand with a razor blade, immersed in a rhodamine and Hoechst staining solution for 30 s, rinsed in DI water and imaged. (a) Transverse cross-section illustrates several myelinated fibers grouped as a single fascicle and surrounded by a thin perineural sheath. (b and c) Close up views from (a) show multiple myelinated fibers as yellow rings. (d) Longitudinal cross section allows visualizing fibers along the length of the nerve. (e) Zoomed in region from (d) illustrates microvasculature revealed by Hoechst staining and evident from tubular structure (arrows). (f) A gap in the myelinated fibers is visible in this close-up view, suggestive of Schimdt-Lanterman clefts or node of Ranvier (asterisk). Scale bars are: 100 µm (a, d) and 20 µm (b, c, e, f).
2D-MUSE image of a cadaver tibial nerve sample, stained with a myelin-specific fluorescent compound (Case Imaging Compound30). Images were color and intensity inverted to improve visual saliency. (a) MUSE image, color and intensity inverted to improve visual saliency. (b, c) Close up views of regions highlighted by white boxes in (a) displaying dark rings, indicative of myelinated fibers. Note the low image contrast from vasculature with this stain (arrow in a). Scale bars are 100 µm in (a) and 50 µm in (b and c).
Block-face 3D-MUSE imaging
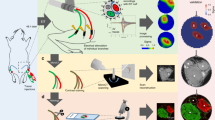
Since 2D-MUSE generated promising images of micro-anatomy in nerve cross-sections, we proceeded to use the block-face, 3D-MUSE imaging system to further reveal spatial organization of peripheral nerves. We used osmium tetroxide and rhodamine B to whole mount stain the nerve, giving contrast to fascicles, connective tissue, myelinated fibers, vasculature, and adipocytes in a human tibial nerve. Individual fascicles along with inner and outer perineural boundaries are visible in the block-face images (Fig. 4a-b). Nerve fibers with diameter greater than 5 µm could be clearly resolved with the current imaging system. An example of a fiber with an axon diameter around 5 µm is illustrated in Fig. 4c. When staining with osmium tetroxide, unsaturated fats in adipocytes react in the same manner as myelin sheaths and therefore, also appear as dark regions in the block-face images. Using a manual segmentation of the inner and outer boundaries of the perineural sheaths, fascicle diameters and perineural sheath thickness could be quantified in the block-face image shown in Fig. 4a. Fascicle diameter was found to be 423 + /− 143 µm and the sheath thickness was 25 + /−11 µm (n = 25 fascicles, Fig. S1). Figure 4d demonstrates a sub-volume with fibers and vasculature within a single nerve fascicle. With manual segmentation, we identify vascular networks within the fascicle (Fig. 4e). Since the volume was sectioned at 5 µm thickness, variations in the shape of structures from one slice to the next were low, allowing determination of structure continuity. Such vascular networks supply nutrients to regions distant from the nerve cell body. When the sub-volume was visualized from the side (XZ plane, longitudinal view), we identify multiple myelinated axons (Supplementary Movie 1). Upon stepping through individual slices in this view, we identify that the full length of the axons is not visible in a single slice but extends over multiple slices. This indicates that axon morphology is not perfectly described by a cylinder and necessitates a more complex 3D description. Volumetric imaging revealed a fascicle merging event (Supplementary Movie 2). The two fascicles are initially seen close to one another and are separated by a perineural sheath. As we move deeper into the nerve, we identify that the sheath gets progressively thinner, until there is merging.
Imaging a cadaver tibial nerve sample with block-face 3D-MUSE imaging. The nerve was post-fixed with osmium tetroxide and counterstained with rhodamine B. Images were acquired with a 10 × 0.45 NA objective and the sample was sectioned at 5 µm thickness. Further details of the system are provided in “Methods”. (a) Representative block face image stitched from twelve individual tiles. Structures of interest include fascicles, venuole (asterisk), and adipose cells (plus). (b and c) Close up views indicate individual fascicles and myelinated fibers respectively. (d) Volume rendering of a region within a fascicle, outlined by a brown box in (a). (e) Manual segmentation of intraneural vasculature from the same volume as in (d). Box size: 800 × 600 µm2. Scale bars are (a) 1 mm (d and e) 250 µm.
Discussion
With 2D-MUSE imaging, we identified several key structures in nerve micro-anatomy including myelinated fibers, blood vessels and connective tissue either with auto-fluorescence or by the addition of suitable fluorescent stains. Measurement of axon diameters and g-ratios was possible. The results are similar to previously published values in literature31 (mean axon diameter: 2.78 µm vs. 2.28 µm, mean g-ratio: 0.55 vs. 0.52). The minor differences are potentially due to varying tissue expansion or shrinkage during sample preparation. However, since a small group of fibers were randomly selected for our measurements, further tests were not performed due to limited statistical power. Rhodamine B staining improved the visibility of myelin, presumably due to a higher solubility in lipophilic media32. The longitudinal cross-section of the nerve showed anatomical structures that are potentially blood vessels and Schmidt-Lanterman clefts and/or nodes of Ranvier, however, further validation is necessary. When staining with the Case Imaging Compound, fluorescence was restricted to the myelinated fibers alone, which could be useful when studying disorders related to the myelin sheath (e.g., multiple sclerosis) and/or assess therapeutic response.
We developed a whole-mount staining approach for visualizing nerve micro-anatomy with 3D-MUSE. Osmium tetroxide preserved lipids/myelin during the dehydration process and produced dark staining for myelinated fibers across depth33. An additional advantage of this stain is its role as a contrast agent in micro-CT34, suggesting suitability in a multimodality imaging pipeline. Combined with rhodamine B counterstaining, the system provided sufficient resolution and contrast to visualize various structures of interest across depth. For example, nerve fascicles were clearly visible and individual fascicle diameters could be quantified. Volumetric imaging also allowed identification of fascicle merging events. Additionally, inner and outer perineural boundaries can be identified, allowing us to quantify sheath thickness. Since these anatomical parameters (fascicle diameter, sheath thickness, fiber distance from the electrode) significantly affect stimulation response, their measurement can guide computational modeling studies for neuromodulation. With suitable segmentation tools, these datasets can provide an accurate representation of nerve micro-anatomy, making them suitable for modeling studies assessing various nerve interface designs. Visualizing nerve fibers across length illustrated the complex 3D morphology of individual axons, consistent with findings from previous reports utilizing electron microscopy35. The combination of osmium tetroxide staining and GMA embedding also produces final tissue blocks with lesser shrinkage compared to paraffin36, thereby making the technique more suitable for morphometric analyses. During our embedding experiments, we noticed that the final blocks occasionally contained air bubbles, potentially due to leftover osmium in the tissue interfering with GMA polymerization37. We aim to address this limitation in future work by testing optimized protocols prescribed in the literature38.
We demonstrated 3D-MUSE block-face imaging and identified advantages of using this system to image peripheral nerves. First, by performing serial section and imaging, long samples can be imaged in a straightforward manner. With the section-and-image approach, the length of nerve imaged could be virtually unlimited. Our current prototype is limited to about 20 mm. Second, since MUSE utilizes simple instrumentation12, the system does not require significant technical expertise to develop and maintain. Third, with a tiling approach, a large field of view (up to 20 × 20 mm with our current motorized stages) can be covered. Finally, with our whole-mount staining procedure described above, we generate images of nerve micro-anatomy with improved resolution and contrast as compared to micro-CT and MRI techniques. For instance, micro-CT imaging provides limited contrast to visualize the perineural sheaths around the fascicles. Diffusion MRI provides maps of fiber tracts but is limited in resolution and contrast to visualize individual fibers. We identify that with our 3D-MUSE approach, both perineural sheaths and large nerve fibers are visible.
Overall, our work describes a microscopic imaging method based on MUSE imaging to visualize nerve micro-anatomy in 2D and 3D. The 2D imaging system offers a simple method to image myelinated fibers at specific locations in the nerve ex vivo, without the need for laborious embedding protocols or complex instrumentation. Extending MUSE to 3D via block-face imaging offers a simple, relatively inexpensive method to analyze these samples when a continuous 3D representation of nerve anatomy is required.
Methods
Tissue preparation
Short segments of sciatic nerves were excised from discarded rat tissues from unrelated studies with IACUC approval at Washington University in St. Louis. Results presented in this work are in accordance with relevant guidelines and regulations (ARRIVE guidelines39). Nerves were fixed in 4% PFA overnight and were subsequently rinsed and stored in 1x PBS. Samples were either cut by hand with a razor blade or were embedded in agarose and sectioned on a Compresstome® (Precisionary, MA) to create a flat surface. Human tibial nerve samples were extracted from embalmed cadavers after their use in medical school cadaver lab training. The study was conducted according to guidelines and regulations laid down in the Declaration of Helsinki. Informed consent was provided by the donors prior to their deaths or by their next of kin. Our study was reviewed by the Case Western Reserve University Institutional Review Board and was determined as non-human subject research based on the nature of the de-identified cadaver donations and other study parameters. Samples were stored in 10% neutral buffered formalin solution at 4 °C for several weeks prior to staining and embedding. Nerves were cut into < 1 mm sections with a razor blade and washed in 1x PBS overnight at 4°C to remove excess fixative.
Sample preparation for 2D-MUSE imaging
Rat sciatic nerve samples were immersed in a solution of 0.5 mg/mL rhodamine B (Sigma) and 0.1 mg/mL Hoechst (Sigma) in 1x PBS for approximately 30 seconds, followed by rinsing under DI water for 10 seconds. Human tibial nerve samples were stained with a myelin-specific fluorescent compound (Case Imaging Compound30). The compound was prepared and stored at a concentration of 10 mM in DMSO. Prior to imaging, the staining solution was made by diluting to a final concentration of 1 mM with 1x PBS. Steps involved in this sample preparation protocol are illustrated in Fig. S2a.
Sample preparation for block-face 3D-MUSE imaging
Human tibial nerve samples were stained with osmium tetroxide and rhodamine B followed by embedding in glycol methacrylate (GMA) resin. The samples were transferred to 2% osmium tetroxide (Electron Microscopy Sciences) and placed on a rotator for 2 hours, forming a black, insoluble precipitate upon reaction with unsaturated fatty acids within the tissue33. The sample was dehydrated in a series of ethanol solutions (30, 60, 90 and 100%, 2h each). Rhodamine B was added at a concentration of 0.5% (w/v) to the ethanol solutions to create a counter contrast. The dehydrated sample was embedded in the GMA resin as previously described40. The samples were placed in a 50:50 mix of ethanol and GMA monomer (Electron Microscopy Sciences) overnight at 4 °C, followed by a freshly prepared GMA infiltration solution (GMA monomer with 1% (w/v) of the initiator added) for 24 h at 4 °C. Samples were then polymerized in 1:15 mix of accelerator and infiltration solution in Teflon molds under vacuum overnight at 20–25 °C. Hardened sample blocks were attached to plastic holders using a fast-curing methyl methacrylate resin (Electron Microscopy Sciences) for microtome sectioning. A detailed protocol is illustrated in Fig. S2b.
2D-MUSE imaging
The setup for 2D-MUSE imaging consists of a UV transparent slide (commonly quartz), 280 nm UV LED for illumination and an inverted microscope system. After the samples were prepared as described above, they were placed on the slide with the surface to be imaged facing downward. The slide is attached to a motorized XYZ stage to provide automated panning and focusing of the sample surface. Since UV light at 280 nm is absorbed by glass, the sample surface is excited with oblique illumination. This is in contrast to routine epifluorescence microscopes which use light in the visible wavelength for excitation with coaxial illumination. The microscope system included a 10x/0.45 NA objective (Nikon Plan Apo), an infinity corrected tube lens (ITL200, Thorlabs) and a CCD color camera (Ximea, MD091CU-SY). Custom control software integrating the various hardware components was developed in the. NET framework. The imaging protocol consists of focusing the microscope at a specific region of interest and capturing an image. Optionally, tiling can be performed to increase the effective field of view. Further details about the imaging system are available in a previous publication by Fereidouni et al.12
Block-face 3D-MUSE imaging
The setup for block-face 3D-MUSE imaging system is based on an automated microtome and a MUSE imaging system. Serial sectioning was performed at 5 µm section thickness with a motorized rotary microtome (Microm HM355 S) equipped with a 16 cm D profile tungsten carbide knife. The GMA-embedded sample was trimmed repeatedly with the microtome. At the end of each cutting cycle, the microtome positions the sample carrier at its upper reversal point (away from the knife) making it convenient to perform serial imaging. An Arduino microcontroller and relay switches were used as the interface between the computer and the microtome. The MUSE imaging system was similar to the original MUSE setup (Fig. S3a)12. For illumination, we use a UV LED with a center wavelength of 285 nm (Thorlabs). Images were collected using a 10x/0.45 objective (Nikon Plan Apo) and a CMOS color camera (Teledyne FLIR, 12 megapixels), providing a field of view of about 1.4 x 1.1 mm. The imaging system is mounted on 3-axis motorized stages (Zaber) to facilitate focusing and tiling. Micro-Manager41, an open-source microscope control software, was used to control serial sample sectioning and image acquisition. Image tiles were set up with an overlap of ten percent to ensure stitching was free of artifacts. A resolution test image of the USAF 1951 target taken with this system is provided in Fig. S3b, indicating resolvability up to Group 9 Element 2 (0.87 µm line spacing).
Image post-processing and visualization
Raw MUSE data were post-processed with the procedure described previously12. Briefly, images were sharpened using unsharp masking in MATLAB. Image contrast was enhanced either with the automatic white balance tool or the curve tool in GIMP. Tile scans were stitched using Image Composite Editor software (Microsoft).
Segmentation and quantification methods
Manual segmentation tools were used to create labeled images for subsequent quantitative analyses. Structures were identified by visually comparing our results with the gold standard of resin embedding and toluidine blue staining42. In order to compute fiber diameters and g-ratios (Fig. 1d and e), large fibers were randomly selected and the inner and outer boundaries of myelin sheaths were delineated using the free hand segmentation tool in Amira software (Thermo Fisher Scientific). A subset of myelinated fibers with diameters ≥ 4 µm could be identified and labeled with this approach. Smaller fibers were difficult to accurately label due to limitations in image resolution. The labeled regions were subsequently analyzed by computing the diameter of a circle with the same area as the region. This was done using the regionprops command in MATLAB. Since the command returns the diameter in terms of pixels, values were multiplied by the calibrated pixel size to obtain the result in physical units (µm).
A similar approach was taken to quantify fascicle diameters and perineural sheath thickness. The inner and outer boundaries of the perineural sheath were delineated to create a labeled image (shown overlaid against the raw image in Fig. S1). We then computed equivalent fascicle diameters. Additionally, perineural sheath thickness was determined by measuring the thickness at 100 points along its boundary using MATLAB code developed in-house. These points were created by drawing rays from the fascicle centroid at equiangular spacing, thereby covering a full 360 degree rotation. This process was repeated for each of the 25 fascicles in Fig. 4a. All values presented in this work indicate mean + /−SD.
Ethical approval
All animal procedures and protocols were performed in accordance with relevant guidelines and regulations (NIH Guide for the Care and Use of Laboratory Animals).
References
Larson, C. & Meng, E. A review for the peripheral nerve interface designer. J. Neurosci. Methods 332, 108523 (2020).
Tyler, D. & Polasek, K. Chapter 17 - electrodes for the neural interface. https://doi.org/10.1016/B978-0-12-374248-3.00018-5. Academic Press, 2009).
Grill, W. & Mortimer, J. Stimulus waveforms for selective neural stimulation. IEEE Eng. Med. Biol. Mag 14, 375–385 (1995).
Grinberg, Y., Schiefer, M. A., Tyler, D. J. & Gustafson, K. J. Fascicular perineurium thickness, size, and position affect model predictions of neural excitation. IEEE Trans. Neural Syst. Rehabil. Eng. 16, 572–581 (2008).
Pelot, N., Behrend, C. & Grill, W. On the parameters used in finite element modeling of compound peripheral nerves. J. Neural Eng 16, 016007 (2019).
Jortner, B. Preparation and analysis of the peripheral nervous system. Toxicol. Pathol. 39, 66–72 (2011).
Pichat, J., Iglesias, J. E., Yousry, T., Ourselin, S. & Modat, M. A survey of methods for 3d histology reconstruction. Med. Image Anal. 46, 73–105 (2018).
Thompson, N. Microct optimisation for imaging fascicular anatomy in peripheral nerves. J. Neurosci. Methods 338, 108652 (2020).
Jeon, T., Fung, M. M., Koch, K. M., Tan, E. T. & Sneag, D. B. Peripheral nerve diffusion tensor imaging: Overview, pitfalls, and future directions. J. Magn. Reson. Imaging 47, 1171–1189 (2018).
Li, S., Li, Y., Yi, R., Liu, L. & Qu, J. Coherent anti-stokes raman scattering microscopy and its applications. Front. Phys. 8, 515 (2020).
Jung, Y. et al. Comprehensive evaluation of peripheral nerve regeneration in the acute healing phase using tissue clearing and optical microscopy in a rodent model. PLoS ONE 9, e94054 (2014).
Fereidouni, F. Microscopy with ultraviolet surface excitation for rapid slide-free histology. Nat. Biomed. Eng 1, 957 (2017).
Fereidouni, F., Mitra, A., Demos, S. & Levenson, R. Microscopy with uv surface excitation (muse) for slide-free histology and pathology imaging. In Optical Biopsy XIII: Toward Real-Time Spectroscopic Imaging and Diagnosis, vol. 9318 93180F (International Society for Optics and Photonics, 2015).
Yoshitake, T. Rapid histopathological imaging of skin and breast cancer surgical specimens using immersion microscopy with ultraviolet surface excitation. Sci. Rep 8, 4476 (2018).
Xie, W. Microscopy with ultraviolet surface excitation for wide-area pathology of breast surgical margins. J. Biomed. Opt 24, 026501 (2019).
Liu, Y., Rollins, A., Levenson, R., Fereidouni, F. & Jenkins, M. Pocket muse: An affordable, versatile and high-performance fluorescence microscope using a smartphone. Commun. Biol. 4, 1–14 (2021).
Zhang, Y. et al. High-throughput, label-free and slide-free histological imaging by computational microscopy and unsupervised learning. Adv. Sci. 9(2), 2102358 (2021).
Walsh, C., Holroyd, N.A., Finnerty, E., Ryan, S.G., Sweeney, P.W., Shipley, R.J., Walker-Samuel, S. Multifluorescence High-Resolution Episcopic Microscopy (MF-HREM) for Three- Dimensional Imaging of Adult Murine Organs. Adv. Photonics Res2(10), 2100110 (2021).
Seiriki, K. Whole-brain block-face serial microscopy tomography at subcellular resolution using fast. Nat. Protoc 14, 1509–1529 (2019).
Ragan, T. Serial two-photon tomography for automated ex vivo mouse brain imaging. Nat. Methods 9, 255–258 (2012).
Roy, D., Steyer, G., Gargesha, M., Stone, M. & Wilson, D. 3d cryo-imaging: A very high-resolution view of the whole mouse. Anat. Rec. Hoboken NJ 292, 342–351 (2007).
Weninger, W. High-resolution episcopic microscopy: A rapid technique for high detailed 3d analysis of gene activity in the context of tissue architecture and morphology. Anat. Embryol Berl. 211, 213–221 (2006).
Mayerich, D., Abbott, L. & McCormick, B. Knife-edge scanning microscopy for imaging and reconstruction of three- dimensional anatomical structures of the mouse brain. J. Microsc. 231, 134–143 (2008).
Xu, F. High-throughput mapping of a whole rhesus monkey brain at micrometer resolution. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-00986-5 (2021).
Wong, T. T. et al. Label-free automated three-dimensional imaging of whole organs by microtomy-assisted photoacoustic microscopy. Nat. Commun. 8, 1–8 (2017).
Krishnamurthi, G., Wang, C., Steyer, G. & Wilson, D. Removal of subsurface fluorescence in cryo-imaging using deconvolution. Opt. Expr. 18, 22324–22338 (2010).
Guo, J., Artur, C., Eriksen, J. & Mayerich, D. Three-dimensional microscopy by milling with ultraviolet excitation. Sci. Rep. 9, 14578 (2019).
Wittenburg, G., Volkel, C., Mai, R. & Lauer, G. Immunohistochemical comparison of differentiation markers on paraffin and plastic embedded human bone samples. J. Physiol. Pharmacol. Off. J. Pol. Physiol. Soc. 60(8), 43–49 (2009).
Rushton, W. A theory of the effects of fibre size in medullated nerve. The J. Physiol. 115, 101–122 (1951).
Wang, Y. In vivo quantification of myelin changes in the vertebrate nervous system. J. Neurosci 29, 14663–14669 (2009).
Kim, P. Collagen nerve protector in rat sciatic nerve repair: A morphometric and histological analysis. Microsurgery 30, 392–396 (2010).
Kasnavia, T., Vu, D. & Sabatini, D. A. Fluorescent dye and media properties affecting sorption and tracer selection. Groundwater 37, 376–381 (1999).
Kiernan, J. Histochemistry of staining methods for normal and degenerating myelin in the central and peripheral nervous systems. J. Histotechnol 30, 87–106. https://doi.org/10.1179/his.2007.30.2.87 (2007).
Masis, J. et al. A micro-ct-based method for characterizing lesions and locating electrodes in small animal brains. JoVE (Journal Vis. Exp. e58585 (2018).
Andersson, M. et al. Axon morphology is modulated by the local environment and impacts the noninvasive investigation of its structure–function relationship. Proc. Natl. Acad. Sci. 117, 33649–33659 (2020).
Schneider, J. P. & Ochs, M. Alterations of mouse lung tissue dimensions during processing for morphometry: a comparison of methods. Am. J. Physiol. Cell. Mol. Physiol. 306, L341–L350 (2014).
Gerrits, P. Staining myelin and myelin-like degradation products in the spinal cords of chronic experimental allergic encephalomyelitis (cr-eae) rats using sudan black b staining of glycol methacrylate-embedded material. J. Neurosci. Methods 45, 99–105 (1992).
Taksir, T. V. et al. Optimized preservation of cns morphology for the identification of glycogen in the pompe mouse model. J. Histochem. & Cytochem. 55, 991–998 (2007).
Edelstein, A. Advanced methods of microscope control using µmanager software. J. Biol. Methods 1, 10–10 (2014).
Percie du Sert, N. et al. The arrive guidelines 2.0: Updated guidelines for reporting animal research. J. Cereb. Blood Flow & Metab. 40, 1769–1777 (2020).
Yeung, E. & Chan, C. Glycol methacrylate: The art of embedding and serial sectioning. Botany 93, 1–8 (2015).
Ghnenis, A. B., Czaikowski, R. E., Zhang, Z. J. & Bushman, J. S. Toluidine blue staining of resin-embedded sections for evaluation of peripheral nerve morphology. J. Vis. Exp. 137, 58031 https://doi.org/10.3791/58031 (2018).
Acknowledgements
The authors would like to acknowledge Dr. Chunying Wu and Dr. Junqing Zhu at the Department of Radiology at Case Western Reserve University for providing the myelin-specific fluorescent stain for our experiments. This research was supported by the National Institutes of Health (NIH)/National Institute of Biomedical Imaging and Bioengineering (NIBIB) under award number R01EB028635, National Cancer Institute under award number R01CA208623, and NIH SPARC 1OT2OD025340-01. The views expressed in this article are those of the authors and do not necessarily reflect the position or policy of the National Institutes of Health, Department of Veterans Affairs, or the United States government.
Author information
Authors and Affiliations
Contributions
C.K. developed the whole-mount nerve staining protocol, built the 3D-MUSE imaging system and conducted the 3D-MUSE experiments. A.T. performed the 2D-MUSE imaging experiments with the rat sciatic nerve sample. A.U. and M.Y.B. provided the cadaver nerve and rat nerve samples respectively. Y.L. provided feedback on the design of the block-face 3D-MUSE imaging system. D.L.W., M.W.J., A.J.S., R.M.L. and F.F. supervised the work and provided crucial guidance. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
R.M.L. is listed as the inventor of two patents related to the MUSE technology described in this manuscript. R.M.L. and F.F. are co-founders of HistoliX, Inc. All remaining authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Video 1.
Supplementary Video 2.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kolluru, C., Todd, A., Upadhye, A.R. et al. Imaging peripheral nerve micro-anatomy with MUSE, 2D and 3D approaches. Sci Rep 12, 10205 (2022). https://doi.org/10.1038/s41598-022-14166-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-14166-1
This article is cited by
-
An end-to-end workflow for nondestructive 3D pathology
Nature Protocols (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.