Abstract
Biogenic monoamines—vital transmitters orchestrating neurological, endocrinal and immunological functions1,2,3,4,5—are stored in secretory vesicles by vesicular monoamine transporters (VMATs) for controlled quantal release6,7. Harnessing proton antiport, VMATs enrich monoamines around 10,000-fold and sequester neurotoxicants to protect neurons8,9,10. VMATs are targeted by an arsenal of therapeutic drugs and imaging agents to treat and monitor neurodegenerative disorders, hypertension and drug addiction1,8,11,12,13,14,15,16. However, the structural mechanisms underlying these actions remain unclear. Here we report eight cryo-electron microscopy structures of human VMAT1 in unbound form and in complex with four monoamines (dopamine, noradrenaline, serotonin and histamine), the Parkinsonism-inducing MPP+, the psychostimulant amphetamine and the antihypertensive drug reserpine. Reserpine binding captures a cytoplasmic-open conformation, whereas the other structures show a lumenal-open conformation stabilized by extensive gating interactions. The favoured transition to this lumenal-open state contributes to monoamine accumulation, while protonation facilitates the cytoplasmic-open transition and concurrently prevents monoamine binding to avoid unintended depletion. Monoamines and neurotoxicants share a binding pocket that possesses polar sites for specificity and a wrist-and-fist shape for versatility. Variations in this pocket explain substrate preferences across the SLC18 family. Overall, these structural insights and supporting functional studies elucidate the mechanism of vesicular monoamine transport and provide the basis to develop therapeutics for neurodegenerative diseases and substance abuse.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The cryo-EM maps have been deposited at the Electron Microscopy Data Bank (EMDB) under accession numbers EMD-41238 (unbound/reserpine), EMD-41241 (reserpine/reserpine), EMD-41237 (dopamine/reserpine), EMD-41240 (noradrenaline/reserpine), EMD-41242 (serotonin/reserpine), EMD-41239 (histamine/reserpine), EMD-41236 (amphetamine/reserpine) and EMD-41235 (MPP+/reserpine). The coordinates have been deposited at the Protein Data Bank (PDB) under accession numbers 8TGJ (unbound/reserpine), 8TGM (reserpine/reserpine), 8TGI (dopamine/reserpine), 8TGL (noradrenaline/reserpine), 8TGN (serotonin/reserpine), 8TGK (histamine/reserpine), 8TGH (amphetamine/reserpine) and 8TGG (MPP+/reserpine). Source data are provided with this paper.
References
Carlsson, A. A half-century of neurotransmitter research: impact on neurology and psychiatry. Biosci. Rep. 21, 691–710 (2001).
Sara, S. J. The locus coeruleus and noradrenergic modulation of cognition. Nat. Rev. Neurosci. 10, 211–223 (2009).
Mawe, G. M. & Hoffman, J. M. Serotonin signalling in the gut-functions, dysfunctions and therapeutic targets. Nat. Rev. Gastroenterol. Hepatol. 10, 473–486 (2013).
Carey, R. M. Renal dopamine system: paracrine regulator of sodium homeostasis and blood pressure. Hypertension 38, 297–302 (2001).
Stone, K. D., Prussin, C. & Metcalfe, D. D. IgE, mast cells, basophils, and eosinophils. J. Allergy Clin. Immunol. 125, S73–S80 (2010).
Erickson, J. D. & Eiden, L. E. Functional identification and molecular cloning of a human brain vesicle monoamine transporter. J. Neurochem. 61, 2314–2317 (1993).
Erickson, J. D., Eiden, L. E. & Hoffman, B. J. Expression cloning of a reserpine-sensitive vesicular monoamine transporter. Proc. Natl Acad. Sci. USA 89, 10993–10997 (1992).
Eiden, L. E., Schafer, M. K., Weihe, E. & Schutz, B. The vesicular amine transporter family (SLC18): amine/proton antiporters required for vesicular accumulation and regulated exocytotic secretion of monoamines and acetylcholine. Pflugers Arch. 447, 636–640 (2004).
Parsons, S. M. Transport mechanisms in acetylcholine and monoamine storage. FASEB J. 14, 2423–2434 (2000).
Liu, Y. et al. A cDNA that suppresses MPP+ toxicity encodes a vesicular amine transporter. Cell 70, 539–551 (1992).
Carlsson, A. & Healy, D. in The Psychopharmacologists (ed. Healy, D.) Vol. 1, 51–80 (CRC Press, 1996).
Baumeister, A. A. The chlorpromazine enigma. J. Hist. Neurosci. 22, 14–29 (2013).
Kenney, C. & Jankovic, J. Tetrabenazine in the treatment of hyperkinetic movement disorders. Expert Rev. Neurother. 6, 7–17 (2006).
Efange, S. M. In vivo imaging of the vesicular acetylcholine transporter and the vesicular monoamine transporter. FASEB J. 14, 2401–2413 (2000).
Erickson, J. D., Schafer, M. K., Bonner, T. I., Eiden, L. E. & Weihe, E. Distinct pharmacological properties and distribution in neurons and endocrine cells of two isoforms of the human vesicular monoamine transporter. Proc. Natl Acad. Sci. USA 93, 5166–5171 (1996).
Sulzer, D., Sonders, M. S., Poulsen, N. W. & Galli, A. Mechanisms of neurotransmitter release by amphetamines: a review. Prog. Neurobiol. 75, 406–433 (2005).
Peter, D., Jimenez, J., Liu, Y., Kim, J. & Edwards, R. H. The chromaffin granule and synaptic vesicle amine transporters differ in substrate recognition and sensitivity to inhibitors. J. Biol. Chem. 269, 7231–7237 (1994).
Erickson, J. D. et al. Functional identification of a vesicular acetylcholine transporter and its expression from a “cholinergic” gene locus. J. Biol. Chem. 269, 21929–21932 (1994).
Lohoff, F. W. et al. Association between polymorphisms in the vesicular monoamine transporter 1 gene (VMAT1/SLC18A1) on chromosome 8p and schizophrenia. Neuropsychobiology 57, 55–60 (2008).
Hu, G. et al. New fluorescent substrate enables quantitative and high-throughput examination of vesicular monoamine transporter 2 (VMAT2). ACS Chem. Biol. 8, 1947–1954 (2013).
Anne, C. & Gasnier, B. Vesicular neurotransmitter transporters: mechanistic aspects. Curr. Top. Membr. 73, 149–174 (2014).
Darchen, F., Scherman, D. & Henry, J. P. Reserpine binding to chromaffin granules suggests the existence of two conformations of the monoamine transporter. Biochemistry 28, 1692–1697 (1989).
Quistgaard, E. M., Low, C., Guettou, F. & Nordlund, P. Understanding transport by the major facilitator superfamily (MFS): structures pave the way. Nat. Rev. Mol. Cell Biol. 17, 123–132 (2016).
Law, C. J., Maloney, P. C. & Wang, D. N. Ins and outs of major facilitator superfamily antiporters. Annu. Rev. Microbiol. 62, 289–305 (2008).
Yaffe, D., Vergara-Jaque, A., Forrest, L. R. & Schuldiner, S. Emulating proton-induced conformational changes in the vesicular monoamine transporter VMAT2 by mutagenesis. Proc. Natl Acad. Sci. USA 113, E7390–E7398 (2016).
Lohoff, F. W. Genetic variants in the vesicular monoamine transporter 1 (VMAT1/SLC18A1) and neuropsychiatric disorders. Methods Mol. Biol. 637, 165–180 (2010).
Lohoff, F. W. et al. Functional genetic variants in the vesicular monoamine transporter 1 modulate emotion processing. Mol. Psychiatry 19, 129–139 (2014).
Sulzer, D. How addictive drugs disrupt presynaptic dopamine neurotransmission. Neuron 69, 628–649 (2011).
Deupree, J. D. & Weaver, J. A. Identification and characterization of the catecholamine transporter in bovine chromaffin granules using [3H]reserpine. J. Biol. Chem. 259, 10907–10912 (1984).
Scherman, D. & Henry, J. P. Reserpine binding to bovine chromaffin granule membranes. Characterization and comparison with dihydrotetrabenazine binding. Mol. Pharmacol. 25, 113–122 (1984).
Vardy, E., Steiner-Mordoch, S. & Schuldiner, S. Characterization of bacterial drug antiporters homologous to mammalian neurotransmitter transporters. J. Bacteriol. 187, 7518–7525 (2005).
Yaffe, D., Forrest, L. R. & Schuldiner, S. The ins and outs of vesicular monoamine transporters. J. Gen. Physiol. 150, 671–682 (2018).
Wimalasena, K. Vesicular monoamine transporters: structure-function, pharmacology, and medicinal chemistry. Med. Res. Rev. 31, 483–519 (2011).
Pidathala, S. et al. Mechanisms of neurotransmitter transport and drug inhibition in human VMAT2. Nature 623, 1086–1092 (2023).
Yaffe, D. et al. Functionally important carboxyls in a bacterial homologue of the vesicular monoamine transporter (VMAT). J. Biol. Chem. 289, 34229–34240 (2014).
Yaffe, D., Radestock, S., Shuster, Y., Forrest, L. R. & Schuldiner, S. Identification of molecular hinge points mediating alternating access in the vesicular monoamine transporter VMAT2. Proc. Natl Acad. Sci. USA 110, E1332–E1341 (2013).
Shirvan, A., Laskar, O., Steiner-Mordoch, S. & Schuldiner, S. Histidine-419 plays a role in energy coupling in the vesicular monoamine transporter from rat. FEBS Lett. 356, 145–150 (1994).
Steiner-Mordoch, S., Shirvan, A. & Schuldiner, S. Modification of the pH profile and tetrabenazine sensitivity of rat VMAT1 by replacement of aspartate 404 with glutamate. J. Biol. Chem. 271, 13048–13054 (1996).
Schuldiner, S. Competition as a way of life for H+-coupled antiporters. J. Mol. Biol. 426, 2539–2546 (2014).
Maron, R., Stern, Y., Kanner, B. I. & Schuldiner, S. Functional asymmetry of the amine transporter from chromaffin granules. J. Biol. Chem. 258, 11476–11481 (1983).
Khare, P., Mulakaluri, A. & Parsons, S. M. Search for the acetylcholine and vesamicol binding sites in vesicular acetylcholine transporter: the region around the lumenal end of the transport channel. J. Neurochem. 115, 984–993 (2010).
Ojeda, A. M., Kolmakova, N. G. & Parsons, S. M. Acetylcholine binding site in the vesicular acetylcholine transporter. Biochemistry 43, 11163–11174 (2004).
Ugolev, Y., Segal, T., Yaffe, D., Gros, Y. & Schuldiner, S. Identification of conformationally sensitive residues essential for inhibition of vesicular monoamine transport by the noncompetitive inhibitor tetrabenazine. J. Biol. Chem. 288, 32160–32171 (2013).
Peter, D., Vu, T. & Edwards, R. H. Chimeric vesicular monoamine transporters identify structural domains that influence substrate affinity and sensitivity to tetrabenazine. J. Biol. Chem. 271, 2979–2986 (1996).
Finn, J. P. 3rd & Edwards, R. H. Multiple residues contribute independently to differences in ligand recognition between vesicular monoamine transporters 1 and 2. J. Biol. Chem. 273, 3943–3947 (1998).
Kanner, B. I., Fishkes, H., Maron, R., Sharon, I. & Schuldiner, S. Reserpine as a competitive and reversible inhibitor of the catecholamine transporter of bovine chromaffin granules. FEBS Lett. 100, 175–178 (1979).
Schuldiner, S., Liu, Y. & Edwards, R. H. Reserpine binding to a vesicular amine transporter expressed in Chinese hamster ovary fibroblasts. J. Biol. Chem. 268, 29–34 (1993).
Kilbourn, M. R., Lee, L. C., Heeg, M. J. & Jewett, D. M. Absolute configuration of (+)-α-dihydrotetrabenazine, an active metabolite of tetrabenazine. Chirality 9, 59–62 (1997).
Dwoskin, L. P. & Crooks, P. A. A novel mechanism of action and potential use for lobeline as a treatment for psychostimulant abuse. Biochem. Pharmacol. 63, 89–98 (2002).
Lohr, K. M. & Miller, G. W. VMAT2 and Parkinson’s disease: harnessing the dopamine vesicle. Expert Rev. Neurother. 14, 1115–1117 (2014).
Rubinstein, J. L. & Brubaker, M. A. Alignment of cryo-EM movies of individual particles by optimization of image translations. J. Struct. Biol. 192, 188–195 (2015).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. D. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Black, C. A. et al. Assessing vesicular monoamine transport and toxicity using fluorescent false neurotransmitters. Chem. Res. Toxicol. 34, 1256–1264 (2021).
Lomize, A. L., Todd, S. C. & Pogozheva, I. D. Spatial arrangement of proteins in planar and curved membranes by PPM 3.0. Protein Sci. 31, 209–220 (2022).
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Van Der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Van Zundert, G. C. P. et al. The HADDOCK2.2 web server: user-friendly integrative modeling of biomolecular complexes. J. Mol. Biol. 428, 720–725 (2016).
Acknowledgements
We thank Andrzej Krezel for comments. W.L. is supported by the Vagelos Endowed Chair, the Established Investigator Award and Collaborative Sciences Award from American Heart Association, the Forefront of Science Award from the W. M. Keck Foundation, the National Heart, Lung and Blood Institute (R01 HL121718), the Children’s Discovery Institute (MCII 2020-854), the National Institute of Allergy and Infectious Diseases (R01 AI158500) and the National Institute of General Medical Sciences (R01 GM131008). B.L. is supported by the Hormel Institute, University of Minnesota. K.W. is supported by Lundbeck foundation (R324-2019-1855). J.X. is supported by National Institute of Neurological Disorders and Stroke (R01 NS092865).
Author information
Authors and Affiliations
Contributions
J.Y. performed cryo-EM and functional experiments with assistance from Y.W., A.A., S.A. and K.W.; B.L., W.L. and J.Y. determined the structures. H.C. and J.Y. performed uptake assays with J.X.’s guidance. W.L. directed the project. W.L. and J.Y. wrote the manuscript with input from all of the authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Simon Newstead, Shimon Schuldiner and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The VMAT1(ΔL1/2) construct enables the protein purification and is active in monoamine uptake.
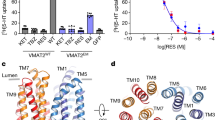
a, FSEC profile comparison of full-length VMAT1 and VMAT1(ΔL1/2). These proteins are tagged with a C-terminal GFP, expressed in HEK293 or Pichia pastoris cells, and extracted in DDM. b, Vesicular uptake of FFN206 by full-length VMAT1 and VMAT1(ΔL1/2). Both can transport this false fluorescence neurotransmitter into the vesicle-like compartments of HEK293 cells (shown in single cells). The activities of full-length VMAT1 and VMAT1(ΔL1/2) are both contingent upon the proton gradient (collapsed by bafilomycin A1, BFA) and inhibitable by reserpine (RSP). c, View of multiple cells for fluorescence quantification of FFN206 transport. Experiments in b and c were repeated independently three times with similar results. d, Dose-dependent FFN206 transport curves. The FFN206 fluorescence is normalized to GFP fluorescence to account for variations in protein expression level. e, Time course of [3H]-dopamine and [3H]-serotonin uptake assays. Data in d and e are shown as mean ± s.e.m. from n = 3 biological replicates.
Extended Data Fig. 2 Reserpine treatment during protein purification induces the dimer formation of human VMAT1.
a, 2D classification showing that VMAT1 protein forms monomers without reserpine treatment (top) and forms dimers with reserpine treatment (bottom). The monomer particles were extracted at box size of 320 pixels (0.664 Å/pixel), and VAMT1 dimer at box size of 320 pixels (0.885 Å/pixel). b, Size-exclusion chromatography (Superose 6) shows a shift to higher molecular weight after reserpine treatment, indicative of dimer formation. The elution profiles are from size-exclusion chromatography after affinity purification (left) and a subsequent rerun (right). c, SDS-PAGE of eluted fractions from the size-exclusion rerun. Uncropped gel is shown in Supplementary Fig. 2. Purifications were repeated independently at least three times with similar results.
Extended Data Fig. 3 Cryo-EM data processing flowchart and density maps of individual regions in VMAT1(ΔL1/2) structure.
The dataset and 3.5 Å density map of VMAT1(ΔL1/2) dimer with amphetamine and reserpine are shown as a representative. Other datasets are similarly processed, and density maps are similar. a, A representative raw cryo-EM image. Similar images are present in the 6,930 movies selected (see c). b, Representative 2D classes. c, The data processing procedure. Three rounds of 2D classification and two rounds of heterogenous refinements generated 3.54 Å and 3.49 Å final maps for two different classes. The maps were sharpened by DeepEMhancer for model building and analysis. d-e, Densities of helices and loops in the lumenal-open (d) and cytoplasmic-open (e) monomer. Contour level = 0.12 in ChimeraX.
Extended Data Fig. 4 Quality of cryo-EM maps.
a-h, VMAT1(ΔL1/2) dimer in unbound form and with reserpine (a); both with reserpine (b); with dopamine (DA) and reserpine (c); with noradrenaline (NE) and reserpine (d); with serotonin (SERT) and reserpine (e); with histamine (HA) and reserpine (f); with amphetamine (AMPH) and reserpine (g); and with MPP+ and reserpine (h). Left to right, Local resolution illustrations of the cryo-EM maps, angular distribution plots, half-map FSC curves, and model-to-map FSCs.
Extended Data Fig. 5 VMAT1 structures with bound reserpine and in unbound form.
a, Cryo-EM density map of the VMAT1 dimer, in which both monomers bind reserpine (density coloured in orange). Map contour level = 0.22 in ChimeraX. b, Helical tube representation of the dual reserpine-bound dimer structure. c, Hydrophobic effect of reserpine binding. Top, molecular structure of reserpine fits into a narrow gap at the bottom of the central cavity. Middle and bottom, The NTD and CTD sides of this gap both contain hydrophobic patches (yellow) that match well with the molecular shape of reserpine. d, Relative rigid-body rotation of the NTD and CTD between the cytoplasmic-open and lumenal-open conformations, and bending of TM5 and TM11. The NTD (left) and CTD (right) from each conformation are superimposed and shown in a side view. e, Top view of the superimposition. f, The NTD and CTD in the lumenal-open state are closely related by a 2-fold rigid body rotation. Left, the overall structure of the lumenal-open state. Right, superimposition of the NTD and CTD. TM5 and TM11, shown in dark colours, are replicated helices related by +6 numbering (as for other NTD and CTD helices). g, The NTD and CTD in the cytoplasmic-open state. Left, overall structure of the cytoplasmic-open state. Right, superimposition of the NTD and CTD.
Extended Data Fig. 6 Sequence alignment of SLC18 transporters.
The sequence alignment of human (h) and rat (r) SCL18 members was generated by ClustalW. The asterisks below the aligned sequences indicate conserved residues, and single and double dots represent varying levels of sequence similarity. Residues participating in substrate and reserpine binding are highlighted with orange shading and green spheres below, respectively. Unique gating residues in the cytoplasmic-open or lumenal-open conformations are indicated by green and blue bars, respectively. Interaction analysis was performed using the CONTACT program from the CCP4i suite. The potential protonation sites are indicated by red triangles. The protein folding topology (cyto: cytoplasmic side, lum: lumenal side), based on the VMAT1 structure, is denoted above the sequences. The sequence logo, produced by WebLogo, depicts the residue conservation level at each position. Initial identification of eukaryotic SLC18 homologues was achieved through the PSI-BLAST program within the Max-Planck Institute Bioinformatics Toolkit, using VMAT1 as the query sequence against a nonredundant (70%) sequence database, eukaryotes NR70. The top 500 proteins were selected, and incomplete sequences were manually removed. The multiple sequence alignment was generated by ClustalΩ. The motifs identified in the DHA-1 drug antiporter family are displayed above the sequence.
Extended Data Fig. 7 Substrate and inhibitor binding.
a-b, Structural superimposition of VMAT1 in unbound form and with bound dopamine showing the overall structures (a) and the substrate-binding pocket (b). c-e, Left, surface representation of the wrist-and-fist binding pocket with bound noradrenaline (c), serotonin (d), and histamine (e). Right, comparison of the binding of these monoamines (orange) with dopamine (dark grey). f, Mutations at the substrate-binding pocket reduces the binding affinity of FFN206. The binding of VMAT1 mutants is assessed by the fluorescence polarization of FFN206. Because most mutants do not achieve saturation in FFN206 binding even at very high protein concentration (100 µM), their relative affinities are estimated through binding potential (Bmax/Kd). Data are shown as mean ± s.e.m. from n = 3 biological replicates. g-h, Left, Amphetamine and MPP+ in the substrate binding pocket (surface representation). Right, comparison of their binding (orange) with dopamine (dark grey). i, Reserpine occupies the monoamine binding pocket. Left, the dopamine (dark grey) bound structure is superimposed onto the reserpine (orange) bound structure by the CTD. The protein sidechains illustrated are from the fixed reserpine-bound structure. Right, a 90-degree rotated view with the binding pocket (from the fixed reserpine-bound structure) shown in surface representation.
Extended Data Fig. 8 Candidate protonation sites.
a, Overall structures of the lumenal-open (left) and cytoplasmic-open conformations (right) with all candidate protonation sites. The lumenal unexposed regions in both conformations are highlighted by the orange circle. b, Local environment of D34 (zoomed view of dashed box in a) in the alternate conformations of VMAT1. c, A representative conformation from the molecular dynamics (MD) simulation of protonated E320 (E320-p, as in Fig. 5e) showing a similar NTD-CTD association as those in the cryo-EM structures. Left, Structure of the E320-p MD conformation (different orange colours). Middle, E320-p conformation is similar to that of the lumenal-open state (different blue colours) at the cytoplasmic side (orange circle). Right, E320-p conformation is also similar to that of the cytoplasmic-open state (different green colours) at the lumenal side (orange circle).
Extended Data Fig. 9 Homology model of VAChT.
Changes in interactions at the altered substrate-binding pocket may enable the recognition of acetylcholine (ACh). Residues in VMAT1 are in parentheses, with altered residues in VAChT highlighted in bold. Surface representation uses the altered binding pocket.
Supplementary information
Supplementary Information
Supplementary Discussion, Supplementary Table 1 and Supplementary Figs. 1 and 2.
Supplementary Video 1 Molecular dynamics simulation of VMAT1 with and without Glu320 protonation.
The simulation with protonated Glu320 (same as in Fig. 5e) shows lumenal NTD–CTD association (orange) similar to that in the cytoplasmic-open state (green) from the cryo-EM structure. By contrast, the unprotonated state (blue) does not show this lumenal NTD–CTD association. The bottom view from the lumenal side is shown, with TM helices indicated by numbers.
Supplementary Video 2 Model of proton antiport and monoamine accumulation.
The lumenal-open structure and cytoplasmic-open structure are superimposed by the CTD. The dopamine molecule is shown as presented in the lumenal-open state. The monoamine-binding pocket is conserved across the alternate conformations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ye, J., Chen, H., Wang, K. et al. Structural insights into vesicular monoamine storage and drug interactions. Nature 629, 235–243 (2024). https://doi.org/10.1038/s41586-024-07290-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-024-07290-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.