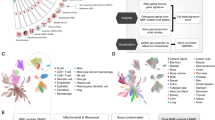
Abstract
We established a relationship among the immune-related genes, tumor-infiltrating immune cells (TIICs), and immune checkpoints in patients with osteosarcoma. The gene expression data for osteosarcoma were downloaded from UCSC Xena and GEO database. Immune-related differentially expressed genes (DEGs) were detected to calculate the risk score. “Estimate” was used for immune infiltrating estimation and “xCell” was used to obtain 64 immune cell subtypes. Furthermore, the relationship among the risk scores, immune cell subtypes, and immune checkpoints was evaluated. The three immune-related genes (TYROBP, TLR4, and ITGAM) were selected to establish a risk scoring system based on their integrated prognostic relevance. The GSEA results for the Hallmark and KEGG pathways revealed that the low-risk score group exhibited the most gene sets that were related to immune-related pathways. The risk score significantly correlated with the xCell score of macrophages, M1 macrophages, and M2 macrophages, which significantly affected the prognosis of osteosarcoma. Thus, patients with low-risk scores showed better results with the immune checkpoints inhibitor therapy. A three immune-related, gene-based risk model can regulate macrophage activation and predict the treatment outcomes the survival rate in osteosarcoma.
Similar content being viewed by others
Introduction
Osteosarcoma (OS) is a common form of high-graded primary malignant bone neoplasm in children and adolescents1,2. Statistical data reveal that incidence of OS is continuously growing by approximately 1.4% annually3. The two most commonly used clinical treatment methods for OS include systemic chemotherapy and local control surgery4. Despite several intensive efforts for a better prognosis, the high recurrence rate and early lung metastatic results in a poor prognosis poor for patients with OS5,6. Therefore, it is important to explore the carcinogenesis and therapeutics of OS.
An increasingly number of studies have demonstrated the significance of the tumor immune microenvironment in tumor progression7,8. TIICs in the tumor microenvironment (TME) influenced the tumor development and progression, which could serve as a potential marker for predicting the prognosis9. In addition, studies report that the expression of immune checkpoints such as PD-1, PD-L1, and CTLA-4 is associated with OS immune tolerance10. Immune checkpoint inhibitors, which restore the immune function of T cells and kill tumor cells, have been used to alleviate the immunosuppressive state of the TME in solid malignancies. Recently, scholars have constantly discussed the immune-related genes in TME, which may regulate TIICs and immune checkpoints. Therefore, it becomes important to enhance the knowledge about the TME, which can later help us identify novel immune therapy biomarkers for OS.
In this study, we constructed a risk model based on the TCGA cohort and validated it using the GSE21257 cohort. The three immune-related genes (TYROBP, TLR4 and ITGAM), the basis for the risk model, were strongly associated with macrophage polarization in the TME. Furthermore, the risk score was negatively correlated with the PD-1, PD-L1, and CTLA-4 immune checkpoint proteins that affect the survival rate and efficacy of the blockade of the immune checkpoint. Therefore, our risk model can predict the efficacy of immune checkpoints blockade in OS.
Results
EMT-related DEGs were mainly involved in inflammation response
Patients in the high immune and stromal score group experienced a better prognosis than those in the low immune and stromal score group (Fig. 1A,B). A total of 765 DEGs (705 up-regulated and 60 down-regulated) were identified in the high immune score group (Fig. 1C), whereas 835 DEGs (756 up-regulated and 79 down-regulated) were identified in the high stromal score group (Fig. 1D). The DEGs (302 up-regulated and 4 down-regulated) common to both the groups were selected for further analysis (Fig. 1E,F).
Immune-related DEGs identification. (A) Kaplan–Meier survival analysis for osteosarcoma patients grouped into the high or low score in immune score determined by the comparison with the median. p = 0.0049 by log-rank test. (B) Kaplan–Meier survival curve for the stromal score with p = 0.048 by log-rank test. (C) Volcano plots for DEGs were generated by comparison of the high score group vs. the low score group in the immune score. (D) Volcano plots for DEGs were generated by comparison of the high score group vs. the low score group in the stromal score. (E,F) Venn plots showing common up-regulated and down-regulated DEGs shared by immune score and stromal score.
The GO and KEGG enrichment analyses showed the involvement of DEGs in inflammatory responses, including mononuclear and lymphocyte proliferation and activation and T cell activation (Supplementary Fig. S1A,B). The complete GO and KEGG analysis results for all DEGs are shown in Supplementary File 1. In addition, DEGs, including in module 1, were mainly involved in regulating mononuclear and lymphocyte proliferation, macrophage activation, and interleukin-1/10 production (Supplementary File 2). DEGs, including in module 2, were mainly involved in the macrophage activation, cell junction disassembly, and the regulation of cell–cell adhesion (Supplementary File 3). DEGs, including in module 3 (Supplementary File 4) and module 4 (Supplementary File 5), had significant involvement in the immune response, inflammation response, and antigen processing and presentation.
A three EMT-related genes signature was established
PPI network for all DEGs included 3891 edges and 282 nodes, as shown in Supplementary Fig. S2A. Module 1 (Supplementary Fig. S2B) network consisted of 336 edges and 32 nodes; Module 2 (Supplementary Fig. S2C) network contained 367 edges and 42 nodes; Module 3 (Supplementary Fig. S2D) network consisted of 128 edges and 36 nodes; and Module 4 (Supplementary Fig. S2E) consisted of 51 edges and 28 nodes. We identified TYROBP, TLR4, TLR8, LCP2, ITGAM, LILRB2, and CD86 as hub genes (Supplementary Fig. S3A). We used univariate Cox regression to analyze 7 DEGs and screened 5 DEGs as significant prognosis factors (Table 1). Kaplan–Meier analysis screened 4 DEGs from 7 DEGs affecting OS prognosis (Supplementary Fig. S3SB–H). These four genes, sharing Cox regression analysis and Kaplan–Meier analysis, were further subjected to LASSO regression analysis to select genes included in the risk model (Fig. 2A,B). A 3 EMT-related DEGs (TYROBP, TLR4, and ITGAM) risk signature was established. Patients with continuous risk scores harbored various clinical outcomes in different groups (Fig. 2C–E). Patients in the high-risk group had lower survival rates (Fig. 2F). Moreover, the time-dependent ROC analysis showed that the area under the curve (AUC) value of the three-gene-based model was 0.669, 0.731, and 0.732 at 1, 3, and 5 years, respectively, after diagnosis in the TCGA cohort (Fig. 2G). Similar results were provided by the GSE21257 cohort, which validated the prognostic model (Fig. 3). Furthermore, Supplementary Fig. S4 illustrates that TYROBP, TLR4, and ITGAM were significantly lower in the tumor cells than in the normal bone.
Establishment of a prognostic gene signature for osteosarcoma. (A) Using 1000-fold cross-validation to the optimal penalty parameter lambda. (B) LASSO coefficient profiles of the 4 DEGs. (C) Classification of patients into different risk groups based on the median risk score. (D) Distribution of patients’ survival time and status. (E) Heatmap of expression profiles of included risk score related genes. (F) Kaplan–Meier survival curves between low- and high-risk groups. (G) ROC curves of the risk score diagnostic ability.
Validation of the prognostic gene signature for osteosarcoma using GSE21257 cohort. (A) Classification of patients into different risk groups based on the median risk score. (B) Distribution of patients’ survival time and status. (C) Heatmap of expression profiles of included risk score related genes. (D) Kaplan–Meier survival curves between low- and high-risk groups in the GSE21257 cohort. (E) ROC curves of the risk score diagnostic ability. GSEA in the Hallmarks (F) and KEGG (G) gene set between group high (n = 42) and low (n = 42) of the risk score in the TCGA cohort.
Immune-related pathways were enriched in the low-risk group
The GSEA results for Hallmark and KEGG pathways revealed that most gene sets focused on immune-related pathways. Hallmark results indicated that a majority of the genes were enriched for allograft rejection, IL6/JAK/STAT3 signaling, IL2/STAT5 signaling, and apoptosis (Fig. 3F), whereas, the KEGG background showed enrichment mainly for antigen processing and presentation, Toll-like receptor signaling pathway, cell adhesion molecules (CAMs), and apoptosis (Fig. 3G) in the low-risk group of TCGA cohort. We obtained similar results in the validation cohort (Fig. 4A,B).
Prognostic-related immune cells identification. GSEA in the Hallmarks (A) and KEGG (B) gene set between group high (n = 27) and low (n = 26) of the risk score in the GSE21257 cohort. Survival analysis for osteosarcoma patients with different macrophages (C), M1 macrophages (D), M2 macrophages (E), plasma cells (F) and skeletal muscle cells (G) group. (H) Comparison of xCell score of the five prognostic related immune cells between high- and low-risk score group in the TCGA cohort.
Macrophages were associated with the prognosis and metastatic of OS
Supplementary File 6 shows the xCell score of 64 immune cells of each sample. Macrophages (Fig. 4C), M1 macrophages (Fig. 4D), M2 macrophages (Fig. 4E), plasma cells (Fig. 4F), and skeletal muscle cells (Fig. 4G) had a significant association with OS prognosis. Moreover, macrophages, M1 macrophages, and M2 macrophages were significantly more in the low-risk group than in the high-risk group (Fig. 4H). In addition, macrophages, M1 macrophages, and M2 macrophages negatively correlated with the risk score (Fig. 5A). Macrophages subtypes were significantly more in the non-relapse and non-metastatic groups (Fig. 5B,C). The GSEA results revealed that a majority of the genes showed enrichment for the allograft rejection, IL6/JAK/STAT3 signaling, IL2/STAT5 signaling, and apoptosis (Fig. 5D), whereas, the KEGG background showed enrichment mainly for antigen processing and presentation, Toll-like receptor signaling pathway, CAMs, and apoptosis (Fig. 5E) in the high-M1 macrophages group of TCGA cohort. Figure 6 illustrates similar results for the GSE21257 cohort.
The correlation of risk score with xCell score. (A) The correlation between the risk score and xCell score in the TCGA cohort. (B,C) Comparison of xCell score of the five prognostic related immune cells between relapse/metastatic and non-relapse/non-metastatic group in the TCGA cohort. GSEA in the Hallmarks (D) and KEGG (E) gene set between group high (n = 42) and low (n = 42) of M1 macrophages in the TCGA cohort.
The correlation of risk score with xCell score in the GSE21257 cohort. (A) Comparison of xCell score of the five prognostic related immune cells between high- and low-risk score group in the GSE21257 cohort. (B) Comparison of xCell score of the five prognostic related immune cells between the metastatic and non-metastatic group in the GSE21257 cohort. (C) The correlation between the risk score and xCell score of macrophages, M1 macrophages, M2 macrophages in the GSE21257 cohort. GSEA in the Hallmarks (D) and KEGG (E) gene set between group high (n = 27) and low (n = 26) of M1 macrophages in the GSE21257 cohort.
Risk score potential as an indicator of immunotherapy response in patients with OS
The expression of PD-1, PD-L1, and CTLA-4 was significantly higher in the low-risk score group than in the high-risk score group in the TCGA cohort (Fig. 7A). Immune checkpoint expression negatively correlated with the risk score (Fig. 7B). Patients with low-risk scores may have a better efficacy for immunotherapy in OS (Fig. 7C–E). Moreover, the immune checkpoints expression was also negatively correlated with the risk score in the validation cohort (Fig. 8A,B). The validation cohort showed a better efficacy for immunotherapy in OS (p > 0.05) for patients with low-risk scores (Fig. 8C–E).
The correlation of risk score with immune checkpoints in the TCGA cohort. (A) Comparison of expression of immune checkpoints between high- and low-risk score group (B) The correlation between the risk score and expression of immune checkpoints. Kaplan–Meier analysis of the four groups based on the risk score and the expression of PD-L1 (C), PD-1 (D) and CTLA-4 (E).
The correlation of risk score with immune checkpoints in the GSE21257 cohort. (A) Comparison of expression of immune checkpoints between high- and low-risk score group (B) The correlation between the risk score and expression of immune checkpoints. Kaplan–Meier analysis of the four groups based on the risk score and the expression of PD-L1 (C), PD-1 (D) and CTLA-4 (E).
Image analysis
All immunohistological images were acquired using an inverted microscope and collected for future use. In addition, we compared the staining of the OS specimens and peritumoral-normal specimens. Representative images in Fig. 9 illustrate that TYROBP, TLR4, and ITGAM were significantly down-regulated in OS than in the peritumoral-normal tissue. In addition, Fig. 9B,D,F showed that a positive rate of immunohistochemical staining for TYROBP, TLR4, and ITGAM in OS, which was significantly lower than that in the peritumoral normal tissue.
Immunohistochemical plots of the three hub genes associated with prognosis and statistical analysis of the positivity rate. (A,C,E) shows the protein expression of each gene in OS and in the paracancer. (B,D,F) shows the statistical analysis of the staining positivity rate for each gene in OS and in the paracancer. *Representative P-value < 0.05, **representative P-value < 0.01.
Discussion
The introduced effective therapy has improved in recent years; however, the 5-year survival rates of OS remains low19,20. Therefore, searching for effective novel therapies is of utmost importance. Immunotherapy revolutionized the treatment of cancer and has been employed in several cancers. As reported, the DEGs and the TIICs in the TME play a critical role in OS development21. Numerous studies have been conducted on OS immunotherapy, such as the TIIC regulation and immune checkpoint blockade22. However, studies lack biomarkers to assess patient prognosis and the efficacy of immunotherapy. The purpose of this study was to establish an immune-related gene signature for predicting the prognosis and efficacy of immunotherapy.
We constructed a three-immune-related gene-based risk model to predict the survival rate of OS patients more precisely. Reports state that TYROBP, TLR4, and ITGAM are involved in several cancer-immune microenvironment-associated pathogeneses, including OS23. The adapter protein TYROBP non-covalently associates with activating receptors on the surface of various immune cells to mediate signaling and cell activation after ligand binding by the receptors24. TYROBP is primarily enriched in natural killer cell-mediated cytotoxicity and osteoclast differentiation, which would lead to tumor cell apoptosis and promote osteoclast differentiation to cause bone resorption around the tumor. According to previous study reports, TYROBP is also involved in the activation of multiple immune cells, including T cells, B cells, and macrophages24,25,26. The low expression of TYROBP may promote the occurrence and progression of OS. As per the risk model, a low expression results in a higher risk.
It has been known that TLR4 plays a fundamental role in pathogen recognition and innate immunity activation27. In this study, TLR4 was involved in multiple biological processes (BP), including regulation of cytokine production and secretion, mononuclear cell proliferation, osteoclast differentiation, and macrophage activation. Furthermore, TLR4 activated the immune response in TME by up-regulating multiple immune cells, including T cells, leading to the anti-tumor effect28,29,30. Moreover, TLR4 facilitates osteoclast differentiation that may cause osteolytic destruction in the tumor surrounding area31,32. Natural barrier destruction may also be the cause of tumor metastasis. In addition, a high expression of TLR4 may improve prognosis through TME regulation that activates the immune system and facilitates tumor cells apoptosis.
It has been reported that ITGAM is implicated in various adhesive interactions of monocytes, macrophages, and granulocytes, as well as in mediating the uptake of complement-coated particles and pathogens. Studies have reported that ITGAM is significantly associated with tumor metastasis33,34. In our study, ITGAM was involved in the cell adhesion molecule pathway and also participated in an important BP, i.e., cell junction disassembly that plays a crucial role in metastasis. Hence, ITGAM may improve the prognosis of OS by inhibiting metastasis.
The literature reports that TYROBP, TLR4, and ITGAM are involved in the BP to activate macrophages that have been reported as tumor-associated and as the main component of the immune environment in OS35,36,37. We found that OS patients with elevated macrophage infiltration in the TME had a better prognosis. Compared to the high-risk score group, patients in the low-risk score group showed a significantly increased number of macrophages, M1 macrophages, and M2 macrophages. Furthermore, compared to non-metastatic/non-relapse cases, the metastatic/relapse cases showed markedly decreased levels of macrophages and M1 macrophages, which satisfactorily suggested that M0 to M1 macrophages polarization levels may be associated with improved outcomes in OS patients. M1 macrophages associated with non-metastasis displayed a pro-inflammatory phenotype and tumoricidal activity in OS10,37. Thus, our study results consolidate the previous data based on the beneficial role of M1 macrophage infiltration in OS.
Common immune checkpoint molecules, PD-1, PD-L1, and CTLA-438,39,40, are known to be related to the OS progress and prognosis. Numerous studies have shown the possibility of immunotherapy in OS41,42,43. Interestingly, the expression of PD‐L1 in OS patients is associated with a higher possibility for obtaining clinical benefits from immunotherapy44,45. Our risk score had a significantly negative correlation with PD-1, PD-L1, and CTLA-4. Moreover, TLR4, being an upstream receptor in the PD-L1 expression and PD-1 checkpoint pathway in cancer, was strongly associated with PD-L1 expression. Survival analysis showed that patients with in the low-risk score group experienced higher efficacy in immune checkpoint blockade.
In this study, the specific immune-related gene expression profile-based risk model could more precisely predict the survival of OS patients. Furthermore, the three immune-related genes regulated TIICs in the TME, especially macrophages activation, facilitating the apoptosis of OS cell. Moreover, our study demonstrated that the risk model could predict the immune checkpoint blockade efficacy in OS.
Conclusion
Overall, a three-immune-related gene-based risk model was constructed that could regulate macrophage activation and predict the survival rate and treatment outcomes to immune checkpoint blockade in OS.
Patients and methods
Microarray data acquisition
The TARGET-OS RNA-sequencing dataset (presented as fragments per kilobase million [FPKM]), as well as corresponding clinical characteristics and prognosis information of patients, were downloaded from UCSC Xena (https://xena.ucsc.edu/). However, patients with expression profiles but without clinical characteristics and prognostic information were excluded. Finally, we included 84 patients with OS in the training set. GSE21257 with 53 OS patients was downloaded from the GEO (www. ncbi.nlm.nih.gov/gds), a public repository at the National Center of Biotechnology Information as a validation cohort. In addition, GSE28424 and GSE19276 were downloaded to validate the differential expression of key genes between the tumor cells and normal bone.
Immune-related DEGs identification and PPI network construction
We applied the “Estimate” package (version 1.0.13) in R language (version 4.0.0) to estimate the immune-stromal component ratio in TME for each sample. TME results consisted of the immune score and stromal score. A higher respective score resulted in a larger ratio of the corresponding component in TME. A total of 84 samples were labeled with either a high score or low score as per the median score obtained for the immune score and stromal score. The “Limma” package (version 3.44.1) was applied to screen DEGs that were defined as an adjusted p-value < 0.05 and |log (fold-change)|> 1. Next, we performed the intersection analysis between the up- and down-regulated genes and included only the overlapping genes in the following analysis. The protein–protein interaction (PPI) network was predicted using an online database (STRING; http://string-db.org) (version 11.0b) search tool to retrieve interacting genes and visualized using Cytoscape (version 3.6.1). Molecular Complex Detection (MCODE) is used for clustering a given topology-based network11. In addition, clusters with nodes < 20 were discarded.
GO and KEGG enrichment analysis
All DEGs, including DEGs in each cluster, were analyzed for enrichment using the “clusterProfiler” package (version 3.16.0) and subsequently visualized using the “ggplot2” package (version 3.3.0) in R language12. GO and KEGG13 terms with the adjusted p-value < 0.05 were considered significantly enriched.
Hub genes identification and prognostic gene signature established
We used the “cytoHubba” in Cytoscape to identify the hub genes. The top 10 nodes ranked by “Degree,” “Betweenness,” and “Closeness” were enrolled in gene selection. The univariate Cox proportional hazard regression analysis and Kaplan–Meier estimates were used to identify prognostic genes. LASSO regression analyses were used to select the genes included in the model. The cutoff point, Lambda.min, brings minimum mean cross-validated error. We selected the genes with the highest lambda values for further analysis14. Next, the risk scores were calculated using the generated coefficients and corresponding expression, and the median risk score was obtained using the prognostic model. Finally, the latter was used to divide the patients into “high-risk” and “low-risk” groups. The diagnostic value of the risk score system was assessed using the time-dependent receiver operating characteristic (ROC) curve and survival analysis.
Relationships among risk score, immune cells, and immune checkpoints
The xCell tool provides 64 cell types, including lymphoid, myeloid, stromal cells, stem cells, and other cells. Hence, the xCell score analysis using the R package “xCell” (https://github.com/dviraran/xCell) allowed us to obtain 64 immune cell type abundance scores15. Furthermore, we performed a Kaplan–Meier analysis to identify prognostic immune cells. The prognostic-immune-cells difference between the two risk groups and clinical characteristics were compared using the t-test. The “corrplot” package (version 0.84) was used to perform Pearson’s correlation analysis between the risk score and the xCell score of immunity cells, as well as among the expression of immune checkpoints. Moreover, the prognostic-immune-cells difference between metastatic/relapse and non-metastatic/non-relapse group, and the difference between each immune checkpoint expression in the high- and low-risk score groups were compared using t-test. The patients were grouped into four groups based on the risk score and immune checkpoint expression. Lastly, their relationships with overall survival were analyzed using Kaplan–Meier analysis. We considered a p-value < 0.05 as the threshold for significance.
Gene set enrichment analysis (GSEA)
The computational method, GSEA (V4.0.3), determines whether a priori defined set of genes shows statistically significant concordant differences between two biological states16. Herein, GSEA investigated the potential biological characteristics between different statuses based on gene sets downloaded from the Molecular Signatures Database17. In this study, we only investigated the Hallmarks (h.all.v72.symbols.gmt) and c2 (c2.cp.kegg.v7.2.symbols.gmt) gene sets. The significant cutoff value was defined as the false discovery rate (FDR) < 0.25 and the normal p < 0.05.
Immunohistochemistry
The subjects volunteering for this study had signed informed consent forms. Moreover, our study was approved by the Ethics Department of the First Affiliated Hospital of Guangxi Medical University and conformed to the World Medical Association Declaration of Helsinki. Six pairs (OS and peritumoral normal tissue) of pathological sections for each gene were immunohistologically analyzed. For immunohistochemical staining, the TYROBP, TLR4 and ITGAM antibodies were purchased from Abcam. The paraffin, hydration, and seals were removed. Next, the specimens were mixed with anti-TYROBP, TLR4, and ITGAM antibodies at various dilution ratios (1:250, 1:200, 1:4000, 1:300, and 1:100) and incubated overnight at 4 °C18. Lastly, we statistically analyzed the positivity rate of the immunohistology images using Image J software and GraphPad Prism 8.
Ethics approval and consent to participate
All subjects volunteered for the study and signed informed consent forms. In order to ensure confidentiality, the names of study participants were not included in the data. Information obtained from the data of the study participants is kept confidential. In addition, the Ethics Committee of The First Affiliated Hospital of Guangxi Medical University approved the study.
Consent to publish
Consent for publication was obtained from all participants.
Data availability
The dataset generated or analyzed during the current study are available in the TCGA dataset repository (https://tcga-data.nci.nih.gov/tcga/) and Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/). And all data and materials for this study shall be availed whenever requested by the editorial team, reviewers, and other users.
Abbreviations
- GSEA:
-
Gene Set Enrichment Analysis
- GO:
-
Gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- DEGs:
-
Differentially expressed genes
- TYROBP:
-
TYRO protein tyrosine kinase binding protein
- TLR4:
-
Toll-like receptor 4
- ITGAM:
-
Integrin subunit alpha M
- PD-1:
-
Programmed Cell Death 1
- PD-L1:
-
Programmed Cell Death 1 Ligand 1
- CTLA-4:
-
Cytotoxic T-Lymphocyte Associated Protein 4
- TCGA:
-
The Cancer Genome Atlas Program
- TME:
-
Tumor microenvironment
- TIICs:
-
Tumor-infiltrating immune cells
- DEGs:
-
Differentially expressed genes
References
Han, K. et al. MicroRNA profiling identifies MiR-195 suppresses osteosarcoma cell metastasis by targeting CCND1. Oncotarget 6(11), 8875–8889. https://doi.org/10.18632/oncotarget.3560 (2015).
Zhuo, W. et al. MicroRNA20a promotes the proliferation and cell cycle of human osteosarcoma cells by suppressing early growth response 2 expression. Mol. Med. Rep. 12(4), 4989–4994. https://doi.org/10.3892/mmr.2015.4098 (2015).
Pei, Y., Yao, Q., Li, Y., Zhang, X. & Xie, B. microRNA-211 regulates cell proliferation, apoptosis and migration/invasion in human osteosarcoma via targeting EZRIN. Cell. Mol. Biol. Lett. 24, 48. https://doi.org/10.1186/s11658-019-0173-x (2019).
Yi, X. J. et al. Aberrant Wnt/beta-catenin signaling and elevated expression of stem cell proteins are associated with osteosarcoma side population cells of high tumorigenicity. Mol. Med. Rep. 12(4), 5042–5048. https://doi.org/10.3892/mmr.2015.4025 (2015).
Bao, J. et al. A Retrospective clinicopathological study of osteosarcoma patients with metachronous metastatic relapse. J. Cancer 10(13), 2982–2990. https://doi.org/10.7150/jca.30750 (2019).
Roy, J., Wycislo, K. L., Pondenis, H., Fan, T. M. & Das, A. Comparative proteomic investigation of metastatic and non-metastatic osteosarcoma cells of human and canine origin. PLoS ONE 12(9), e0183930. https://doi.org/10.1371/journal.pone.0183930 (2017).
Mao, Y. et al. Low tumor infiltrating mast cell density confers prognostic benefit and reflects immunoactivation in colorectal cancer. Int. J. Cancer 143(9), 2271–2280. https://doi.org/10.1002/ijc.31613 (2018).
Kurbatov, V. et al. Digital inference of immune microenvironment reveals low-risk subtype of early lung adenocarcinoma. Ann. Thorac. Surg. 109(2), 343–349. https://doi.org/10.1016/j.athoracsur.2019.08.050 (2020).
Gajewski, T. F., Schreiber, H. & Fu, Y. X. Innate and adaptive immune cells in the tumor microenvironment. Nat. Immunol. 14(10), 1014–1022. https://doi.org/10.1038/ni.2703 (2013).
Torabi, A., Amaya, C. N., Wians, F. H. Jr. & Bryan, B. A. PD-1 and PD-L1 expression in bone and soft tissue sarcomas. Pathology 49(5), 506–513. https://doi.org/10.1016/j.pathol.2017.05.003 (2017).
Yi, X. H., Zhang, B., Fu, Y. R. & Yi, Z. J. STAT1 and its related molecules as potential biomarkers in Mycobacterium tuberculosis infection. J. Cell Mol. Med. 24(5), 2866–2878. https://doi.org/10.1111/jcmm.14856 (2020).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2009).
Kanehisa, M., Furumichi, M., Sato, Y., Ishiguro-Watanabe, M. & Tanabe, M. KEGG: Integrating viruses and cellular organisms. Nucleic Acids Res. 49(D1), D545–D551. https://doi.org/10.1093/nar/gkaa970 (2021).
Chen, P. et al. Identification of prognostic immune-related genes in the tumor microenvironment of endometrial cancer. Aging 12(4), 3371–3387. https://doi.org/10.18632/aging.102817 (2020).
Aran, D. Cell-type enrichment analysis of bulk transcriptomes using xCell. Methods Mol. Biol. 2120, 263–276. https://doi.org/10.1007/978-1-0716-0327-7_19 (2020).
Tang, G. & Yin, W. Development of an immune infiltration-related prognostic scoring system based on the genomic landscape analysis of glioblastoma multiforme. Front. Oncol. 10, 154. https://doi.org/10.3389/fonc.2020.00154 (2020).
Lin, K. et al. Development of a prognostic index and screening of potential biomarkers based on immunogenomic landscape analysis of colorectal cancer. Aging 12(7), 5832–5857. https://doi.org/10.18632/aging.102979 (2020).
Jiang, J. et al. TRIM68, PIKFYVE, and DYNLL2: The possible novel autophagy- and immunity-associated gene biomarkers for osteosarcoma prognosis. Front. Oncol. 11, 643104. https://doi.org/10.3389/fonc.2021.643104 (2021).
Zhang, J., Lan, Q. & Lin, J. Identification of key gene modules for human osteosarcoma by co-expression analysis. World J. Surg. Oncol. 16(1), 89. https://doi.org/10.1186/s12957-018-1381-y (2018).
Robl, B. et al. Evaluation of F8-TNF-α in models of early and progressive metastatic osteosarcoma. Transl. Oncol. 10(3), 419–430. https://doi.org/10.1016/j.tranon.2017.02.005 (2017).
Zhang, C. et al. Profiles of immune cell infiltration and immune-related genes in the tumor microenvironment of osteosarcoma. Aging 12(4), 3486–3501. https://doi.org/10.18632/aging.102824 (2020).
Zhang, J., Li, Y., Yang, S., Zhang, L. & Wang, W. Anti-CD40 mAb enhanced efficacy of anti-PD1 against osteosarcoma. J. Bone Oncol. 17, 100245. https://doi.org/10.1016/j.jbo.2019.100245 (2019).
Pan, Y., Lu, L., Chen, J., Zhong, Y. & Dai, Z. Identification of potential crucial genes and construction of microRNA-mRNA negative regulatory networks in osteosarcoma. Hereditas 155, 21. https://doi.org/10.1186/s41065-018-0061-9 (2018).
Lanier, L. L., Corliss, B., Wu, J. & Phillips, J. H. Association of DAP12 with activating CD94/NKG2C NK cell receptors. Immunity 8(6), 693–701. https://doi.org/10.1016/s1074-7613(00)80574-9 (1998).
Ono, Y. et al. Graft-infiltrating PD-L1(hi) cross-dressed dendritic cells regulate antidonor T cell responses in mouse liver transplant tolerance. Hepatology 67(4), 1499–1515. https://doi.org/10.1002/hep.29529 (2018).
Zheng, L. et al. A humanized Lym-1 CAR with novel DAP10/DAP12 signaling domains demonstrates reduced tonic signaling and increased antitumor activity in B-cell lymphoma models. Clin. Cancer Res. 26(14), 3694–3706. https://doi.org/10.1158/1078-0432.Ccr-19-3417 (2020).
Shetab Boushehri, M. A. & Lamprecht, A. TLR4-Based Immunotherapeutics in Cancer: A Review of the Achievements and Shortcomings. Mol Pharm 15(11), 4777–4800. https://doi.org/10.1021/acs.molpharmaceut.8b00691 (2018).
Bock, M. et al. The posttraumatic activation of CD4+ T regulatory cells is modulated by TNFR2- and TLR4-dependent pathways, but not by IL-10. Cell. Immunol. 331, 137–145. https://doi.org/10.1016/j.cellimm.2018.06.009 (2018).
Dias, A. S. O. et al. TLR-2 and TLR-4 agonists favor expansion of CD4(+) T cell subsets implicated in the severity of neuromyelitis optica spectrum disorders. Mult. Scler. Relat. Disord. 34, 66–76. https://doi.org/10.1016/j.msard.2019.06.018 (2019).
Yahiro, K. et al. Activation of TLR4 signaling inhibits progression of osteosarcoma by stimulating CD8-positive cytotoxic lymphocytes. Cancer Immunol. Immunother. 69(5), 745–758. https://doi.org/10.1007/s00262-020-02508-9 (2020).
Xie, H., Cao, L., Ye, L., Shan, G. & Song, W. The miR-1906 mimic attenuates bone loss in osteoporosis by down-regulating the TLR4/MyD88/NF-κB pathway. Physiol. Int. https://doi.org/10.1556/2060.2020.00042 (2021).
Jiang, H. et al. TREM-2 promotes acquired cholesteatoma-induced bone destruction by modulating TLR4 signaling pathway and osteoclasts activation. Sci. Rep. 6, 38761. https://doi.org/10.1038/srep38761 (2016).
Boguslawska, J. et al. Expression of genes involved in cellular adhesion and extracellular matrix remodeling correlates with poor survival of patients with renal cancer. J. Urol. 195(6), 1892–1902. https://doi.org/10.1016/j.juro.2015.11.050 (2016).
Lyu, T. et al. SMYD3 promotes implant metastasis of ovarian cancer via H3K4 trimethylation of integrin promoters. Int. J. Cancer 146(6), 1553–1567. https://doi.org/10.1002/ijc.32673 (2020).
Withers, S. S. et al. Association of macrophage and lymphocyte infiltration with outcome in canine osteosarcoma. Vet. Comp. Oncol. 17(1), 49–60. https://doi.org/10.1111/vco.12444 (2019).
Wolf-Dennen, K., Gordon, N. & Kleinerman, E. S. Exosomal communication by metastatic osteosarcoma cells modulates alveolar macrophages to an M2 tumor-promoting phenotype and inhibits tumoricidal functions. Oncoimmunology 9(1), 1747677. https://doi.org/10.1080/2162402x.2020.1747677 (2020).
Gomez-Brouchet, A. et al. CD163-positive tumor-associated macrophages and CD8-positive cytotoxic lymphocytes are powerful diagnostic markers for the therapeutic stratification of osteosarcoma patients: An immunohistochemical analysis of the biopsies fromthe French OS2006 phase 3 trial. Oncoimmunology 6(9), e1331193. https://doi.org/10.1080/2162402X.2017.1331193 (2017).
Luo, R. et al. Cisplatin facilitates radiation-induced abscopal effects in conjunction with PD-1 checkpoint blockade through CXCR3/CXCL10-mediated T-cell recruitment. Clin. Cancer Res. 25(23), 7243–7255. https://doi.org/10.1158/1078-0432.Ccr-19-1344 (2019).
Donia, M. et al. PD-1(+) polyfunctional T cells dominate the periphery after tumor-infiltrating lymphocyte therapy for Cancer. Clin. Cancer Res. 23(19), 5779–5788. https://doi.org/10.1158/1078-0432.Ccr-16-1692 (2017).
Wei, S. C. et al. Distinct cellular mechanisms underlie anti-CTLA-4 and Anti-PD-1 checkpoint blockade. Cell 170(6), 1120-1133.e1117. https://doi.org/10.1016/j.cell.2017.07.024 (2017).
Silva, M. A. et al. Biomarker recommendation for PD-1/PD-L1 immunotherapy development in pediatric cancer based on digital image analysis of PD-L1 and immune cells. J. Pathol. Clin. Res. 6(2), 124–137. https://doi.org/10.1002/cjp2.152 (2020).
Zheng, B. et al. PD-1 axis expression in musculoskeletal tumors and antitumor effect of nivolumab in osteosarcoma model of humanized mouse. J. Hematol. Oncol. 11(1), 16. https://doi.org/10.1186/s13045-018-0560-1 (2018).
Toda, Y. et al. PD-L1 and IDO1 expression and tumor-infiltrating lymphocytes in osteosarcoma patients: Comparative study of primary and metastatic lesions. J. Cancer Res. Clin. Oncol. https://doi.org/10.1007/s00432-020-03242-6 (2020).
Zhang, M. L., Chen, L., Li, Y. J. & Kong, D. L. PD-L1/PD-1 axis serves an important role in natural killer cell-induced cytotoxicity in osteosarcoma. Oncol. Rep. 42(5), 2049–2056. https://doi.org/10.3892/or.2019.7299 (2019).
Liu, X. et al. Blocking the PD-1/PD-L1 axis enhanced cisplatin chemotherapy in osteosarcoma in vitro and in vivo. Environ. Health Prev. Med. 24(1), 79. https://doi.org/10.1186/s12199-019-0835-3 (2019).
Acknowledgements
We are grateful to Dr. Xinli Zhan (Spine and Osteopathy Ward, The First Affiliated Hospital of Guangxi Medical University) for his kindly assistance in all stages of the present study.
Funding
This work was sponsored by the National Natural Science Foundation of China (81560359); National Natural Science Foundation of China (81860393). Funding bodies had no role in the study design, collection, analysis, and interpretation of the data or in writing the manuscript.
Author information
Authors and Affiliations
Contributions
X.Z., T.L.: Conceptualization, Methodology. J.C., G.Y.X., Z.Z.: Data curation, Investigation. J.X., H.Z., J.J.: Formal analysis, Software. Z.Q., L.H., .T.C., Z.Y., .Y.N.: Visualization. T.L., C.L.: Writing- Reviewing and Editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liang, T., Chen, J., Xu, G. et al. TYROBP, TLR4 and ITGAM regulated macrophages polarization and immune checkpoints expression in osteosarcoma. Sci Rep 11, 19315 (2021). https://doi.org/10.1038/s41598-021-98637-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-98637-x
This article is cited by
-
Programmed Cell Death-Related Gene Signature Associated with Prognosis and Immune Infiltration and the Roles of HMOX1 in the Proliferation and Apoptosis were Investigated in Uveal Melanoma
Genes & Genomics (2024)
-
The role of SPI1-TYROBP-FCER1G network in oncogenesis and prognosis of osteosarcoma, and its association with immune infiltration
BMC Cancer (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.