Featured
-
-
Article
| Open Access Regulation of Rad52-dependent replication fork recovery through serine ADP-ribosylation of PolD3
Regulation of Rad52-dependent replication fork recovery through serine ADP-ribosylation of PolD3
Here the authors identify that PARP1 maintains genome integrity by regulating replication fork recovery by break-induced replication. Mechanistically, this is achieved through MRE11-dependent PARP1 activation and site-specific ADP-ribosylation of PolD3.
- Frederick Richards
- , Marta J. Llorca-Cardenosa
- & Nicholas D. Lakin
-
Article
| Open Access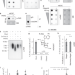 Mammalian N1-adenosine PARylation is a reversible DNA modification
Mammalian N1-adenosine PARylation is a reversible DNA modification
Poly-ADP-ribosylation (PARylation) is a well-known posttranslational modification of proteins. Here the authors show that beyond proteins also mammalian single-stranded DNA is PARylated in vitro and in vivo.
- Michael U. Musheev
- , Lars Schomacher
- & Christof Niehrs
-
Article
| Open Access Deubiquitinating enzymes and the proteasome regulate preferential sets of ubiquitin substrates
Deubiquitinating enzymes and the proteasome regulate preferential sets of ubiquitin substrates
Deubiquitinases (DUBs) remove ubiquitin from its target proteins. Here, authors compare the regulatory effects of the proteasome and DUBs on the ubiquitinated proteome. They find preferential sets of substrates regulated by DUBs or by the proteasome. Moreover, they find that PARP1 is hyper-ubiquitinated in response to DUB inhibition, which increases its enzymatic activity.
- Fredrik Trulsson
- , Vyacheslav Akimov
- & Alfred C. O. Vertegaal
-
Article
| Open Access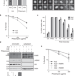 Linking DNA repair and cell cycle progression through serine ADP-ribosylation of histones
Linking DNA repair and cell cycle progression through serine ADP-ribosylation of histones
Poly(ADP-ribose)-polymerases (PARPs) are a cornerstone of the DNA damage response that promote DNA repair by modifying target proteins with ADP-ribose. Here, the authors show serine ADP-ribosylation of the H3 variant H3b maintains genome stability by coupling DNA repair with mitotic entry in Dictyostelium by regulating double strand break repair by nonhomologous end-joining (NHEJ).
- Julien Brustel
- , Tetsuya Muramoto
- & Nicholas D. Lakin
-
Article
| Open Access HPF1 dynamically controls the PARP1/2 balance between initiating and elongating ADP-ribose modifications
HPF1 dynamically controls the PARP1/2 balance between initiating and elongating ADP-ribose modifications
HPF1 controls the ADP-ribosylation activity of PARP1/2 in response to DNA breaks. Here, the authors show that HPF1 regulates the balance between ADP-ribose initiation and elongation through a dynamic interaction that accelerates the initiation rate on serine residues.
- Marie-France Langelier
- , Ramya Billur
- & John M. Pascal
-
Article
| Open Access SPINDOC binds PARP1 to facilitate PARylation
SPINDOC binds PARP1 to facilitate PARylation
SPINDOC is known to interact with Spindlin1 (SPIN1), a histone code effector protein. Here, the authors show that SPINDOC is distributed between two distinct protein complexes, one comprising SPIN1 and the other one with PARP1. Their results suggest a role for SPINDOC in the regulation of PARP1- mediated PARylation and the DNA damage response.
- Fen Yang
- , Jianji Chen
- & Mark T. Bedford
-
Article
| Open Access The regulatory landscape of the human HPF1- and ARH3-dependent ADP-ribosylome
The regulatory landscape of the human HPF1- and ARH3-dependent ADP-ribosylome
ADP-ribosylation is regulated by HPF1 and ARH3, but the cellular target spectrum of these enzymes is not fully understood. Here, the authors use quantitative proteomics to define the HPF1- and ARH3-dependent ADP-ribosylome, providing evidence that mono-ADP-ribosylation of serine predominates in cells.
- Ivo A. Hendriks
- , Sara C. Buch-Larsen
- & Michael L. Nielsen
-
Article
| Open Access Mechanistic insights into the three steps of poly(ADP-ribosylation) reversal
Mechanistic insights into the three steps of poly(ADP-ribosylation) reversal
PARG and ARH3 are the main hydrolases to reverse serine poly(ADP-ribosylation) yet their activities in the process differ. Here, the authors synthesise linear and branched poly(ADP-ribose) molecules, perform structure-function analysis and elucidate the mechanistic differences between PARG and ARH3.
- Johannes Gregor Matthias Rack
- , Qiang Liu
- & Ivan Ahel
-
Article
| Open Access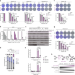 Serine-linked PARP1 auto-modification controls PARP inhibitor response
Serine-linked PARP1 auto-modification controls PARP inhibitor response
PARP inhibitors function by trapping PARP1 protein on DNA breaks, which has cytotoxic consequences to cancer cells. Here the authors identify three serine residues within PARP1 as key sites whose efficient HPF1-dependent modification counters PARP1 trapping and contributes to inhibitor tolerance.
- Evgeniia Prokhorova
- , Florian Zobel
- & Ivan Ahel
-
Article
| Open Access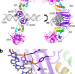 Activation of PARP2/ARTD2 by DNA damage induces conformational changes relieving enzyme autoinhibition
Activation of PARP2/ARTD2 by DNA damage induces conformational changes relieving enzyme autoinhibition
Poly(ADP-ribose) polymerase 2 (PARP2) is activated by 5′-phosphorylated DNA breaks but the molecular mechanism is not fully understood. Here, the authors report a crystal structure of PARP2 bound to an activating DNA fragment, providing insights into the structural changes that lead to PARP2 activation.
- Ezeogo Obaji
- , Mirko M. Maksimainen
- & Lari Lehtiö
-
Article
| Open Access Cells recognize osmotic stress through liquid–liquid phase separation lubricated with poly(ADP-ribose)
Cells recognize osmotic stress through liquid–liquid phase separation lubricated with poly(ADP-ribose)
Cells experience various osmotic perturbation, but cellular osmosensing mechanisms remain obscure. Here, the authors report that cells recognize osmotic stress from the inside through macromolecular crowding-driven and poly(ADP-ribose)-conditioned liquid–liquid phase separation.
- Kengo Watanabe
- , Kazuhiro Morishita
- & Hidenori Ichijo
-
Article
| Open Access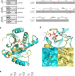 Engineering Af1521 improves ADP-ribose binding and identification of ADP-ribosylated proteins
Engineering Af1521 improves ADP-ribose binding and identification of ADP-ribosylated proteins
ADP-ribose binding macro domains facilitate the enrichment and detection of cellular ADP-ribosylation. Here, the authors generate an engineered macro domain with increased ADP-ribose affinity, improving the identification of ADP-ribosylated proteins by proteomics, western blot and immunofluorescence.
- Kathrin Nowak
- , Florian Rosenthal
- & Michael O. Hottiger
-
Article
| Open Access Real-time monitoring of PARP1-dependent PARylation by ATR-FTIR spectroscopy
Real-time monitoring of PARP1-dependent PARylation by ATR-FTIR spectroscopy
The mechanism of PARP1-dependent poly-ADP-ribosylation in response to DNA damage is still under debate. Here, the authors use ATR-FTIR spectroscopy to provide time-resolved insights into the molecular details of this process under near physiological conditions.
- Annika Krüger
- , Alexander Bürkle
- & Aswin Mangerich
-
Article
| Open Access Tankyrase inhibition preserves osteoarthritic cartilage by coordinating cartilage matrix anabolism via effects on SOX9 PARylation
Tankyrase inhibition preserves osteoarthritic cartilage by coordinating cartilage matrix anabolism via effects on SOX9 PARylation
Osteoarthritis results from the progressive destruction of cartilage matrix. Here, Kim et al. identify tankyrase as a regulator of cartilage matrix anabolism, and find that tankyrase inhibition, by preventing SOX9 PARylation, protects from cartilage destruction in a mouse model of osteoarthritis.
- Sukyeong Kim
- , Sangbin Han
- & Jin-Hong Kim
-
Article
| Open Access A ribose-functionalized NAD+ with unexpected high activity and selectivity for protein poly-ADP-ribosylation
A ribose-functionalized NAD+ with unexpected high activity and selectivity for protein poly-ADP-ribosylation
The study of NAD+ dependent ADP-ribosylation can be challenging. Here the authors report on the development of NAD+ analogues, using chemo-enzymatic methods, which can be used as probes to label the substrate proteins of poly-ADP-ribose polymerase.
- Xiao-Nan Zhang
- , Qinqin Cheng
- & Yong Zhang
-
Article
| Open Access Structural and biochemical evidence supporting poly ADP-ribosylation in the bacterium Deinococcus radiodurans
Structural and biochemical evidence supporting poly ADP-ribosylation in the bacterium Deinococcus radiodurans
Poly-ADP-ribosylation (PARylation) is a well-known regulatory event in eukaryotes but has not yet been observed in bacteria. Here, the authors solve the structure of a bacterial PAR-glycohydrolase and provide evidence for a prokaryotic PARylation machinery involved in the response to genotoxic stress.
- Chao-Cheng Cho
- , Chia-Yu Chien
- & Chun-Hua Hsu
-
Review Article
| Open Access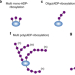 Emerging roles of eraser enzymes in the dynamic control of protein ADP-ribosylation
Emerging roles of eraser enzymes in the dynamic control of protein ADP-ribosylation
ADP-ribose erasing enzymes are increasingly recognized as critical regulators of protein ADP-ribosylation dynamics in living systems. Here, the authors review recent advances in the discovery and characterization of ADP-ribose erasers and discuss their role within the cellular ADP-ribosylation machinery.
- Julia O’Sullivan
- , Maria Tedim Ferreira
- & Guy G. Poirier
-
Article
| Open Access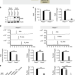 PARP2 mediates branched poly ADP-ribosylation in response to DNA damage
PARP2 mediates branched poly ADP-ribosylation in response to DNA damage
PARP1 and PARP2 of the PARP family enzymes are involved in DNA damage response. Here the authors report PARP2 activation mechanisms and its role in the formation of branched poly(ADP-ribose) chains in response to DNA damage.
- Qian Chen
- , Muzaffer Ahmad Kassab
- & Xiaochun Yu
-
Article
| Open Access Coupling bimolecular PARylation biosensors with genetic screens to identify PARylation targets
Coupling bimolecular PARylation biosensors with genetic screens to identify PARylation targets
Poly ADP-ribosylation (PARylation) is a highly dynamic post-translation protein modification, but most methods only detect stable PARylation events. Here the authors develop a split-GFP-based sensor for PARylation detection in live cells and use it to identify a new centrosomal PARylation target.
- Dragomir B. Krastev
- , Stephen J. Pettitt
- & Christopher J. Lord
-
Article
| Open Access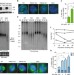 Whole proteome analysis of human tankyrase knockout cells reveals targets of tankyrase-mediated degradation
Whole proteome analysis of human tankyrase knockout cells reveals targets of tankyrase-mediated degradation
Tankyrase 1 and 2 are poly(ADP-ribose) polymerases that mark proteins for degradation, but there is a current lack of knowledge about their distinct functions and substrates. Here, the authors elucidate the cellular roles and substrates of these polymerases using comparative functional and proteomics analyses of tankyrase knockout cell lines.
- Amit Bhardwaj
- , Yanling Yang
- & Susan Smith
-
Article
| Open Access Proteomic analyses identify ARH3 as a serine mono-ADP-ribosylhydrolase
Proteomic analyses identify ARH3 as a serine mono-ADP-ribosylhydrolase
Protein ADP-ribosylation has emerged as a key post translational modification that regulates several stress responses. Here the authors characterize ARH3 as a major serine-specific mono–ADP-ribosylhydrolase and use a proteomics approach to identify the cellular targets of ARH3.
- Jeannette Abplanalp
- , Mario Leutert
- & Michael O. Hottiger
-
Article
| Open Access PARP1 promotes gene expression at the post-transcriptional level by modulating the RNA-binding protein HuR
PARP1 promotes gene expression at the post-transcriptional level by modulating the RNA-binding protein HuR
PARP1, in addition to its role in DNA repair, has a role in regulating gene transcription via PARylation of target proteins. Here the authors show that HuR is targeted after lipopolysaccharide exposure to regulate the inflammatory gene expression at post-transcriptional level.
- Yueshuang Ke
- , Yanlong Han
- & Xianlu Zeng
-
Article
| Open Access Proteome-wide identification of the endogenous ADP-ribosylome of mammalian cells and tissue
Proteome-wide identification of the endogenous ADP-ribosylome of mammalian cells and tissue
ADP-ribosylation is a reversible post-translational protein modification involved in many cellular processes. Here the authors describe a sensitive approach for the analysis of ADP-ribosylation sites under physiologic conditions and identify lysine residues as in vivotargets of ADP-ribosylation.
- Rita Martello
- , Mario Leutert
- & Michael L. Nielsen
-
Article
| Open Access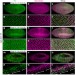 Wnt pathway activation by ADP-ribosylation
Wnt pathway activation by ADP-ribosylation
Wnt/β-catenin signalling directs several developmental processes and is aberrantly activated in several cancers. Here the authors implicate Tankyrase—previously shown to target the scaffolding protein Axin for proteolysis—in early Wnt signalling by promoting the interaction between Axin and the Wnt co-receptor LRP6.
- Eungi Yang
- , Ofelia Tacchelly-Benites
- & Yashi Ahmed
-
Article |
 Family-wide analysis of poly(ADP-ribose) polymerase activity
Family-wide analysis of poly(ADP-ribose) polymerase activity
The poly(ADP-ribose) polymerase family of enzymes control many aspects of cellular signalling by covalently modifying proteins with either poly- or mono-(ADP-ribose). Vyas et al.catalogue the catalytic specificity of this family, and reveal that the majority of these enzymes generate only mono(ADP-ribose).
- Sejal Vyas
- , Ivan Matic
- & Paul Chang
-
Article |
 A systematic analysis of the PARP protein family identifies new functions critical for cell physiology
A systematic analysis of the PARP protein family identifies new functions critical for cell physiology
The poly(ADP-ribose) polymerase (PARP) family includes 17 proteins in humans, many of which have no known function. Vyas et al.systematically characterize the localization and function of each human PARP and identify PARP14 as a regulator of focal adhesions.
- Sejal Vyas
- , Melissa Chesarone-Cataldo
- & Paul Chang
-
Article
| Open Access Visualization of poly(ADP-ribose) bound to PARG reveals inherent balance between exo- and endo-glycohydrolase activities
Visualization of poly(ADP-ribose) bound to PARG reveals inherent balance between exo- and endo-glycohydrolase activities
Poly-ADP-ribosylation is a post-translational modification that is countered by poly(ADP-ribose) glycohydrolases (PARGs). In this study, the authors present the crystal structure of poly(ADP-ribose) glycohydrolase (PARGs) in complex with a poly(ADP-ribose) substrate, and reveal that poly(ADP-ribose) glycohydrolase (PARGs) enzymes act predominantly as exo- rather than as endo-glycohydrolases.
- Eva Barkauskaite
- , Amy Brassington
- & David Leys

 Preserving ester-linked modifications reveals glutamate and aspartate mono-ADP-ribosylation by PARP1 and its reversal by PARG
Preserving ester-linked modifications reveals glutamate and aspartate mono-ADP-ribosylation by PARP1 and its reversal by PARG