Abstract
Bone marrow (BM) cells depend on their niche for growth and survival. However, the genes modulated by niche stimuli have not been discriminated yet. For this purpose, we investigated BM aspirations from patients with various hematological malignancies. Each aspirate was fractionated, and the various samples were fixed at different time points and analyzed by microarray. Identification of niche-modulated genes relied on sustained change in expression following loss of niche regulation. Compared with the reference (‘authentic’) samples, which were fixed immediately following aspiration, the BM samples fixed after longer stay out-of-niche acquired numerous changes in gene-expression profile (GEP). The overall genes modulated included a common subset of functionally diverse genes displaying prompt and sustained ‘switch’ in expression irrespective of the tumor type. Interestingly, the ‘switch’ in GEP was reversible and turned ‘off-and-on’ again in culture conditions, resuming cell–cell–matrix contact versus respread into suspension, respectively. Moreover, the resuming of contact prolonged the survival of tumor cells out-of-niche, and the regression of the ‘contactless switch’ was followed by induction of a new set of genes, this time mainly encoding extracellular proteins including angiogenic factors and extracellular matrix proteins. Our data set, being unique in authentic expression design, uncovered niche-modulated and niche-modulating genes capable of controlling homing, expansion and angiogenesis.
Similar content being viewed by others
Introduction
In contrast to immortal cell lines, investigation of the authentic primary counterparts, including malignant bone marrow (BM) cells, is limited by the short survival of the latter ex vivo.1 This obstacle reflects the strict dependence of primary BM cells on their niche, which contributes the space, supplies the nutrients and provides the conditions required for cell–cell interactions and soluble factor exchange.1, 2 Newer investigation tools, for example, molecular profiling of cell populations improved the discrimination between different subtypes of the same tumor and provided prognostic classifications to various hematological malignancies.3, 4, 5, 6, 7, 8, 9
As the loss of niche stimuli is one of the limiting factors in growing primary cells, we searched for the target genes in BM cells that respond to the niche stimuli and did it in the setting of BM malignancy. This selection was made in order to assist in designing new targeted therapy directed at critical niche signals required for growth and survival of the primary tumor cells. For this purpose, we analyzed the changes in gene-expression profile (GEP) following interruption of cell–niche interactions due to BM aspiration. The sustained changes in expression disclosed the subset of genes, which are sensitive to the BM microenvironment. The nature of the microenvironment stimuli regulating the BM cells was further specified using isolated culture conditions.
Materials and methods
Sample collections
BM aspiration was collected into heparinized syringe, and two drops (∼100 μl) were transferred immediately into an Eppendorf tube and flash frozen with liquid nitrogen. Subsequent samples were frozen after longer incubation in the syringe pool (left in room temperature). Alternatively, two drops were transferred into tubes containing 1.2 ml RNAlater (Ambion Diagnostics, Austin, TX, USA).
Patients
Newly diagnosed patients with clinical and laboratory manifestations typical for multiple myeloma (MM) with a high tumor load (severe cytopenias, leukemic phase, extreme paraprotein levels, multiple lytic lesions and hypercalcemia) entered the study. Also, patients with acute leukemia presenting with severe cytopenias (grade 4) and no mature leukocytes seen in the peripheral blood were included. The samples from the leukemia cases were analyzed only if cytospine smears from the aspirates confirmed complete ‘replacement’ (∼100%) by tumor cells. Another case had leukemic relapse of diffuse large B-cell lymphoma (DLBCL). The patients provided written informed consent. The study was approved by Assaf Harofeh Institute, Zerifin, Israel, review boards.
Long-term cultures
Residual BM aspirate was placed in a vertical tube for ⩾30 min, and then the fluid layer above the red blood cell sediment was collected and 300 μl fractions (containing 20–100 × 103 cell per μl) were seeded into 24-well plates (BD Falcon 24-well Multiwell Plate, Bactlab Diagnostics, Caesarea, Israel). After >15 min incubation (room temperature, Bactlab Diagnostics, Caesarea, Israel), the cultures were supplemented with 1 ml RPMI1640 containing 2 mM L-glutamine (Sigma, Rehovot, Israel), 50 U/ml penicillin, 50 μg/ml streptomycin (Sigma) and 20% (v/v) heat-inactivated fetal bovine serum (Biological Industries, Kibbutz Beit Haemek, Israel). Cells were maintained at 37 °C in a humidified 5% CO2 incubator. Cultures were fed biweekly by replacing 50% of the medium with fresh supplement. The proportion of tumor cells was determined morphologically using cytospine smears. For RNA extraction, most of the medium was removed (leaving only 200–300 μl), and after vigorous pippetation, the released cells were collected into an Eppendorf tube. Fractions of ∼70 μl were flash frozen at different time points, starting immediately following release. The cells were reincubated between freezing procedures.
RNA extraction:
Frozen samples were lyzed by adding 300 μl lysis buffer to tubes. Samples stored in RNAlater were fractionated and lyzed by adding 10 × volume (v/v) lysis buffer. RNA was isolated by MagNA Pure Compact RNA Isolation procedure using MagNA Pure Compact instrument (Roche). Integrity of RNA was examined by Agilent 2100 Bioanalyzer (Santa Clara, CA, USA).
Gene-expression profile
Biotin-labeled cRNA was generated from 200 ng total RNA, hybridized onto GeneChip Human Gene1.0 ST Array (Affymetrix, Santa Clara, CA,USA) and the data were processed with the Affymetrix GeneChip Scanner 3000 and Affymetrix Expression Console. Normalization was done by the Robust Multi-array Average (RMA) method, and fold-change results were calculated relative to the reference sample of each case (which was fixed without delay). The entire GEP data is accessible from NCBI Gene Expression Omnibus GSE36036 and GSE39184. http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE36036. http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE39184.
Quantitative real-time PCR
cDNA was obtained by Transcriptor First strand cDNA synthesis kit (Roche Diagnostics, Mannheim, Germany), according to the manufacturer’s instruction. Quantitative real-time PCR with various primers and LightCycler 480 SYBR Green I Master kit (Roche Diagnostics) was done using LightCycler 480 Instrument II (Roche Diagnostics) according to the manufacturer’s instructions. PCR primers and thermal cycling parameters were designed using Primer3 software (Supplementary Table 1). Standard curves were generated using 0.2, 0.5, 1, 2 and 5 ng/μl of control GAPDH cDNA.
Results
BM aspirations were obtained from an overall group of 27 patients with MM, 14 patients with acute leukemia and one case with leukemic phase of DLBCL. The aspirates remained in their syringes, and on each time point, a fraction was transferred into an empty tube and fixed (either by flash freezing or using RNAlater). Therefore, each case had its own reference sample fixed immediately following aspiration and many additional samples (6–40 samples) whose fixation was delayed (up to 24 h). GEP was analyzed from whole BM samples of the 10 MM cases, with the highest proportion of plasma cells (60–95%) and the best quality of purified RNA (RIN ⩾6). The latter group of patients presented typical characteristics of symptomatic myeloma in terms of age distribution, gender, clinical and laboratory abnormalities (Table 1a). The relatively high proportion of tumor cells, despite skipping the usual cell-separation procedures, which would prevent the immediate fixation required to record the authentic ‘in vivo’ expression, was the consequence of strict patient selection and using of the initial aspiration fraction while the majority of residual normal cells were granulocytes. GEP was also analyzed from whole BM samples of four acute myeloid leukemia (AML) cases for whom the proportion of tumor cells was nearly 100% by morphology and from a case with leukemic relapse of testicular DLBC with 70% tumor cells (Table 1b).
Prompt ‘switch’ in GEP
The first participant had MM (case A), and the BM samples selected for microarray analysis were those frozen immediately following aspiration, at 17 min later and following 120 min spent in the syringe pool. The comparison of the three GEPs accepted revealed prompt induction of numerous genes after 17 min ex vivo, which included upregulation of 363 genes >2-folds, 34 genes >3-folds, 10 genes >4-folds, 7 genes >5-folds and 3 genes >10-folds compared with the reference sample frozen immediately following aspiration. Moreover, after 120 min ex vivo, nearly two-third of the former upregulated genes (at 17 min) remained overexpressed and many of which displayed further increase in expression signals. Surprisingly, a very similar ‘switch’ signature was recorded from the aspiration samples of an AML patient (case A′) after 20 min and 120 min spent out-of-niche compared with his reference GEP (Table 2). In both patients, downregulation was milder and involved fewer genes.
To trace the earliest onset of gene modulation and view the dynamic of changes, we used quantitative real-time PCR analysis, directed at mRNA levels of three inducible genes. The results revealed an almost immediate induction onset with progressive increase in transcript levels during the first 120 min ex vivo (Table 3).
The ‘switch’ in GEP is sustained
The annotation database revealed that many of the promptly induced genes in the aspirates from the first two cases examined are classified as ‘immediate early genes’ (FOS, FOSB, RGS1, DUSP1, JUN, CD69, EGR1, PTGS2 and NR4A2). This class of genes is characterized by rapid induction onset upon stimulation but transient expression pattern.10 To follow the expression of these and related genes at more distant time intervals following aspiration, we analyzed BM samples from additional cases and extended the time-to-fixation delays. It was found that the ‘switch’ in expression of the above genes is still recognizable even after 11 h ex vivo (Figure 1). Furthermore, the majority of the early GEP changes (⩾2-folds) recorded during the first 120 min ex vivo persisted after 260 min (case C′), 420 min (case F), 480 min (case G) and 540 min (case B) spent in the syringe pool, and, frequently, the changes continued to increase (Figure 2). The ‘switch’ signature was also ‘fixed’ in a sense that no late-onset induction or downregulation of genes was evident during all our records, including caspases or other proapoptotic genes expected to be rising. Finally, the tumor type had no visible landmarks on the ‘switch’ signature, and the top modulated genes were similar among different individuals whether diagnosed with MM, AML or DLBCL.
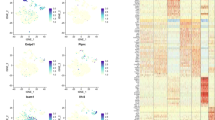
Sustained ‘switch’ in GEP following BM aspiration. BM samples from 10 patients with MM, 4 with AML and 1 case with DLBCL (BCL) were fractionated and fixed (either by flash freezing or using RNAlater) at different time points following aspiration. The samples were analyzed by microarray, and the common changes in gene expression (⩾2-folds) are displayed as fold changes relative to the signals recorded from the corresponding reference samples (fixed immediately following aspiration).
Induction of AREG. The signal intensities are depicted against the time to RNA fixation following BM aspirations obtained from patients with MM (cases A–J), leukemic phase of DLBCL (case B′) and AML (cases A′, C′–E′) or alternatively following mobilization from the bottom well into suspension of cultured BM cells from cases B and C (see details in the next section).
The ‘switch’ signature is reversible and contact dependent
In theory, the ‘switch’ in GEP could be secondary to any of the following insults: (1) Stress of aspiration and cell transfer to ambient conditions. (2) Loss of regulatory cues from the BM niche proper. (3) Loss of external inputs from habitual cell–cell interactions among adjacent BM cells due to cell spread, dilution with red blood cells and absence of extracellular matrix to adhere. To search for the real cause, we turned to isolated culture conditions. Our culture model was designed to enable recovery of the usual cell–cell contact relationships via seeding of intact BM samples (excluding red blood cells) on a matrix and adding the medium only after adherence of the whole sample in order to avoid cell dilution. In our experience, such conditions can delay apoptosis of primary MM cells for variable periods of time, although cell proliferation remains limited (as evident from the failure of cellular areas on the bottom well to fill adjacent empty areas).11, 12 Moreover, owing to the short life span of normal granulocytes, the purity of the MM cell populations increased over culture days and, eventually, the cultures became homotypic, excluding scattered macrophages (Supplementary Figure 1). As anticipated, the prolonged viability of the MM cell populations in our culture model was associated with near-complete regression of the changes in GEP, which evolved following aspiration. Specifically, in all analyzed cases: J, B and C, where their aspiration samples were cultured for 2, 7 and 29 days, respectively, the ‘switch’ in GEP fade over culture days except for IL8, which remained overexpressed and its signals even increased further (by 61.6-, 7.6- and 3.9-folds, respectively). Moreover, the ‘switch’ signature recurred after leaving the cultured cells in suspension for additional 60 min (case B) or 120 min (case C), following their release from the bottom well compared with the cells that were frozen immediately following release (Figure 3), and the mobilized cells died within 24 h. These observations indicate that the ‘switch’ phenomenon is reversible and basically contact dependent.
The ‘switch’ in GEP is contact dependent. Cultured BM cells (CC) were flash frozen at different time points following their mobilization from the bottom well into suspension. Pleural fluid (PF) cell sediment containing pure MM cell population was split with one part left in the bottom tube and the second was transferred into another tube and maintained in suspension after redilution with pleural fluid. Both parts of the sediment were flash frozen at 270 min after the split. The frozen samples were analyzed by microarray, and the differences in signal intensities are displayed as fold changes relative to the GEPs recorded from the reference samples (fixed immediately following mobilization into suspension). Unannotated genes are denoted by their Affymetrix probe set ID number. The results showed similar changes in GEP following cell mobilization and maintenance in suspension as evolved following BM aspiration, indicating that the switch in GEP is related to the loss of cell–cell–matrix contact relationships.
Another source of pure MM cell population was malignant pleural effusion that developed in case C. The pleural fluid obtained formed thick sediment on the bottom tube, which by morphology contained only sheets of MM cells without normal cells. On that occasion, the GEP comparison was between sediment cells that were flash frozen after respreading and maintenance in suspension for additional 270 min versus intact sediment cells flash frozen without any spread. Once again, the spread of the crowding tumor cells into suspension reinduced their ‘contactless’ signature, which already started to regress when the cells were allowed to cluster together on the bottom tube (Figure 3).
‘Contactless switch’ during remission
To clarify if the ‘contactless switch’ is tumor related or actually represents a more generalized phenomenon, we repeated the GEP comparison during remission. The patient selected (case B) entered complete remission following treatment with bortezomib and dexamethasone (with drop in her monoclonal protein from 6.41 g/dl to undetectable levels). It was found that the switch in GEP recorded before treatment recurred following aspiration during remission, although many of the modulated genes were discordant. Consequently, the overall GEP changes could be categorized into three different groups: (1) Genes that were modulated during remission only; in this group, the top modulated gene was CXCL12 (data not shown) which has a major role in homing of myeloid progenitors. (2) Genes modulated in pretreatment samples only (Figure 4, lower field); in this group, the top cited gene in the context of MM was the homing receptor CXCR4.13 (3) Concordant genes which were modulated in both disease stages. The latter category included 22 genes, with many of which being recognizable as immediate early genes (Figure 4, upper field).
‘Switch’ in GEP during remission versus before treatment. A comparison between the top post-aspiration changes in GEP (⩾2-folds) evolved in the BM samples from case B during complete remission (CR) (right column) versus the changes recorded before treatment (MM). The results are displayed as fold changes relative to corresponding reference samples (fixed immediately following aspiration). The lower field demonstrates the genes modulated in the pretreatment samples only. The upper field demonstrates overlapping genes modulated in both disease stages. Unannotated genes are denoted by their Affymetrix probe set ID number.
Unannotated niche-modulated genes
In addition to known genes, the ‘contactless’ signature consisted of many nameless and unannotated genes (Figure 1). The prominent member in this category was MOP-1, showing remarkable induction in all ‘contactless’ situations. Other genes (that is, Affymetrix probe sets 8017096 and 8015796) were only induced in BM samples that were largely ‘replaced’ by tumor cells. The top unannotated genes induced in the aspirates from the 10 MM cases are presented on Supplementary Table 2. This list likely includes newer immediate early genes waiting for validation.
Niche-modulating genes
Interestingly, the regression of the ‘contactless’ switch during culture was followed by induction of a new set of genes, this time mostly encoding extracellular proteins (Figure 5); among which were angiogenic factors (IL8 and CXCL5) and extracellular matrix proteins (SPP1 and FN1), as well as additional chemokines and growth factors (CXCL5, CCL2, CCL20 and IL6). This adaptive response uncovered inducible genes becoming overexpressed permanently, presumably due to failure of the tumor cells to recruit the stimulated niche components (for example, vascular vessels and extracellular matrix scaffold). Some of the above niche-modulating genes can be considered therefore attractive therapeutic targets, especially SPP1 (or its product osteopontin) and IL8 because of their strongest induction. Likewise, inhibitors to niche-modulated gene products, such as amphiregulin (encoded by AREG) and COX-2 (encoded by PTGS2), may have a role in reducing the tumor expansion in certain BM malignancies. Conversely, suppression of niche-modulated genes like JUN, EGR1 and KLF4 may improve the culture success.
Niche-modulating genes. The top induced genes in cultured BM samples from three patients with MM are listed . The changes were recorded after culture for 1, 7 and 29 days and are presented as fold changes (green) relative to the signal intensities recorded from corresponding fresh BM samples flash frozen immediately following aspiration. The genes induced during culture were totally different from the ‘contact-modulated genes’ except for IL8, which was also induced following aspiration.
Discussion
This study exhibited pitfalls in gene-expression profiling when RNA fixation is not executed immediately following interruption of homeostasis. In addition, the changes in GEP recorded following aspiration from the whole BM samples of a heterogeneous group of patients with diverse hematological malignancies, and disease stages were integrated into a uniform ‘switch’ signature that was recognizable as early as few minutes following aspiration, persisted for many hours and regressed only after cell–cell–matrix contact was resumed. The latter observation indicates that cell–cell contact per se controls the expression of numerous genes in BM cells, while interruption of the contact relationships relieves cell–to–cell regulation and permits full expression of the ‘default’ genetic program inherent to each individual cell.
The resuming of contact that suppressed the ‘contactless switch’ signature contributed in our culture model to increased viability of the tumor cells. Such protective effect may enhance tumor cell crowding in niche, and the contact may also serve to distribute vital signals from unreachable niche components like osteoclasts, vascular vessels and stromal cells, thereby overcoming the limited access to these regulators. The survival advantage for tumor cells gained by the overwhelming cell density was balanced, however, by stasis and lack of any expansion features, suggesting contact inhibition of cell cycle combined. Indeed, tumor expansion is more likely favored by the ‘contactless’ genetic program with its overexpressed oncogenes (FOS, JUN), homing genes (RGS1, CD69), angiogenic genes (IL8, AREG) and growth-promoting genes (PTGS2), whereas suppression of these genes presumably stimulates cell migration and inhibits proliferation. Nevertheless, the real physiological set up in which switch to the ‘contactless’ signature is allowed remains obscured. One possible scenario is that the dual function of contact, which delays apoptosis of crowding tumor cells but also inhibits their proliferation combined together with contact suppression of homing genes, ultimately, directs cell spreading and tumor expansion toward less populated niche regions where the ‘contactless’ gene signature is supposed to be less suppressed.
In data analysis of the niche-modulated genes based on the Ingenuity software (Redwood, CA, USA), the top function annotation category was “cancer” (25 molecules) followed by “cell death” (23 molecules) and “proliferation of cells” (20 molecules) (Supplementary Table 3). However, “proliferation of vascular tissue” yielded only one molecule (PTGS2), whereas our literature search yielded six molecules. Further analysis of the data from a niche-regulation perspective reasoned classification of the niche-modulated genes (Figure 1) into four functional categories:
Homing
Both RGS1 and CD69 regulate leukocyte retention in niche. RGS1 attenuates migration and retains cells in niche by regulation of G-protein signaling and reduction of B-cell response to chemokines such as CXCL12.14 Overexpression of CD69 retains immature B-cells in the BM and reduces their number in blood.15
Angiogenesis and stromagenesis
Several niche-modulated genes, for example, AREG, PTGS2, RGS1, RGS2, NR4A2 and IL8 support angiogenesis and recruitment of stromal cells.16, 17, 18, 19, 20, 21, 22, 23, 24
Expansion of BM cells
Amphiregulin (encoded by AREG) also supports tumor cells directly and promotes MM cell growth.25, 26 The upregulation of FOS, JUN, NR4A2, TRIB1 and CCL3 and the downregulation of MIR15A can also enhance leukocyte expansion and tumorigenesis.27, 28, 29, 30, 31, 32
Suppression of BM cells
As pointed above, the reduced viability of primary tumor cells out-of-niche was not preceded during all our records by induction of any specific proapoptotic gene like caspase. Still, constitutive overexpression of master regulatory genes like JUN, tumor-suppressor genes like EGR1 and KLF4 and less-defined genes like FOSB, CD69, PPP1R15A, TRIB1 and NFKBIA can be lethal by itself,10, 33, 34, 35, 36, 37, 38, 39 unless contact is resumed on time before translation/protein activation.
In reviewing the literature, we were unable to find any other series with the same GEP design. Still, the study by Chen et al.36 who observed downregulation of EGR1 and JUN in coculture of purified MM cells with osteoclasts compared with culture without osteoclasts is consistent with our data concerning the downregulation of both genes when contact is resumed. Irrespective of this, Chen et al.36 also found correlation between the outcome of MM patients to JUN and EGR1 expression by their tumor cells. The latter observation, however, did not consider the timing of RNA fixation, which in our series strongly influenced JUN and EGR1 expression. Furthermore, in most published GEP data sets analyzing purified MM or AML cells, the expression of the inducible contact-modulated genes was generally high as opposed to the ‘silence’ of these genes when recorded during homeostasis (immediately following aspiration). For example, in GDS531,40 the signal intensities recorded from FOSB, FOS, DUSP1, NR4A2, EGR1 and RGS1 approached the levels of GAPDH (Supplementary Table 4). Therefore, expression analysis may need some standardization in the future.
In conclusion, despite the limited number of cases analyzed in the present study, the serial methodology used and the reproducibility of results substantiated our contact regulation model. The major questions left are: (1) Who is the sensor of contact? and (2) How is the message transmitted and how does it regulate transcription of so many genes simultaneously?
References
Yaccoby S, Barlogie B, Epstein J . Primary myeloma cells growing in SCID-hu mice: a model for studying the Biology and treatment of myeloma and its manifestations. Blood 1998; 92: 2908–2913.
Nefedova Y, Landowski TH, Dalton WS . Bone marrow stromal-derived soluble factors and direct cell contact contribute to de novo drug resistance of myeloma cells by distinct mechanisms. Leukemia 2003; 17: 1175–1182.
Kohlmann A, Bullinger L, Thiede C, Schaich M, Schnittger S, Döhner K et al. Gene expression profiling in AML with normal karyotype can predict mutations for molecular markers and allows novel insights into perturbed biological pathways. Leukemia 2010; 24: 1216–1220.
Mahtouk K, Hose D, De Vos J, Moreaux J, Jourdan M, Rossi JF et al. Input of DNA microarrays to identify novel mechanisms in multiple myeloma biology and therapeutic applications. Clin Cancer Res 2007; 13: 7289–7295.
Bergsagel PL, Kuehl WM, Zhan F . Cyclin D dysregulation: an early and unifying pathogenic event in multiple myeloma. Blood 2005; 106: 296–303.
Zhan F, Huang Y, Colla S . The molecular classification of multiple myeloma. Blood 2006; 108: 2020–2028.
Zhan F, Hardin J, Kordsmeier B . Global gene expression profiling of multiple myeloma, monoclonal gammopathy of undetermined significance, and normal bone marrow plasma cells. Blood 2002; 99: 1745–1757.
Munshi NC, Hideshima T, Carrasco D . Identification of genes modulated in multiple myeloma using genetically identical twin samples. Blood 2004; 103: 1799–1806.
Mahtouk K, Hose D, Raynaud P . Heparanase influences expression and shedding of syndecan-1, and its expression by the bone marrow environment is a bad prognostic factor in multiple myeloma. Blood 2007; 109: 4914–4923.
Shan Y, Carlock LR, Walker PD . NMDA receptor overstimulation triggers a prolonged wave of immediate early gene expression: relationship to excitotoxicity. Exp Neurol 1997; 144: 406–415.
Cohen Y, Hardan Y, Nagler A, Zipori D Establishment of an in vitro culture method for multiple myeloma cells Blood (ASH Ann Meet Abs) 2004, Abstract no. 4878.
Cohen y, Zipori D Extremely peculiar nuclear changes in cultured primary myeloma cells, including bulging and duplication/multiplication of nuclei and cell expansion without apparent mitotic forms Blood (ASH Annu Meet Abs) 2004, Abstract no. 4879.
Nie Y, Waite J, Brewer F, Sunshine MJ, Littman DR, Zou YR . The role of CXCR4 in maintaining peripheral B cell compartments and humoral immunity. J Exp Med 2004; 200: 1145–1156.
Han SB, Moratz C, Huang NN, Kelsall B, Cho H, Shi CS et al. Rgs1 and Gnai2 regulate the entrance of B lymphocytes into lymph nodes and B cell motility within lymph node follicles. Immunity 2005; 22: 343–354.
Allende ML, Tuymetova G, Lee BG, Bonifacino E, Wu YP, Proia RL . S1P1 receptor directs the release of immature B cells from bone marrow into blood. J Exp Med 2010; 207: 1113–1124.
Mahtouk K, Hose D, Rème T, De Vos J, Jourdan M, Moreaux J et al. Expression of EGF-family receptors and amphiregulin in multiple myeloma. Amphiregulin is a growth factor for myeloma cells. Oncogene 2005; 24: 3512–3524.
Müller-Decker K, Fürstenberger G . The cyclooxygenase-2-mediated prostaglandin signaling is causally related to epithelial carcinogenesis. Mol Carcinog 2007; 46: 705–710.
Pai R, Soreghan B, Szabo IL, Pavelka M, Baatar D, Tarnawski AS . Prostaglandin E2 transactivates EGF receptor: a novel mechanism for promoting colon cancer growth and gastrointestinal hypertrophy. Nat Med 2002; 8: 289–293.
Aherne CM, McMorrow J, Kane D, FitzGerald O, Mix KS, Murphy EP . Identification of NR4A2 as a transcriptional activator of IL-8 expression in human inflammatory arthritis. Mol Immunol 2009; 46: 3345–3357.
Mizukami Y, Jo WS, Duerr EM, Gala M, Li J, Zhang X et al. Induction of interleukin-8 preserves the angiogenic response in HIF-1 deficient colon cancer cells. Nat Med 2005; 11: 992–997.
Koch AE, Polverini PJ, Kunkel SL, Harlow LA, DiPietro LA, Elner VM et al. Interleukin-8 as a macrophage-derived mediator of angiogenesis. Science 1992; 258: 1798–1801.
Zhao D, Desai S, Zeng H . VEGF stimulates PKD-mediated CREB-dependent orphan nuclear receptor Nurr1 expression: role in VEGF-induced angiogenesis. Int J Cancer 2011; 128: 2602–2612.
Rangel J, Nosrati M, Leong SP, Haqq C, Miller JR, Sagebiel RW et al. Novel role for RGS1 in melanoma progression. Am J Surg Pathol 2008; 32: 1207–1712.
Boelte KC, Gordy LE, Joyce S, Thompson MA, Yang L, Lin PC . Rgs2 mediates pro-angiogenic function of myeloid derived suppressor cells in the tumor microenvironment via upregulation of MCP-1. PLoS One 2011; 6: e18534.
Mahtouk K, Cremer FW, Rème T, Jourdan M, Baudard M, Moreaux J et al. Heparan sulphate proteoglycans are essential for the myeloma cell growth activity of EGF-family ligands in multiple myeloma. Oncogene. 2006; 25: 7180–7191.
Willmarth NE, Ethier SP . Amphiregulin as a novel target for breast cancer therapy. J Mammary Gland Biol Neoplasia 2008; 13: 171–179.
Behrens A, Sibilia M, David JP, Möhle-Steinlein U, Tronche F, Schütz G et al. Impaired postnatal hepatocyte proliferation and liver regeneration in mice lacking c-jun in the liver. EMBO J 2002; 21: 1782–1790.
Li X, Tai HH . Activation of thromboxane A(2) receptors induces orphan nuclear receptor Nurr1 expression and stimulates cell proliferation in human lung cancer cells. Carcinogenesis 2009; 30: 1606–1613.
Yokoyama T, Nakamura T . Tribbles in disease: Signaling pathways important for cellular function and neoplastic transformation. Cancer Sci 2011; 102: 1115–1522.
Gilby DC, Sung HY, Winship PR, Goodeve AC, Reilly JT, Kiss-Toth E . Trib1 and Evi1 cooperate with Hoxa and Meis1 in myeloid leukemogenesis. Blood 2007; 109: 3998–4005.
Dussmann P, Pagel JI, Vogel S, Magnusson T, Zimmermann R, Wagner E et al. Live in vivo imaging of Egr-1 promoter activity during neonatal development, liver regeneration and wound healing. BMC Dev Biol 2011; 20: 28.
Cimmino A, Calin GA, Fabbri M, Iorio MV, Ferracin M, Shimizu M et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA 2005; 102: 13944–13949.
Preston GA, Lyon TT, Yin Y, Lang JE, Solomon G, Annab L et al. Induction of apoptosis by c-Fos protein. Mol Cell Biol 1996; 16: 211–218.
Baumann S, Hess J, Eichhorst ST, Krueger A, Angel P, Krammer PH et al. An unexpected role for FosB in activation-induced cell death of T cells. Oncogene 2003; 22: 1333–1339.
Podar K, Raab MS, Tonon G, Sattler M, Barilà D, Zhang J et al. Up-regulation of c-Jun inhibits proliferation and induces apoptosis via caspase-triggered c-Abl cleavage in human multiple myeloma. Cancer Res 2007; 67: 1680–1688.
Chen L, Wang S, Zhou Y, Wu X, Entin I, Epstein J et al. Identification of early growth response protein 1 (EGR-1) as a novel target for JUN-induced apoptosis in multiple myeloma. Blood 2010; 115: 61–70.
Joslin JM, Fernald AA, Tennant TR, Davis EM, Kogan SC, Anastasi J et al. Haploinsufficiency of EGR1, a candidate gene in the del(5q), leads to the development of myeloid disorders. Blood 2007; 110: 719–726.
Kharas MG, Yusuf I, Scarfone VM, Yang VW, Segre JA, Huettner CS . KLF4 suppresses transformation of pre-B cells by ABL oncogene. Blood 2007; 109: 747–755.
Gilby DC, Sung HY, Winship PR, Goodeve AC, Reilly JT, Kiss-Toth E . Tribbles-1 and -2 are tumour suppressors, down-regulated in human acute myeloid leukaemia. Immunol Lett 2010; 130: 115–124.
Tian E, Zhan F, Walker R, Rasmussen E, Ma Y, Barlogie B et al. The role of the Wnt-signaling antagonist DKK1 in the development of osteolytic lesions in multiple myeloma. N Engl J Med 2003; 349: 2483–2494.
Acknowledgements
We thank Professor Dov Zipory for the training and teaching of the first author in the Weitzmann Institute of Science, Dr Izhar Hardan from Meir Medical Center for his kind technical support, Dr Relly Forer, Dr Inna Vulin and Dr Dina Volodarsky from Dyn diagnostics Ltd for microarray and Bioinformatic service, Miri Rotchild for flow cytometry, and Dr Naomy Levine, Dr Marina Izak, Dr Katerine Hertzog, Dr Irena Esterlis and Dr Odit Gutwein their for assistance in patient recruitment.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Blood Cancer Journal website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Cohen, Y., Garach-Jehoshua, O., Bar-Chaim, A. et al. Niche-modulated and niche-modulating genes in bone marrow cells. Blood Cancer Journal 2, e97 (2012). https://doi.org/10.1038/bcj.2012.42
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bcj.2012.42