Article
|
Open Access
Featured
-
-
Article
| Open Access Inferring the molecular and phenotypic impact of amino acid variants with MutPred2
Inferring the molecular and phenotypic impact of amino acid variants with MutPred2
Identifying variants capable of causing genetic disease is challenging. The authors use semisupervised learning to predict pathogenic missense variants and their impacts on protein structure and function, enabling a molecular mechanism-driven approach to studying different types of human disease.
- Vikas Pejaver
- , Jorge Urresti
- & Predrag Radivojac
-
Article
| Open Access Transforming protein-polymer conjugate purification by tuning protein solubility
Transforming protein-polymer conjugate purification by tuning protein solubility
Therapeutic proteins are often conjugated with polymers, but separating the conjugate from unconjugated protein and free polymer is a major challenge. Here, the authors discover that proteins conjugated to charged or zwitterionic polymers maintain solubility in 100% ammonium sulfate, greatly simplifying purification.
- Stefanie L. Baker
- , Aravinda Munasinghe
- & Alan J. Russell
-
Article
| Open Access Deep learning extends de novo protein modelling coverage of genomes using iteratively predicted structural constraints
Deep learning extends de novo protein modelling coverage of genomes using iteratively predicted structural constraints
Prediction of protein structures on the scale of genomes remains a challenge. Here the authors introduce a protein structure prediction method that uses deep learning to predict inter-atomic distances, torsion angles and hydrogen bonds, and apply it to predict the structures of 1475 Pfam domains.
- Joe G. Greener
- , Shaun M. Kandathil
- & David T. Jones
-
Article
| Open Access Environmental conditions shape the nature of a minimal bacterial genome
Environmental conditions shape the nature of a minimal bacterial genome
Minimal bacterial genomes still contain hundreds of genes of unknown function. Here the authors use in silico annotation methods and identify the environmental factors shaping a minimal genome.
- Magdalena Antczak
- , Martin Michaelis
- & Mark N. Wass
-
Article
| Open Access Rapid determination of quaternary protein structures in complex biological samples
Rapid determination of quaternary protein structures in complex biological samples
Protein structure determination in complex biological samples is still challenging. Here, the authors develop a computational modeling-guided cross-linking mass spectrometry method, obtaining a high-resolution model of a 1.8 MDa protein assembly from cross-links detected in a mixture of human plasma and bacteria.
- Simon Hauri
- , Hamed Khakzad
- & Lars Malmström
-
Article
| Open Access Automated NMR resonance assignments and structure determination using a minimal set of 4D spectra
Automated NMR resonance assignments and structure determination using a minimal set of 4D spectra
Further automation of NMR structure determination is needed to increase the throughput and accessibility of this method. Here the authors present 4D-CHAINS/autoNOE-Rosetta, a complete pipeline that allows rapid and fully automated structure determination from two highly complementary NMR datasets.
- Thomas Evangelidis
- , Santrupti Nerli
- & Konstantinos Tripsianes
-
Article
| Open Access MHC class II complexes sample intermediate states along the peptide exchange pathway
MHC class II complexes sample intermediate states along the peptide exchange pathway
MHCII proteins bind and present both foreign and self-antigens to potentially activate CD4+ T cells via cognate T cell receptors (TCRs) during the adaptive immune response. Here, the authors combine NMR-detected H/D exchange with Markov modelling analysis to shed light on the dynamics of MHCII peptide exchange.
- Marek Wieczorek
- , Jana Sticht
- & Christian Freund
-
Article
| Open Access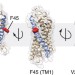 Dimerization deficiency of enigmatic retinitis pigmentosa-linked rhodopsin mutants
Dimerization deficiency of enigmatic retinitis pigmentosa-linked rhodopsin mutants
Retinitis pigmentosa is often caused by mutations that affect the activity or transport of rhodopsin, but some mutations cause disease even though an apparently functional protein is produced. Here the authors show that three such enigmatic mutants retain scramblase activity but are unable to dimerize.
- Birgit Ploier
- , Lydia N. Caro
- & Anant K. Menon
-
Article
| Open Access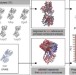 Prediction and validation of protein intermediate states from structurally rich ensembles and coarse-grained simulations
Prediction and validation of protein intermediate states from structurally rich ensembles and coarse-grained simulations
Protein conformational changes are key to a wide range of cellular functions but remain difficult to access experimentally. Here the authors describe eBDIMS, a novel approach to predict intermediates observed in structural transition pathways from experimental ensembles.
- Laura Orellana
- , Ozge Yoluk
- & Erik Lindahl
-
Article
| Open Access Prediction of allosteric sites and mediating interactions through bond-to-bond propensities
Prediction of allosteric sites and mediating interactions through bond-to-bond propensities
Allostery is a key molecular mechanism underpinning control and modulation in a variety of cellular processes. Here, the authors present a method that can be used to predict allosteric sites and the mediating interactions that connect them to the active site of the protein.
- B. R. C. Amor
- , M. T. Schaub
- & M. Barahona
-
Article |
 Evolutionary-guided de novo structure prediction of self-associated transmembrane helical proteins with near-atomic accuracy
Evolutionary-guided de novo structure prediction of self-associated transmembrane helical proteins with near-atomic accuracy
While the transmembrane regions of single-pass transmembrane proteins play critical roles in receptor signalling, they remain difficult to characterize structurally. Here the authors present a computational approach for accurate structure prediction of associated single-pass transmembrane helical proteins.
- Y. Wang
- & P. Barth

 Integrative modeling of membrane-associated protein assemblies
Integrative modeling of membrane-associated protein assemblies