Featured
-
-
Article |
 Single yeast cells vary in transcription activity not in delay time after a metabolic shift
Single yeast cells vary in transcription activity not in delay time after a metabolic shift
Individual cells respond differently to environmental stressors. Here, Schwabe et al.expose yeast cells to sulphur stress and show that small variations in response time combined with a high transient variability in transcript number contribute to stochasticity in response to this stress.
- Anne Schwabe
- & Frank J. Bruggeman
-
Article |
 Contribution of RNA polymerase concentration variation to protein expression noise
Contribution of RNA polymerase concentration variation to protein expression noise
The quantitative relationship between the fluctuation of specific extrinsic and intrinsic factors, and stochastic fluctuations in gene expression - or noise - has not been clearly established. Here, Yang et al.demonstrate that intrinsic noise is independent of - while extrinsic noise scales linearly with - variation in RNA polymerase abundance.
- Sora Yang
- , Seunghyeon Kim
- & Nam Ki Lee
-
Article |
 Telomerase stimulates ribosomal DNA transcription under hyperproliferative conditions
Telomerase stimulates ribosomal DNA transcription under hyperproliferative conditions
Several recent studies suggest that telomerase has key physiologic functions beyond its well-known role telomere maintenance. Here, Garcia Gonzalez et al. implicate telomerase in the regulation of ribosomal DNA transcription by RNA polymerase I.
- Omar Garcia Gonzalez
- , Robin Assfalg
- & Sebastian Iben
-
Article |
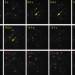 Single-molecule analysis of transcription factor binding at transcription sites in live cells
Single-molecule analysis of transcription factor binding at transcription sites in live cells
Despite numerous previous studies, the nature of the interaction of transcription factors (TF) with their endogenous response elements (REs) has remained unclear. Here the authors characterize the binding of p53 and glucocorticoid receptor to their endogenous REs and find that transcriptionally productive interactions are transient and involve only a small fraction of cellular TF molecules.
- Tatsuya Morisaki
- , Waltraud G. Müller
- & James G. McNally
-
Article |
 Conserved architecture of the core RNA polymerase II initiation complex
Conserved architecture of the core RNA polymerase II initiation complex
It was suggested that despite the conservation of their components, yeast and human pol II initiation complexes diverged in architecture. Mühlbacher et al.now demonstrate that the yeast and human core complexes are structurally conserved and provide insight into the conformations adopted by TFIIF during initiation.
- Wolfgang Mühlbacher
- , Sarah Sainsbury
- & Patrick Cramer
-
Article |
 DSIF and NELF interact with Integrator to specify the correct post-transcriptional fate of snRNA genes
DSIF and NELF interact with Integrator to specify the correct post-transcriptional fate of snRNA genes
The elongation factors DSIF and NELF have established roles in polymerase pausing, elongation and 3'-end processing of replication-dependent histone mRNAs. Here the authors demonstrate that DSIF and NELF form a complex with Integrator and allow proper 3'-processing of snRNA transcripts by preventing the recruitment of CstF.
- Junichi Yamamoto
- , Yuri Hagiwara
- & Yuki Yamaguchi
-
Article |
 The palindromic DNA-bound USP/EcR nuclear receptor adopts an asymmetric organization with allosteric domain positioning
The palindromic DNA-bound USP/EcR nuclear receptor adopts an asymmetric organization with allosteric domain positioning
Nuclear receptors use DNA- and ligand-binding to regulate gene expression. Here, Maletta et al. report the first structural description of a full inverted repeat-bound nuclear receptor complex, which shows that the protein structure is asymmetric, despite the symmetry of the bound DNA.
- Massimiliano Maletta
- , Igor Orlov
- & Bruno P. Klaholz
-
Article
| Open Access Cell cycle transition from S-phase to G1 in Caulobacter is mediated by ancestral virulence regulators
Cell cycle transition from S-phase to G1 in Caulobacter is mediated by ancestral virulence regulators
The bacterium Caulobacter crescentus divides asymmetrically to generate a replicative stalk cell and a quiescent swarmer cell. Fumeaux et al. show that MucR zinc-finger transcription factors, which regulate virulence in other species, also control re-entry into quiescence in Caulobacter.
- Coralie Fumeaux
- , Sunish Kumar Radhakrishnan
- & Patrick H. Viollier
-
Article |
 A source of the single-stranded DNA substrate for activation-induced deaminase during somatic hypermutation
A source of the single-stranded DNA substrate for activation-induced deaminase during somatic hypermutation
The process of somatic hypermutation, used by B cells to increase antibody diversity, is catalysed by the activation-induced deaminase (AID), which needs to access single-stranded DNA to mediate its function. Here, the authors propose a mechanism for the generation of single-stranded DNA substrate required for AID activity.
- Xiaohua Wang
- , Manxia Fan
- & Matthew D. Scharff
-
Article |
 Cooperativity and equilibrium with FOXA1 define the androgen receptor transcriptional program
Cooperativity and equilibrium with FOXA1 define the androgen receptor transcriptional program
The pioneer factor FOXA1 contributes to androgen receptor (AR)-dependent gene expression by opening chromatin to facilitate AR binding. Here, Jin et al.show that because FOXA1 promotes AR association to many low-affinity sites, excessive FOXA1 can lead to reduced AR availability for specific sites.
- Hong-Jian Jin
- , Jonathan C. Zhao
- & Jindan Yu
-
Article |
 The Arabidopsis transcription factor bZIP11 activates auxin-mediated transcription by recruiting the histone acetylation machinery
The Arabidopsis transcription factor bZIP11 activates auxin-mediated transcription by recruiting the histone acetylation machinery
Histone acetylation has been proposed to be crucial for the regulation of gene transcription controlled by the plant hormone auxin. Here, the authors show that the bZIP11 transcription factor activates auxin-mediated transcription by recruiting the histone acetylation machinery.
- Christoph Weiste
- & Wolfgang Dröge-Laser
-
Article
| Open Access The tapetal AHL family protein TEK determines nexine formation in the pollen wall
The tapetal AHL family protein TEK determines nexine formation in the pollen wall
The nexine is a conserved layer of the pollen wall in land plants. The authors show that the AHL family protein TRANSPOSABLE ELEMENT SILENCING VIA AT-HOOK (TEK) is necessary for nexine formation in Arabidopsis, acting downstream of the transcription factor ABORTED MICROSPORES (AMS).
- Yue Lou
- , Xiao-Feng Xu
- & Zhong-Nan Yang
-
Article
| Open Access GATA-dependent regulatory switches establish atrioventricular canal specificity during heart development
GATA-dependent regulatory switches establish atrioventricular canal specificity during heart development
The atrioventricular canal partitions the developing vertebrate heart. Here, the authors show that the cardiac transcription factor Gata4 together with histone modification enzymes and localized co-factors binds atrioventricular canal-specific enhancers, thereby repressing gene activity in the cardiac chambers.
- Sonia Stefanovic
- , Phil Barnett
- & Vincent M. Christoffels
-
Article
| Open Access The structure and substrate specificity of human Cdk12/Cyclin K
The structure and substrate specificity of human Cdk12/Cyclin K
Cyclin-dependent kinase 12 (Cdk12) phosphorylates the C-terminal domain (CTD) of RNA polymerase II to regulate transcription. Here, the authors solve the crystal structure of the Cdk12 kinase domain and show that Cdk12 has its highest activity on a CTD substrate that carries a serine 7 phosphorylation.
- Christian A. Bösken
- , Lucas Farnung
- & Matthias Geyer
-
Article
| Open Access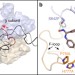 CBR antimicrobials alter coupling between the bridge helix and the β subunit in RNA polymerase
CBR antimicrobials alter coupling between the bridge helix and the β subunit in RNA polymerase
Bacterial RNA polymerase (RNAP) is crucial for cellular gene expression and a validated target for antimicrobial drugs. Here, Malinen et al. explore the effects of the CBR class of RNAP inhibitors on the E. coliRNAP transcription cycle and provide detailed mechanistic insight into their antibacterial action.
- Anssi M. Malinen
- , Monali NandyMazumdar
- & Georgiy A Belogurov
-
Article |
 MED18 interaction with distinct transcription factors regulates multiple plant functions
MED18 interaction with distinct transcription factors regulates multiple plant functions
Arabidopsiscontains multiple mediator proteins that function as transcriptional activators. Here, the authors show that MED18 has roles in plant immunity, control of flowering time and response to hormones, suggesting that mediator proteins control multiple pathways.
- Zhibing Lai
- , Craig M. Schluttenhofer
- & Tesfaye Mengiste
-
Article |
 Darwinian evolution in a translation-coupled RNA replication system within a cell-like compartment
Darwinian evolution in a translation-coupled RNA replication system within a cell-like compartment
Molecular evolution events are vital for the development of cellular complexity. Here the authors construct an evolvable artificial cell model, and observe that Darwinian evolution leads to more efficient RNA replication over time.
- Norikazu Ichihashi
- , Kimihito Usui
- & Tetsuya Yomo
-
Article |
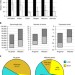 Disparity between microRNA levels and promoter strength is associated with initiation rate and Pol II pausing
Disparity between microRNA levels and promoter strength is associated with initiation rate and Pol II pausing
MicroRNAs are known to be transcribed by RNA polymerase II. The authors show that microRNA promoters driven by TATA-box or NF-κB have increased rates of transcription re-initiation, which leads to local crowding of RNA polymerase II and lower efficiency of microRNA synthesis.
- Nadav Marbach-Bar
- , Amitai Ben-Noon
- & Rivka Dikstein
-
Article
| Open Access The thermodynamic patterns of eukaryotic genes suggest a mechanism for intron–exon recognition
The thermodynamic patterns of eukaryotic genes suggest a mechanism for intron–exon recognition
The thermodynamics of unwinding polynucleotide duplexes can be determined from energy changes for DNA and mRNA interactions. Here the authors show that the ratio between mRNA/DNA and DNA/DNA duplex stability upstream of the 3′- spice sites is a characteristic that can contribute to intron–exon recognition.
- Marina N. Nedelcheva-Veleva
- , Mihail Sarov
- & Stoyno S. Stoynov
-
Article |
 Ribosomal protein S1 functions as a termination factor in RNA synthesis by Qβ phage replicase
Ribosomal protein S1 functions as a termination factor in RNA synthesis by Qβ phage replicase
Protein S1, a subunit of the Qß phage RNA-directed RNA polymerase, was thought to only initiate copying of the phage RNA plus strand. Here, the authors show that S1 stimulates replication of any cognate template by promoting release of the newly synthesized product strand.
- Nikita N. Vasilyev
- , Zarina S. Kutlubaeva
- & Alexander B. Chetverin
-
Article
| Open Access Topoisomerase IIα promotes activation of RNA polymerase I transcription by facilitating pre-initiation complex formation
Topoisomerase IIα promotes activation of RNA polymerase I transcription by facilitating pre-initiation complex formation
Topoisomerases facilitate the progress of elongating polymerases during transcription. Zomerdijk and colleagues now demonstrate an additional role for this enzyme; their data suggest that Top2 can cleave DNA inducing topological changes at the ribosomal DNA promoter, which assists de novoassembly of the RNA polymerase I pre-initiation complex.
- Swagat Ray
- , Tatiana Panova
- & Joost C. B. M. Zomerdijk
-
Article |
 Annotation of microsporidian genomes using transcriptional signals
Annotation of microsporidian genomes using transcriptional signals
Microsporidia are widespread human parasites, but limited genome annotation has hampered efforts to understand their biology. Peyretailladeet al. use sequence motifs upstream of start codons to annotate or re-annotate microsporidian genomes and find new genes potentially involved in interactions with the host.
- Eric Peyretaillade
- , Nicolas Parisot
- & Pierre Peyret
-
Article |
 Protein sliding and DNA denaturation are essential for DNA organization by human mitochondrial transcription factor A
Protein sliding and DNA denaturation are essential for DNA organization by human mitochondrial transcription factor A
The mitochondrial transcription factor A (TFAM) mediates both mitochondrial transcription and DNA compaction, but how it achieves these two functions is unknown. In this study, TFAM is shown to slide along DNA and cause local melting, suggesting a mechanism for how TFAM modulates both transcription and compaction.
- Géraldine Farge
- , Niels Laurens
- & Gijs J.L. Wuite
-
Article |
 Comprehensive interrogation of natural TALE DNA-binding modules and transcriptional repressor domains
Comprehensive interrogation of natural TALE DNA-binding modules and transcriptional repressor domains
The peptide sequence of transcription activator-like effectors (TALEs) can be customized to tailor the binding of TALEs to specific DNA sequences. Conget al. improve TALE specificity for guanine binding and use a genetic construct based on TALEs to efficiently repress expression of a target gene.
- Le Cong
- , Ruhong Zhou
- & Feng Zhang
-
Article
| Open Access Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
Phosphorylation of the carboxy-terminal domain of RNA polymerase II is important for controlling gene transcription. In this study, the transcription elongation factor Tefb is shown to phosphorylate serine-5 and its activity is enhanced when the polymerase is already phosphorylated on serine-7.
- Nadine Czudnochowski
- , Christian A. Bösken
- & Matthias Geyer
-
Article
| Open Access FAD-dependent lysine-specific demethylase-1 regulates cellular energy expenditure
FAD-dependent lysine-specific demethylase-1 regulates cellular energy expenditure
Lysine-specific demethylase 1 (LSD1) removes methyl groups from mono-methylated and dimethylated lysine 4 of histone H3 and represses transcription. In this study, a role for LSD1 in the regulation of genes involved in energy expenditure in adipocytes is reportedin vitroand in mice fed on a high-fat diet.
- Shinjiro Hino
- , Akihisa Sakamoto
- & Mitsuyoshi Nakao
-
Article |
 Multiple exposures to drought 'train' transcriptional responses in Arabidopsis
Multiple exposures to drought 'train' transcriptional responses in Arabidopsis
Whether plants can remember their transcriptional response to stress is unknown. By repeatedly exposingArabidopsisto drought, we show that the plants remember their transcriptional response to stress and that the altered genes retain the epigenetic mark H3K4me3 and stalled phosphorylated polymerase II.
- Yong Ding
- , Michael Fromm
- & Zoya Avramova
-
Article
| Open Access ELL facilitates RNA polymerase II pause site entry and release
ELL facilitates RNA polymerase II pause site entry and release
The super elongation complex, which is involved in transcriptional elongation, contains the Eleven-nineteen Lysine-rich Leukemia protein (ELL). In this study, ELL is shown to stabilize RNA polymerase II prior to recruitment into the super elongation complex, suggesting ELL has a role in early transcription elongation.
- Jung S. Byun
- , Temesgen D. Fufa
- & Kevin Gardner
-
Article
| Open Access IKKβ regulates essential functions of the vascular endothelium through kinase-dependent and -independent pathways
IKKβ regulates essential functions of the vascular endothelium through kinase-dependent and -independent pathways
IKK kinases activate nuclear factor-κB, and the activated form of this transcription factor is found in endothelial cells in diseased tissue. In this study, mice lacking IKKβ in the endothelium are generated, and it is shown that defects in endothelial cell function are both IKK kinase activity dependent and independent.
- Noboru Ashida
- , Sucharita SenBanerjee
- & Anthony Rosenzweig
-
Article
| Open Access Parallel evolution of the make–accumulate–consume strategy in Saccharomyces and Dekkera yeasts
Parallel evolution of the make–accumulate–consume strategy in Saccharomyces and Dekkera yeasts
Saccharomycesyeasts can produce ethanol from sugars in the presence of oxygen. In this study, the authors demonstrate thatDekkera bruxellensis, a distantly related yeast, can also produce and consume ethanol due to the loss of a cis-regulatory element from the promoters of genes crucial for respiration.
- Elżbieta Rozpędowska
- , Linda Hellborg
- & Jure Piškur
-
Article
| Open Access The nuclear orphan receptor Nr4a2 induces Foxp3 and regulates differentiation of CD4+ T cells
The nuclear orphan receptor Nr4a2 induces Foxp3 and regulates differentiation of CD4+ T cells
Regulatory T cells are characterized by the expression of Foxp3, however, how the expression of this protein is controlled is unclear. Here, the authors show that the nuclear orphan receptor, Nr4a2, is a transcriptional activator of Foxp3, and suggest that it is required for the function of regulatory T cells.
- Takashi Sekiya
- , Ikkou Kashiwagi
- & Akihiko Yoshimura
-
Article |
 Wwp2 is essential for palatogenesis mediated by the interaction between Sox9 and mediator subunit 25
Wwp2 is essential for palatogenesis mediated by the interaction between Sox9 and mediator subunit 25
Sox9 is an important transcription factor in the formation of cartilage chondrogenesis that occurs during skeletal development. Nakamuraet al.show that Sox9 interacts with Wwp2 and Med25 to form a complex and that loss of either protein in zebrafish results in altered palate chondrogenesis.
- Yukio Nakamura
- , Koji Yamamoto
- & Haruhiko Akiyama

 Functional characterization of the TERRA transcriptome at damaged telomeres
Functional characterization of the TERRA transcriptome at damaged telomeres