Featured
-
-
Article
| Open Access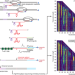 Observation of coordinated RNA folding events by systematic cotranscriptional RNA structure probing
Observation of coordinated RNA folding events by systematic cotranscriptional RNA structure probing
RNA begins to fold as the nascent transcript emerges from a transcribing RNA polymerase. Here, the authors develop a concise method for mapping RNA folding pathways that couples in vitro transcription with high-throughput RNA chemical probing.
- Courtney E. Szyjka
- & Eric J. Strobel
-
Article
| Open Access Structural and dynamic mechanisms for coupled folding and tRNA recognition of a translational T-box riboswitch
Structural and dynamic mechanisms for coupled folding and tRNA recognition of a translational T-box riboswitch
T-box riboswitches are RNA-based gene regulators, composed of highly structured noncoding RNAs: the T-box and a tRNA ligand. Here, the authors assess the folding of a translational T-box aptamer and dissect the role of Mg2+, intra- and intermolecular RNA-RNA interactions in modulating its folding and function.
- Xiaolin Niu
- , Zhonghe Xu
- & Xianyang Fang
-
Article
| Open Access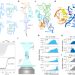 Direct observation of tRNA-chaperoned folding of a dynamic mRNA ensemble
Direct observation of tRNA-chaperoned folding of a dynamic mRNA ensemble
T-box riboswitch RNAs directly bind to specific tRNA and regulate the transcription or translation of downstream genes in bacteria. Using single-molecule FRET and ensemble biophysical analyses, here the authors uncover a Venus flytrap-like mechanism where tRNA binding to a T-box riboswitch mRNA triggers its rapid domain closure.
- Krishna C. Suddala
- , Janghyun Yoo
- & Jinwei Zhang
-
Article
| Open Access Automated design of protein-binding riboswitches for sensing human biomarkers in a cell-free expression system
Automated design of protein-binding riboswitches for sensing human biomarkers in a cell-free expression system
Cell-free genetically encoded biosensors have been developed to detect small molecules and nucleic acids, but they have yet to be reliably engineered to detect proteins. Here the authors develop an automated platform to convert protein-binding RNA aptamers into riboswitch sensors that operate within low-cost cell-free assays.
- Grace E. Vezeau
- , Lipika R. Gadila
- & Howard M. Salis
-
Article
| Open Access Observation of structural switch in nascent SAM-VI riboswitch during transcription at single-nucleotide and single-molecule resolution
Observation of structural switch in nascent SAM-VI riboswitch during transcription at single-nucleotide and single-molecule resolution
Observation of single nascent RNA at single-nucleotide resolution enables building a conformation landscape of how RNAs change during transcription. Here the authors develop a method to monitor the conformational change of individual SAM-VI riboswitch RNA (riboSAM) during transcription at single-nucleotide resolution as it is synthesized by transcription.
- Yanyan Xue
- , Jun Li
- & Yu Liu
-
Article
| Open Access Visualizing orthogonal RNAs simultaneously in live mammalian cells by fluorescence lifetime imaging microscopy (FLIM)
Visualizing orthogonal RNAs simultaneously in live mammalian cells by fluorescence lifetime imaging microscopy (FLIM)
No multi-color RNA fluorescent tags are currently available for use in live cells. Here, the authors show that fluorescence lifetime imaging microscopy is advantageous for multiplexed RNA visualization while achieving robust cellular contrast.
- Nadia Sarfraz
- , Emilia Moscoso
- & Esther Braselmann
-
Article
| Open Access Real-time monitoring of single ZTP riboswitches reveals a complex and kinetically controlled decision landscape
Real-time monitoring of single ZTP riboswitches reveals a complex and kinetically controlled decision landscape
Many RNAs become functional before their synthesis completes. Here the authors employ a single-molecule vectorial folding assay mimicking RNA transcription and show that the ZTP riboswitch is kinetically controlled and activated by slower unwinding and strategic pausing.
- Boyang Hua
- , Christopher P. Jones
- & Taekjip Ha
-
Article
| Open Access A SAM-I riboswitch with the ability to sense and respond to uncharged initiator tRNA
A SAM-I riboswitch with the ability to sense and respond to uncharged initiator tRNA
Riboswitches consist of an aptamer domain and expression platform, which senses a signal and regulates gene expression, respectively. Here the authors show that the expression platform of a SAM-I riboswitch from the Gram-negative bacteria can sense and bind uncharged an initiator Met tRNA to control the met operon.
- Dong-Jie Tang
- , Xinyu Du
- & Ji-Liang Tang
-
Article
| Open Access A riboswitch gives rise to multi-generational phenotypic heterogeneity in an auxotrophic bacterium
A riboswitch gives rise to multi-generational phenotypic heterogeneity in an auxotrophic bacterium
Auxotrophic bacteria rely on transporters to acquire essential compounds from their environment. Here, Hernandez-Valdes et al. show that a riboswitch generates long-term, stable heterogenous expression of a high-affinity methionine transporter in auxotrophic Lactococcus lactis.
- Jhonatan A. Hernandez-Valdes
- , Jordi van Gestel
- & Oscar P. Kuipers
-
Article
| Open Access Massively parallel RNA device engineering in mammalian cells with RNA-Seq
Massively parallel RNA device engineering in mammalian cells with RNA-Seq
Synthetic RNA-based devices can dynamically control a wide range of processes. Here the authors develop a quantitative and high-throughput mammalian cell-based RNA-seq assay to efficiently engineer ribozyme switches.
- Joy S. Xiang
- , Matias Kaplan
- & Christina D. Smolke
-
Article
| Open Access Local-to-global signal transduction at the core of a Mn2+ sensing riboswitch
Local-to-global signal transduction at the core of a Mn2+ sensing riboswitch
Riboswitches bind intracellular metabolites and control bacterial gene expression. Here, by using X-ray crystallography, molecular dynamics simulations, and single-molecule fluorescence resonance energy transfer, the authors show how a local Mn2+ ion-binding signal is transduced across the yybP-ykoY riboswitch from Xanthomonas oryzae.
- Krishna C. Suddala
- , Ian R. Price
- & Nils G. Walter
-
Article
| Open Access A tetracycline-dependent ribozyme switch allows conditional induction of gene expression in Caenorhabditis elegans
A tetracycline-dependent ribozyme switch allows conditional induction of gene expression in Caenorhabditis elegans
Tools for conditional induction of gene expression in C. elegans are limited compared to other organisms. Here the authors present a tetracycline-dependent ribozyme that allows conditional control of a gene of interest.
- Lena A. Wurmthaler
- , Monika Sack
- & Martin Gamerdinger
-
Article
| Open Access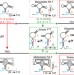 Life times of metastable states guide regulatory signaling in transcriptional riboswitches
Life times of metastable states guide regulatory signaling in transcriptional riboswitches
Riboswitches are RNA-based regulatory elements, which regulate downstream gene expression by binding of small molecular weight ligands. Here the authors demonstrate the molecular mechanism of a transcriptional riboswitch that integrates changes in transcription rates, metabolite concentration, and kinetic on- and off-rates of ligand binding.
- Christina Helmling
- , Dean-Paulos Klötzner
- & Harald Schwalbe
-
Article
| Open Access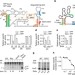 Transcriptional pausing at the translation start site operates as a critical checkpoint for riboswitch regulation
Transcriptional pausing at the translation start site operates as a critical checkpoint for riboswitch regulation
Riboswitches are non-coding RNA elements that detect metabolites and control expression by regulating mRNA levels or translation. Here, the authors provide evidence that theE. coli thiC riboswitch has a pause site in the translation initiation region that acts as a checkpoint for thiCexpression.
- Adrien Chauvier
- , Frédéric Picard-Jean
- & Daniel A. Lafontaine
-
Article
| Open Access The Shine-Dalgarno sequence of riboswitch-regulated single mRNAs shows ligand-dependent accessibility bursts
The Shine-Dalgarno sequence of riboswitch-regulated single mRNAs shows ligand-dependent accessibility bursts
In response to intracellular signals, bacterial translational riboswitches embedded in mRNAs can regulate gene expression through inhibition of translation initiation. Here, the authors describe SiM-KARTS, a novel approach for detecting changes in the structure of single RNA molecules in response to a ligand.
- Arlie J. Rinaldi
- , Paul E. Lund
- & Nils G. Walter
-
Article
| Open Access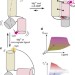 Rapid RNA–ligand interaction analysis through high-information content conformational and stability landscapes
Rapid RNA–ligand interaction analysis through high-information content conformational and stability landscapes
The structure and biological properties of RNAs are a function of changing cellular conditions. Here, Baird et al.report a high-throughput Förster resonance energy transfer (FRET) method to rapidly compare RNA structure modulation by cognate and non-cognate ligands across multiplexed solution conditions.
- Nathan J. Baird
- , James Inglese
- & Adrian R. Ferré-D’Amaré
-
Article
| Open Access Synthetic human cell fate regulation by protein-driven RNA switches
Synthetic human cell fate regulation by protein-driven RNA switches
The control of cell fate and apoptosis is a continuing challenge in synthetic biology. In this study, systems are developed in which an intracellularly expressed genome-encoded protein simultaneously achieves up- and downregulation of two distinct apoptosis pathways.
- Hirohide Saito
- , Yoshihiko Fujita
- & Tan Inoue

 A nascent riboswitch helix orchestrates robust transcriptional regulation through signal integration
A nascent riboswitch helix orchestrates robust transcriptional regulation through signal integration