Featured
-
-
Article
| Open Access A high-speed search engine pLink 2 with systematic evaluation for proteome-scale identification of cross-linked peptides
A high-speed search engine pLink 2 with systematic evaluation for proteome-scale identification of cross-linked peptides
The identification of cross-linked peptides at a proteome scale for interactome analyses represents a complex challenge. Here the authors report an efficient and reliable search engine pLink 2 for proteome-scale cross-linking mass spectrometry analyses, and demonstrate how to systematically evaluate the credibility of search engines.
- Zhen-Lin Chen
- , Jia-Ming Meng
- & Si-Min He
-
Article
| Open Access TurboID-based proximity labeling reveals that UBR7 is a regulator of N NLR immune receptor-mediated immunity
TurboID-based proximity labeling reveals that UBR7 is a regulator of N NLR immune receptor-mediated immunity
Plant NLR receptors trigger immune signaling following recognition of pathogen effectors. Here, Zhang et al. optimize a TurboID-based proximity labeling approach and show that it can be used to identify interacting partners of N, an NLR that confers resistance to Tobacco mosaic virus.
- Yongliang Zhang
- , Gaoyuan Song
- & Savithramma P. Dinesh-Kumar
-
Article
| Open Access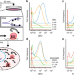 Parallel analysis of tri-molecular biosynthesis with cell identity and function in single cells
Parallel analysis of tri-molecular biosynthesis with cell identity and function in single cells
Simultaneous quantification of DNA, RNA and protein at the single cell level has not yet been possible. Here the authors introduce a molecular labelling and detection strategy to quantify synthesis of these biomolecules and couple it to transient cell states through parallel quantification of state-dependent biomolecules.
- Samuel C. Kimmey
- , Luciene Borges
- & Sean C. Bendall
-
Article
| Open Access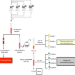 Dynamic molecular changes during the first week of human life follow a robust developmental trajectory
Dynamic molecular changes during the first week of human life follow a robust developmental trajectory
The first week of life impacts health for all of life, but the mechanisms are little-understood. Here the authors extract multi-omic data from small volumes of blood to study the dynamic molecular changes during the first week of life, revealing a robust developmental trajectory common to different populations.
- Amy H. Lee
- , Casey P. Shannon
- & Tobias R. Kollmann
-
Article
| Open Access Topological scoring of protein interaction networks
Topological scoring of protein interaction networks
Inferring direct protein−protein interactions (PPIs) and modules in PPI networks remains a challenge. Here, the authors introduce an algorithm to infer potential direct PPIs from quantitative proteomic AP-MS data by identifying enriched interactions of each bait relative to the other baits.
- Mihaela E. Sardiu
- , Joshua M. Gilmore
- & Michael P. Washburn
-
Article
| Open Access Combining LOPIT with differential ultracentrifugation for high-resolution spatial proteomics
Combining LOPIT with differential ultracentrifugation for high-resolution spatial proteomics
Spatial proteomics allows studying cellular protein localisations at system-wide scale. Here, the authors show that combining the previously developed hyperLOPIT method with differential centrifugation yields protein localisation maps at suborganellar resolution while reducing analysis time and input material.
- Aikaterini Geladaki
- , Nina Kočevar Britovšek
- & Kathryn S. Lilley
-
Article
| Open Access Global phosphoproteomic analysis reveals ARMC10 as an AMPK substrate that regulates mitochondrial dynamics
Global phosphoproteomic analysis reveals ARMC10 as an AMPK substrate that regulates mitochondrial dynamics
AMPK is regulates cellular energy and has been extensively studied, although our knowledge of downstream substrates is incomplete. Here, Chen et al. perform global quantitative analysis for AMPK-dependent sites and validate one hit, ARMC10, as a direct AMPK effector of mitochondrial dynamics.
- Zhen Chen
- , Caoqi Lei
- & Junjie Chen
-
Article
| Open Access Loss of peroxiredoxin-2 exacerbates eccentric contraction-induced force loss in dystrophin-deficient muscle
Loss of peroxiredoxin-2 exacerbates eccentric contraction-induced force loss in dystrophin-deficient muscle
In the mdx mouse model of Duchenne muscular dystrophy, muscle contractions lead to force loss, which is attributed to myofibre damage. Here, the authors show that force loss is instead mediated by a redox circuit involving NOX2, PROX1, myoglobin and cytoplasmic actins, and suggest that it may be a protective mechanism to prevent excessive contraction-induced myofibre damage.
- John T. Olthoff
- , Angus Lindsay
- & James M. Ervasti
-
Article
| Open Access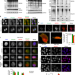 NEDDylation promotes nuclear protein aggregation and protects the Ubiquitin Proteasome System upon proteotoxic stress
NEDDylation promotes nuclear protein aggregation and protects the Ubiquitin Proteasome System upon proteotoxic stress
Protein NEDDylation increases upon proteotoxic stress but the function of this response remains to be elucidated. Here, the authors show that NEDDylation contributes to the cellular defence against proteotoxicity by promoting nuclear protein aggregation and protecting the ubiquitin proteasome system.
- Chantal M. Maghames
- , Sofia Lobato-Gil
- & Dimitris P. Xirodimas
-
Article
| Open Access Combining discovery and targeted proteomics reveals a prognostic signature in oral cancer
Combining discovery and targeted proteomics reveals a prognostic signature in oral cancer
Oral cancer has region-specific histopathological and molecular characteristics, complicating its classification by the standard tumor-node-metastasis system. Here, the authors combine discovery and targeted proteomics with IHC to identify region-specific and saliva biomarkers for oral cancer prognosis.
- Carolina Moretto Carnielli
- , Carolina Carneiro Soares Macedo
- & Adriana Franco Paes Leme
-
Article
| Open Access The dual methyltransferase METTL13 targets N terminus and Lys55 of eEF1A and modulates codon-specific translation rates
The dual methyltransferase METTL13 targets N terminus and Lys55 of eEF1A and modulates codon-specific translation rates
Eukaryotic elongation factor 1 alpha (eEF1A) is subject to extensive post-translational methylation but not all responsible enzymes are known. Here, the authors identify METTL13 as an eEF1A methyltransferase with dual specificity, which is involved in the codon-specific modulation of mRNA translation.
- Magnus E. Jakobsson
- , Jędrzej M. Małecki
- & Pål Ø. Falnes
-
Article
| Open Access The identification of carbon dioxide mediated protein post-translational modifications
The identification of carbon dioxide mediated protein post-translational modifications
Carbon dioxide can interact with proteins to form carbamate post-translational modifications. Here, the authors developed a strategy to identify carbamate post-translational modifications by trapping carbon dioxide and subsequently identifying the carbamylated proteins.
- Victoria L. Linthwaite
- , Joanna M. Janus
- & Martin J. Cann
-
Article
| Open Access Immuno-detection by sequencing enables large-scale high-dimensional phenotyping in cells
Immuno-detection by sequencing enables large-scale high-dimensional phenotyping in cells
Detecting proteins and post-translational modifications is important for drug screens, but the number of proteins measurable simultaneously is limited. Here the authors use antibodies tagged with DNA barcodes and high-throughput sequencing to detect up to 70 (phospho-)proteins in stem cells.
- Jessie A. G. van Buggenum
- , Jan P. Gerlach
- & Klaas W. Mulder
-
Article
| Open Access Deducing the presence of proteins and proteoforms in quantitative proteomics
Deducing the presence of proteins and proteoforms in quantitative proteomics
An accurate quantitation of different proteoforms remains challenging due to incomplete protein sequence coverage in mass spectrometry datasets. Here the authors describe a method to facilitate the discovery of proteoforms that may otherwise not be considered due to incomplete protein coverage or ambiguities in mapping peptides back to proteins.
- Casimir Bamberger
- , Salvador Martínez-Bartolomé
- & John R. Yates III
-
Article
| Open Access Reactive-site-centric chemoproteomics identifies a distinct class of deubiquitinase enzymes
Reactive-site-centric chemoproteomics identifies a distinct class of deubiquitinase enzymes
Deubiquitinases are proteases that cleave after the C-terminus of ubiquitin to hydrolyze ubiquitin chains and cleave ubiquitin from substrates. Here the authors describe a reactive-site-centric chemoproteomics approach to studying deubiquitinase activity, and expand the repertoire of known deubiquitinases.
- David S. Hewings
- , Johanna Heideker
- & Ingrid E. Wertz
-
Article
| Open Access Nanodroplet processing platform for deep and quantitative proteome profiling of 10–100 mammalian cells
Nanodroplet processing platform for deep and quantitative proteome profiling of 10–100 mammalian cells
There is a great need of developing highly sensitive mass spectrometry-based proteomics analysis for small cell populations. Here, the authors establish a robotically controlled chip-based nanodroplet processing platform and demonstrate its ability to profile the proteome from 10–100 mammalian cells.
- Ying Zhu
- , Paul D. Piehowski
- & Ryan T. Kelly
-
Article
| Open Access The STUbL RNF4 regulates protein group SUMOylation by targeting the SUMO conjugation machinery
The STUbL RNF4 regulates protein group SUMOylation by targeting the SUMO conjugation machinery
SUMO and ubiquitin are key signal transducers in several cellular processes including the DNA-damage response. Here the authors describe a method for selective enrichment of ubiquitin substrates for E3 ligases from complex cellular proteomes and identify the SUMO conjugation machinery as direct RNF4 substrates.
- Ramesh Kumar
- , Román González-Prieto
- & Alfred C. O. Vertegaal
-
Article
| Open Access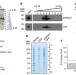 Plasma membrane-derived extracellular microvesicles mediate non-canonical intercellular NOTCH signaling
Plasma membrane-derived extracellular microvesicles mediate non-canonical intercellular NOTCH signaling
ARMMs are extracellular vesicles that bud directly at the plasma membrane; their function is poorly understood. Here the authors purify and carryout a proteomics analysis of the protein components of ARMMs, and show that NOTCH receptors are recruited into ARMMs and can be transferred to recipient cells to mediate NOTCH signaling.
- Qiyu Wang
- & Quan Lu
-
Article
| Open Access RNA localization is a key determinant of neurite-enriched proteome
RNA localization is a key determinant of neurite-enriched proteome
Subcellular localization of RNAs and proteins is important for polarized cells such as neurons. Here the authors differentiate mouse embryonic stem cells into neurons, and analyze the local transcriptome, proteome, and translated transcriptome in their cell bodies and neurites, providing a unique resource for future studies on neuronal polarity.
- Alessandra Zappulo
- , David van den Bruck
- & Marina Chekulaeva
-
Article
| Open Access pGlyco 2.0 enables precision N-glycoproteomics with comprehensive quality control and one-step mass spectrometry for intact glycopeptide identification
pGlyco 2.0 enables precision N-glycoproteomics with comprehensive quality control and one-step mass spectrometry for intact glycopeptide identification
Protein glycosylation is a heterogeneous post-translational modification that generates greater proteomic diversity that is difficult to analyze. Here the authors describe pGlyco 2.0, a workflow for the precise one step identification of intact N-glycopeptides at the proteome scale.
- Ming-Qi Liu
- , Wen-Feng Zeng
- & Peng-Yuan Yang
-
Article
| Open Access Multi-laboratory assessment of reproducibility, qualitative and quantitative performance of SWATH-mass spectrometry
Multi-laboratory assessment of reproducibility, qualitative and quantitative performance of SWATH-mass spectrometry
SWATH-mass spectrometry consists of a data-independent acquisition and a targeted data analysis strategy that aims to maintain the favorable quantitative characteristics on the scale of thousands of proteins. Here, using data generated by eleven groups worldwide, the authors show that SWATH-MS is capable of generating highly reproducible data across different laboratories.
- Ben C. Collins
- , Christie L. Hunter
- & Ruedi Aebersold
-
Article
| Open Access A scalable double-barcode sequencing platform for characterization of dynamic protein-protein interactions
A scalable double-barcode sequencing platform for characterization of dynamic protein-protein interactions
Protein-protein interactions (PPIs) play a major role in defining biological functions. Here, the authors present PPiSeq, a method to quantitatively score dynamic PPIs in yeast combining fragment complementation assays, genomic double-barcoding, and an analytical framework to precisely call fitness from barcode lineage trajectories.
- Ulrich Schlecht
- , Zhimin Liu
- & Sasha F. Levy
-
Article
| Open Access An engineered high affinity Fbs1 carbohydrate binding protein for selective capture of N-glycans and N-glycopeptides
An engineered high affinity Fbs1 carbohydrate binding protein for selective capture of N-glycans and N-glycopeptides
Protein glycosylation is an essential post-translational modification which analysis is complicated by the diversity of glycan composition and heterogeneity at individual attachment sites. Here the authors describe a method to selectively enrich N-linked glycopeptides to facilitate the detection of micro-heterogeneity in N-glycosylation.
- Minyong Chen
- , Xiaofeng Shi
- & James C. Samuelson
-
Article
| Open Access Uncovering the SUMOylation and ubiquitylation crosstalk in human cells using sequential peptide immunopurification
Uncovering the SUMOylation and ubiquitylation crosstalk in human cells using sequential peptide immunopurification
Ubiquitylation and SUMOylation are two important related post-translational modifications. Here the authors present an approach for the simultaneous identification and quantification of protein-wide SUMO and ubiquitin sites from a single sample, uncovering widespread crosstalk between the two modifications.
- Frédéric Lamoliatte
- , Francis P. McManus
- & Pierre Thibault
-
Article
| Open Access Identification of microsporidia host-exposed proteins reveals a repertoire of rapidly evolving proteins
Identification of microsporidia host-exposed proteins reveals a repertoire of rapidly evolving proteins
Unbiased identification of proteins from pathogens that are exposed to a host can provide insight into host–pathogen interaction. Here, the authors use an enzymatic tagging method and mass spectrometry to identify rapidly evolvingNematocida microsporidia proteins when infecting C. elegans.
- Aaron W. Reinke
- , Keir M. Balla
- & Emily R. Troemel
-
Article
| Open Access A bead-based western for high-throughput cellular signal transduction analyses
A bead-based western for high-throughput cellular signal transduction analyses
Dissecting cellular signalling requires the analysis of large numbers of proteins. Here the authors describe DigiWest, a high-throughput protein detection method that combines the concept of western and widely-used bead array systems that allows rapid quantification of hundreds of specific proteins.
- Fridolin Treindl
- , Benjamin Ruprecht
- & Markus F. Templin
-
Article
| Open Access A proteomic approach reveals integrin activation state-dependent control of microtubule cortical targeting
A proteomic approach reveals integrin activation state-dependent control of microtubule cortical targeting
Integrins are activated by many extracellular cues and respond by assembling diverse signalling complexes. Byron et al.use activation state-specific antibodies to proteomically characterize these complexes, and provide insight into integrin-dependent microtubule stabilization.
- Adam Byron
- , Janet A. Askari
- & Martin J. Humphries
-
Article
| Open Access Proteomic screen reveals Fbw7 as a modulator of the NF-κB pathway
Proteomic screen reveals Fbw7 as a modulator of the NF-κB pathway
Fbw7 is a ubiquitin-ligase, which targets several oncoproteins for proteolysis, and is therefore important for the control and prevention of tumorigenesis. In this study, Arabi and colleagues carry out a proteomic screen of the targets of Fbw7, and identify Nuclear Factor of κ-B2 as a substrate.
- Azadeh Arabi
- , Karim Ullah
- & Olle Sangfelt

 Haploinsufficiency in the ANKS1B gene encoding AIDA-1 leads to a neurodevelopmental syndrome
Haploinsufficiency in the ANKS1B gene encoding AIDA-1 leads to a neurodevelopmental syndrome