Article
|
Open Access
Featured
-
-
Article |
 The E3 ligase adapter cereblon targets the C-terminal cyclic imide degron
The E3 ligase adapter cereblon targets the C-terminal cyclic imide degron
C-terminal cyclic imides are physiological degrons that enable the ubiquitin E3 ligase adapter protein cereblon to target substrates for degradation.
- Saki Ichikawa
- , Hope A. Flaxman
- & Christina M. Woo
-
Article |
 Structural insights into Ubr1-mediated N-degron polyubiquitination
Structural insights into Ubr1-mediated N-degron polyubiquitination
Structures of Ubr1 in complex with Ubc2, ubiquitin and two N-degron peptides reveal a Ubc2-binding region and an acceptor ubiquitin-binding loop on Ubr1, providing mechanistic insights into the initiation and elongation steps of ubiquitination catalysed by Ubr1.
- Man Pan
- , Qingyun Zheng
- & Minglei Zhao
-
Article |
 CRL4AMBRA1 is a master regulator of D-type cyclins
CRL4AMBRA1 is a master regulator of D-type cyclins
Biochemical and genetics studies identify CRL4AMBRA1 as the ubiquitin ligase that has a key role in regulating the stability of D-type cyclins during cell-cycle progression.
- Daniele Simoneschi
- , Gergely Rona
- & Michele Pagano
-
Article |
 The AMBRA1 E3 ligase adaptor regulates the stability of cyclin D
The AMBRA1 E3 ligase adaptor regulates the stability of cyclin D
AMBRA1 is the main regulator of the degradation of D-type cyclins, and loss of AMBRA1 promotes cell proliferation and tumour growth, and reduces the sensitivity of cancer cells to inhibition of CDK4 and CDK6.
- Andrea C. Chaikovsky
- , Chuan Li
- & Julien Sage
-
Article |
 Small-molecule-induced polymerization triggers degradation of BCL6
Small-molecule-induced polymerization triggers degradation of BCL6
Binding of the small molecule BI-3802 to the oncogenic transcription factor B cell lymphoma 6 (BCL6) induces polymerization of BCL6, leading to its ubiquitination by SIAH1 and proteasomal degradation.
- Mikołaj Słabicki
- , Hojong Yoon
- & Benjamin L. Ebert
-
Article |
 NEDD8 nucleates a multivalent cullin–RING–UBE2D ubiquitin ligation assembly
NEDD8 nucleates a multivalent cullin–RING–UBE2D ubiquitin ligation assembly
A cryo-electron microscopy structure provides insights into the activation of cullin–RING E3 ligases by NEDD8 and the consequent catalysis of ubiquitylation reactions.
- Kheewoong Baek
- , David T. Krist
- & Brenda A. Schulman
-
Article |
 Structural plasticity of D3–D14 ubiquitin ligase in strigolactone signalling
Structural plasticity of D3–D14 ubiquitin ligase in strigolactone signalling
The plant F-box protein D3 has a C-terminal α-helix that switches between two conformational states, allowing the α/β hydrolase D14 to recruit the transcription repressor D53 for strigolactone-dependent degradation.
- Nitzan Shabek
- , Fabrizio Ticchiarelli
- & Ning Zheng
-
Letter |
 Distinct proteostasis circuits cooperate in nuclear and cytoplasmic protein quality control
Distinct proteostasis circuits cooperate in nuclear and cytoplasmic protein quality control
Ubiquitin chains linked to cytoplasmic misfolded proteins are different from those linked to nuclear misfolded proteins, each requiring a distinct combination of molecular chaperones and ubiquitination circuitries.
- Rahul S. Samant
- , Christine M. Livingston
- & Judith Frydman
-
Article |
 Lenalidomide induces ubiquitination and degradation of CK1α in del(5q) MDS
Lenalidomide induces ubiquitination and degradation of CK1α in del(5q) MDS
Lenalidomide, a derivative of thalidomide, is an effective drug for myelodysplastic syndrome; lenalidomide binds the CRL4CRBN E3 ubiquitin ligase and promotes degradation of casein kinase 1a, on which the malignant cells rely for survival.
- Jan Krönke
- , Emma C. Fink
- & Benjamin L. Ebert
-
Letter |
 Synergistic blockade of mitotic exit by two chemical inhibitors of the APC/C
Synergistic blockade of mitotic exit by two chemical inhibitors of the APC/C
Simultaneous disruption of two different protein–protein interactions within the (APC/C–Cdc20)–substrate complex can synergistically inhibit APC/C-dependent proteolysis and mitotic exit.
- Katharine L. Sackton
- , Nevena Dimova
- & Randall W. King
-
Letter |
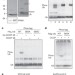 Ubiquitin is phosphorylated by PINK1 to activate parkin
Ubiquitin is phosphorylated by PINK1 to activate parkin
Ubiquitin, known for its role in post-translational modification of other proteins, undergoes post-translational modification itself; after a decrease in mitochondrial membrane potential, the kinase enzyme PINK1 phosphorylates ubiquitin at Ser 65, and the phosphorylated ubiquitin then interacts with ubiquitin ligase (E3) enzyme parkin, which is also phosphorylated by PINK1, and this process is sufficient for full activation of parkin enzymatic activity.
- Fumika Koyano
- , Kei Okatsu
- & Noriyuki Matsuda
-
Letter |
 BTB-ZF factors recruit the E3 ligase cullin 3 to regulate lymphoid effector programs
BTB-ZF factors recruit the E3 ligase cullin 3 to regulate lymphoid effector programs
The E3 ubiquitin ligase cullin 3 is shown to bind BTB-zinc finger transcription factors to direct the ubiquitination of nuclear chromatin-associated factors to control transcription and cell-fate decisions in B- and T-cell populations.
- Rebecca Mathew
- , Michael P. Seiler
- & Albert Bendelac
-
Article |
 Structure of a RING E3 ligase and ubiquitin-loaded E2 primed for catalysis
Structure of a RING E3 ligase and ubiquitin-loaded E2 primed for catalysis
This study presents the crystal structure of a RING-type E3 ligase bound to ubiquitin-loaded E2; the structure reveals how ubiquitin binding to E2 leads to changes in the catalytic site, priming it for catalysis by the E3 enzyme.
- Anna Plechanovová
- , Ellis G. Jaffray
- & Ronald T. Hay
-
Letter |
 GlcNAcylation of histone H2B facilitates its monoubiquitination
GlcNAcylation of histone H2B facilitates its monoubiquitination
Genome-wide analysis shows that H2B S112 O-linked to N-acetylglucosamine is frequently located near transcribed genes, suggesting that histone GlcNAcylation facilitates transcription of the genes.
- Ryoji Fujiki
- , Waka Hashiba
- & Shigeaki Kato
-
Letter |
 Control of Drosophila endocycles by E2F and CRL4CDT2
Control of Drosophila endocycles by E2F and CRL4CDT2
- Norman Zielke
- , Kerry J. Kim
- & Bruce A. Edgar
-
Letter |
 UBCH7 reactivity profile reveals parkin and HHARI to be RING/HECT hybrids
UBCH7 reactivity profile reveals parkin and HHARI to be RING/HECT hybrids
- Dawn M. Wenzel
- , Alexei Lissounov
- & Rachel E. Klevit

 Stress response silencing by an E3 ligase mutated in neurodegeneration
Stress response silencing by an E3 ligase mutated in neurodegeneration