Article
|
Open Access
Featured
-
-
Article
| Open Access Synchrony of Bird Migration with Global Dispersal of Avian Influenza Reveals Exposed Bird Orders
Synchrony of Bird Migration with Global Dispersal of Avian Influenza Reveals Exposed Bird Orders
Highly pathogenic avian influenza virus subtype H5 is an important pathogen of wild birds and poultry that has also caused infection in humans and other mammals. Here the authors use wild bird movement tracking data and virus genome sequences to quantify how seasonal bird migration facilitates global dispersal of the virus.
- Qiqi Yang
- , Ben Wang
- & Bryan Grenfell
-
Article
| Open Access Diversity and dissemination of viruses in pathogenic protozoa
Diversity and dissemination of viruses in pathogenic protozoa
Heeren et al study the evolutionary genomics of leishmaniasis in Peru and Bolivia to show that parasite hybridization increases the prevalence, diversity and spread of viruses that have been previously associated with disease severity and treatment failure.
- Senne Heeren
- , Ilse Maes
- & Frederik Van den Broeck
-
Article
| Open Access Phylotranscriptomics unveil a Paleoproterozoic-Mesoproterozoic origin and deep relationships of the Viridiplantae
Phylotranscriptomics unveil a Paleoproterozoic-Mesoproterozoic origin and deep relationships of the Viridiplantae
Evolutionary relationships among green plants are unresolved and, in particular, the phylogenetic position of Prasinodermophyta remains controversial. Here, the authors conduct phylogenomic analyses to resolve relationships within Viridiplantae, suggesting that this group diverged between the Great Oxidation Event and the Neoproterozoic Oxygenation Event.
- Zhiping Yang
- , Xiaoya Ma
- & Bojian Zhong
-
Article
| Open Access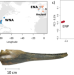 Ancient dolphin genomes reveal rapid repeated adaptation to coastal waters
Ancient dolphin genomes reveal rapid repeated adaptation to coastal waters
The chronology and mode of parallel evolution remain unclear. Here, the authors compare mid-Holocene and contemporary bottlenose dolphin adaptations between pelagic and coastal ecosystems with paleogenomics, finding rapid adaptation to newly emerged habitat from standing genetic variation.
- Marie Louis
- , Petra Korlević
- & Andrew D. Foote
-
Article
| Open Access Phylodynamic of SARS-CoV-2 during the second wave of COVID-19 in Peru
Phylodynamic of SARS-CoV-2 during the second wave of COVID-19 in Peru
The second SARS-CoV-2 wave in Peru had a high case fatality rate with Lambda and Gamma causing most cases. Using phylodynamics, the authors here show that Lambda most likely originated in Peru from where it spread to other South American countries and that the center of Peru played a key role in transmission to other regions.
- Santiago Justo Arevalo
- , Carmen Sofia Uribe Calampa
- & Joao Renato Rebello Pinho
-
Article
| Open Access Origin of minicircular mitochondrial genomes in red algae
Origin of minicircular mitochondrial genomes in red algae
While the organelle genome is commonly considered to be a single circular DNA molecule, extensive variation exists. Here, the authors report multipartite minicircular genomes in red algae and indicate an origin driven by recombination due to loss of DNA replication, recombination, and repair genes.
- Yongsung Lee
- , Chung Hyun Cho
- & Hwan Su Yoon
-
Article
| Open Access The evolution and international spread of extensively drug resistant Shigella sonnei
The evolution and international spread of extensively drug resistant Shigella sonnei
An increase in shigellosis cases among men who have sex with men in the United Kingdom has been linked to an extensively drug-resistant strain of Shigella sonnei. In this genomic epidemiology study, the authors investigate the genetic basis, evolutionary history, and international dissemination of the outbreak strain.
- Lewis C. E. Mason
- , David R. Greig
- & Kate S. Baker
-
Article
| Open Access Key innovations and the diversification of Hymenoptera
Key innovations and the diversification of Hymenoptera
Hymenoptera is an incredibly diverse order, with numerous behavioral and morphological innovations. Here, the authors compile a time-calibrated Hymenoptera phylogeny and find that secondary transitions to phytophagy, plant feeding, are associated with significant increases in diversification rate in this group.
- Bonnie B. Blaimer
- , Bernardo F. Santos
- & Matthew L. Buffington
-
Article
| Open Access Muscle5: High-accuracy alignment ensembles enable unbiased assessments of sequence homology and phylogeny
Muscle5: High-accuracy alignment ensembles enable unbiased assessments of sequence homology and phylogeny
Multiple sequence alignments are widely used to predict protein structure, function, and phylogeny, but are uncertain with more diverged sequences. Muscle5 generates ensembles of alternative high-accurate alignments, enabling novel confidence estimates in alignments, trees, and other inferences.
- Robert C. Edgar
-
Article
| Open Access Tracing the international arrivals of SARS-CoV-2 Omicron variants after Aotearoa New Zealand reopened its border
Tracing the international arrivals of SARS-CoV-2 Omicron variants after Aotearoa New Zealand reopened its border
In March 2022, Aotearoa New Zealand re-opened its border allowing quarantine-free travel for many travellers. Here, the authors describe circulating Omicron sub-variants before and after the reopening of the border and show that the rate of viral introductions grew roughly linearly with the increase in daily international travel.
- Jordan Douglas
- , David Winter
- & Jemma L. Geoghegan
-
Article
| Open Access Genomic insights into rapid speciation within the world’s largest tree genus Syzygium
Genomic insights into rapid speciation within the world’s largest tree genus Syzygium
The relative importance of the mechanisms underlying species radiation remains unclear. Here, the authors combine reference genome assembly and population genetics analyses to show that neutral forces have contributed to the radiation of the most species-rich tree genus Syzygium.
- Yee Wen Low
- , Sitaram Rajaraman
- & Victor A. Albert
-
Article
| Open Access Delineating Mycobacterium abscessus population structure and transmission employing high-resolution core genome multilocus sequence typing
Delineating Mycobacterium abscessus population structure and transmission employing high-resolution core genome multilocus sequence typing
Mycobacterium abscessus is an emerging infection of increasing public health concern due to outbreaks and intrinsic multidrug-resistance. Here, the authors develop and evaluate a core-genome multilocus sequence typing scheme for this pathogen to facilitate standardised molecular surveillance.
- Margo Diricks
- , Matthias Merker
- & Florian P. Maurer
-
Article
| Open Access Vibrio cholerae O139 genomes provide a clue to why it may have failed to usher in the eighth cholera pandemic
Vibrio cholerae O139 genomes provide a clue to why it may have failed to usher in the eighth cholera pandemic
The O139 Vibrio cholerae serogroup emerged in the 1990s and spread rapidly but did not become globally dominant. Here, the authors describe the genomic epidemiology of this strain and identify changes in virulence and antimicrobial resistance characteristics that they hypothesise may have contributed to its decline.
- Thandavarayan Ramamurthy
- , Agila Kumari Pragasam
- & Ankur Mutreja
-
Article
| Open Access Genomic distances reveal relationships of wild and cultivated beets
Genomic distances reveal relationships of wild and cultivated beets
While a large amount of genomic resources is available, the phylogeny of wild and cultivated beets remains unclear. Here, the authors use the k-mer-based Mash method to analyze resequenced genomes of 606 accessions of the genus Beta and reveal Greece as the domestication site of sugar beet.
- Felix L. Sandell
- , Nancy Stralis-Pavese
- & Juliane C. Dohm
-
Article
| Open Access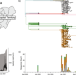 Multiple expansions of globally uncommon SARS-CoV-2 lineages in Nigeria
Multiple expansions of globally uncommon SARS-CoV-2 lineages in Nigeria
SARS-CoV-2 genomic surveillance has been important for informing pandemic responses but many regions remain under-sampled, limiting knowledge of circulating strains. Here, the authors sequence 378 isolates from Nigeria and identify two strains that appear to be important locally though globally uncommon.
- Egon A. Ozer
- , Lacy M. Simons
- & Ramon Lorenzo-Redondo
-
Article
| Open Access Population structure analysis and laboratory monitoring of Shigella by core-genome multilocus sequence typing
Population structure analysis and laboratory monitoring of Shigella by core-genome multilocus sequence typing
Lab-based surveillance of Shigella has traditionally been based on serotyping but increasing availability of whole genome sequencing could enable higher resolution typing. Here, the authors apply a core genome multilocus sequence typing scheme to Shigella sequence data and describe its population structure.
- Iman Yassine
- , Sophie Lefèvre
- & François-Xavier Weill
-
Article
| Open Access Chloranthus genome provides insights into the early diversification of angiosperms
Chloranthus genome provides insights into the early diversification of angiosperms
Chloranthales remain the last lineage of core angiosperms that lacks a nuclear genome assembly. Here, the authors report the genome assembly of Chloranthus spicatus and show its contribution to deepen our understanding on diversification, phylogeny, and genome evolution in angiosperms.
- Xing Guo
- , Dongming Fang
- & Huan Liu
-
Article
| Open Access Genomic sequencing of SARS-CoV-2 in Rwanda reveals the importance of incoming travelers on lineage diversity
Genomic sequencing of SARS-CoV-2 in Rwanda reveals the importance of incoming travelers on lineage diversity
Genomic surveillance of SARS-CoV-2 can inform regional transmission dynamics and inform public health interventions. Here, the authors sequence ~200 samples from Rwanda, identify shifts in predominating strains from May 2020 to February 2021, and infer geographic origins.
- Yvan Butera
- , Enatha Mukantwari
- & Nadine Rujeni
-
Article
| Open Access Global phylogenomic analyses of Mycobacterium abscessus provide context for non cystic fibrosis infections and the evolution of antibiotic resistance
Global phylogenomic analyses of Mycobacterium abscessus provide context for non cystic fibrosis infections and the evolution of antibiotic resistance
Mycobacterium abscessus is an emerging infection that usually affects patients with structural lung diseases such as cystic fibrosis (CF). Here, the authors use phylogenetic analyses to demonstrate close relationships between isolates from CF and non-CF patients and identify antibiotic resistance markers.
- Ryan A. Bronson
- , Chhavi Gupta
- & Keira A. Cohen
-
Article
| Open Access Tracking the introduction and spread of SARS-CoV-2 in coastal Kenya
Tracking the introduction and spread of SARS-CoV-2 in coastal Kenya
SARS-CoV-2 was first detected in Kenya in March 2020 and there was evidence of local transmission in the following months. Here, the authors characterise the early stages of the epidemic in coastal Kenya using phylogenetics and find evidence of multiple strain importations from international points of entry.
- George Githinji
- , Zaydah R. de Laurent
- & Charles N. Agoti
-
Article
| Open Access Population genomics of apricots unravels domestication history and adaptive events
Population genomics of apricots unravels domestication history and adaptive events
The evolutionary and domestication history of apricots is poorly understood. Here, the authors provide four apricot high-quality genome assemblies, the genomes of 578 accessions from natural and cultivated populations, and show that Chinese and European apricots constitute two different gene pools, resulting from independent domestication events.
- Alexis Groppi
- , Shuo Liu
- & Véronique Decroocq
-
Article
| Open Access Population genomics provides insights into the evolution and adaptation to humans of the waterborne pathogen Mycobacterium kansasii
Population genomics provides insights into the evolution and adaptation to humans of the waterborne pathogen Mycobacterium kansasii
Mycobacterium kansasii can cause serious pulmonary disease. Here, the authors present a population genomics analysis of 358 environmental and clinical isolates from around the world, supporting the idea that municipal water is a main source of infection, and shedding light into the pathogen’s diversity and adaptation to the human host.
- Tao Luo
- , Peng Xu
- & Qian Gao
-
Article
| Open Access Accommodating individual travel history and unsampled diversity in Bayesian phylogeographic inference of SARS-CoV-2
Accommodating individual travel history and unsampled diversity in Bayesian phylogeographic inference of SARS-CoV-2
Spatiotemporal sampling gaps in existing pathogen genomic data limits their use in understanding epidemiological patterns. Here, the authors apply a phylogeographic approach with SARS-CoV-2 genomes to accurately reproduce pathogen spread by accounting for spatial biases and travel history of the individual.
- Philippe Lemey
- , Samuel L. Hong
- & Marc A. Suchard
-
Article
| Open Access Origin and adaptation to high altitude of Tibetan semi-wild wheat
Origin and adaptation to high altitude of Tibetan semi-wild wheat
Mechanism of high altitude adaptation of wheat remains unknown. Here, the authors assemble the draft genome of a Tibetan semi-wild wheat accession and resequence 245 wheat accessions to reveal that Tibetan semi-wild wheat has been de-domesticated from local landraces to adapt to high altitude.
- Weilong Guo
- , Mingming Xin
- & Qixin Sun
-
Article
| Open Access Inference of person-to-person transmission of COVID-19 reveals hidden super-spreading events during the early outbreak phase
Inference of person-to-person transmission of COVID-19 reveals hidden super-spreading events during the early outbreak phase
Although SARS-CoV-2 has spread rapidly, the contribution of super-spreading events to transmission is unclear. Here, the authors show that the number of secondary infections arising from an individual infection in the early phase of the outbreak was highly skewed, indicating that super-spreading events occurred.
- Liang Wang
- , Xavier Didelot
- & Yuhai Bi
-
Article
| Open Access Discovery of EMRE in fungi resolves the true evolutionary history of the mitochondrial calcium uniporter
Discovery of EMRE in fungi resolves the true evolutionary history of the mitochondrial calcium uniporter
The mitochondrial calcium uptake system, crucial for cellular processes, evolved in ancient eukaryotes. Here, authors perform a phylogenomic analysis across 1,156 eukaryotes, and show that previously identified animal and fungal genes in this system originated from an ancestral duplication.
- Alexandros A. Pittis
- , Valerie Goh
- & Toni Gabaldón
-
Article
| Open Access Undinarchaeota illuminate DPANN phylogeny and the impact of gene transfer on archaeal evolution
Undinarchaeota illuminate DPANN phylogeny and the impact of gene transfer on archaeal evolution
The evolutionary relationships within Archaea remain unresolved. Here, the authors used genomic approaches to study the Undinarchaeota, a previously uncharacterized clade of DPANN, shed light on their position in an updated archaeal phylogeny and illuminate the history of archaeal genome evolution.
- Nina Dombrowski
- , Tom A. Williams
- & Anja Spang
-
Article
| Open Access Nested whole-genome duplications coincide with diversification and high morphological disparity in Brassicaceae
Nested whole-genome duplications coincide with diversification and high morphological disparity in Brassicaceae
As one of the most successful angiosperm clades with ~4000 species, the mustard family has been diversifying into many evolutionary lineages. Here, the authors construct plastid-based phylogeny and show nested whole-genome duplications coincide with diversification and high morphological disparity.
- Nora Walden
- , Dmitry A. German
- & Marcus A. Koch
-
Article
| Open Access Genome-wide analysis of Cushion willow provides insights into alpine plant divergence in a biodiversity hotspot
Genome-wide analysis of Cushion willow provides insights into alpine plant divergence in a biodiversity hotspot
Exceptional alpine plant diversity exists in the Hengduan Mountains. Here, through genome assembly and population genomics studies, the authors find notable intraspecific divergence among Cushion willow populations isolated by the sky island-like habitats and consider it contributes to speciation and biodiversity.
- Jia-hui Chen
- , Yuan Huang
- & Hang Sun
-
Article
| Open Access The chromosome-scale reference genome of black pepper provides insight into piperine biosynthesis
The chromosome-scale reference genome of black pepper provides insight into piperine biosynthesis
Black pepper (Piper nigrum) belongs to the long-isolated lineage of basal angiosperm and its fruit has been used for food spice and phytomedicines for thousands of years. Here, the authors assemble the reference genome of this species and analyze gene families associated with piperine biosynthesis.
- Lisong Hu
- , Zhongping Xu
- & Shuangxia Jin
-
Article
| Open Access Increasing species sampling in chelicerate genomic-scale datasets provides support for monophyly of Acari and Arachnida
Increasing species sampling in chelicerate genomic-scale datasets provides support for monophyly of Acari and Arachnida
Morphological and molecular data have led to conflicting phylogenetic hypotheses for the Chelicerata. Here, the authors reconstruct the phylogeny of the Chelicerata using genomic-scale datasets, finding evidence for a monophyletic Acari and a single terrestrialisation of Arachnida.
- Jesus Lozano-Fernandez
- , Alastair R. Tanner
- & Davide Pisani
-
Article
| Open Access Genomic analysis on pygmy hog reveals extensive interbreeding during wild boar expansion
Genomic analysis on pygmy hog reveals extensive interbreeding during wild boar expansion
The pygmy hog (Porcula salvania), now highly endangered and restricted in a small region at the southern foothills of the Himalaya, is the only suid species in mainland Eurasia that outlived the expansion of wild boar (Sus scrofa). Here, the authors analyze genomes of pygmy hog and related suid species, and identify signals of introgression among these species.
- Langqing Liu
- , Mirte Bosse
- & Ole Madsen
-
Article
| Open Access Genome sequences of two diploid wild relatives of cultivated sweetpotato reveal targets for genetic improvement
Genome sequences of two diploid wild relatives of cultivated sweetpotato reveal targets for genetic improvement
Sweetpotato is an important food security crop providing rich source of macro- and micronutrients including carbohydrates and vitamins. Here, the authors assemble of the two diploid relatives of cultivated sweetpotato and identify genes and alleles associated with carotenoid biosynthesis from breeding lines.
- Shan Wu
- , Kin H. Lau
- & Zhangjun Fei
-
Article
| Open Access Gene synthesis allows biologists to source genes from farther away in the tree of life
Gene synthesis allows biologists to source genes from farther away in the tree of life
Gene synthesis has expanded the ability to modify and create DNA sequences, with implications for biosurveillance. The authors use machine learning and codon theory to identify synthetic genes in Addgene data, and show that synthesis accelerates human-directed gene transfer across the tree of life.
- Aditya M. Kunjapur
- , Philipp Pfingstag
- & Neil C. Thompson
-
Article
| Open Access Human adaptation and population differentiation in the light of ancient genomes
Human adaptation and population differentiation in the light of ancient genomes
Detecting the targets of positive selection in the human genome is challenging. Here, the authors combine modern and ancient genomes to show that alleles strongly differentiated between Africans and Europeans mediated local adaptation in European populations, and were mostly contributed by ancient hunter-gatherers.
- Felix M. Key
- , Qiaomei Fu
- & Aida M. Andrés
-
Article
| Open Access Unique features of a global human ectoparasite identified through sequencing of the bed bug genome
Unique features of a global human ectoparasite identified through sequencing of the bed bug genome
The bed bug, Cimex lectularius, is a ubiquitous human ectoparasite with global distribution. Here, the authors sequence the genome of the bed bug and identify reductions in chemosensory genes, expansion of genes associated with blood digestion and genes linked to pesticide resistance.
- Joshua B. Benoit
- , Zach N. Adelman
- & Stephen Richards
-
Article
| Open Access Genome assembly and geospatial phylogenomics of the bed bug Cimex lectularius
Genome assembly and geospatial phylogenomics of the bed bug Cimex lectularius
The common bedbug is a pest for humans, yet its molecular biology is poorly understood. Here, the authors sequence the common bedbug genome and profile gene expression across all life stages to show major changes in gene expression after feeding on human blood.
- Jeffrey A. Rosenfeld
- , Darryl Reeves
- & Christopher E. Mason
-
Article
| Open Access Phylogenomic and biogeographic reconstruction of the Trichinella complex
Phylogenomic and biogeographic reconstruction of the Trichinella complex
Trichinellosis is a globally important food-borne disease caused by roundworms of the Trichinella complex. Here the authors present genomic sequences representing all 12 recognized Trichinellaspecies and genotypes, and reconstruct their phylogeny and biogeography.
- Pasi K. Korhonen
- , Edoardo Pozio
- & Robin B. Gasser
-
Article
| Open Access Phylodynamics of H1N1/2009 influenza reveals the transition from host adaptation to immune-driven selection
Phylodynamics of H1N1/2009 influenza reveals the transition from host adaptation to immune-driven selection
Influenza A H1N1/2009 virus emerged from swine and rapidly replaced the seasonal H1N1 virus. Here, the authors show that natural selection acting on H1N1/2009 after introduction into humans was driven by adaptation to the new host but later selection has been driven by immunological escape.
- Yvonne C. F. Su
- , Justin Bahl
- & Gavin J. D. Smith
-
Article |
 A primase subunit essential for efficient primer synthesis by an archaeal eukaryotic-type primase
A primase subunit essential for efficient primer synthesis by an archaeal eukaryotic-type primase
Archaea encode an eukaryotic-type primase comprising a catalytic subunit PriS and a noncatalytic subunit PriL. Here, the authors identify in an archaeon Sulfolobus solfataricus an essential noncatalytic subunit of primase, PriX, that forms PriSLX trimer and increases the efficiency of primer synthesis.
- Bing Liu
- , Songying Ouyang
- & Li Huang
-
Article
| Open Access Evolutionary analysis of the female-specific avian W chromosome
Evolutionary analysis of the female-specific avian W chromosome
The evolution of non-recombining chromosomes is poorly understood. Here, the authors sequence the collared flycatcher female-specific W chromosome and show nonrandom survival of genes during W chromosome degeneration which is due to selection for maintaining gene dose and expression levels of essential genes.
- Linnéa Smeds
- , Vera Warmuth
- & Hans Ellegren
-
Article |
 Biology of a widespread uncultivated archaeon that contributes to carbon fixation in the subsurface
Biology of a widespread uncultivated archaeon that contributes to carbon fixation in the subsurface
Research on microbes that inhabit the Earth's subsurface is mostly based on metagenomic information only. Here, Probst et al. combine metagenomics with ultrastructural and functional analyses to study the biology of a group of uncultivated subsurface archaea, the SM1 Euryarchaeon lineage.
- Alexander J. Probst
- , Thomas Weinmaier
- & Christine Moissl-Eichinger
-
Article |
 The plastid ancestor originated among one of the major cyanobacterial lineages
The plastid ancestor originated among one of the major cyanobacterial lineages
Chloroplasts originate from endosymbiosis between a cyanobacterium and a eukaryotic mitochondriate ancestor. Here, the authors show that the plastid ancestor is related to a cyanobacterial lineage that include N2-fixing filamentous cyanobacteria and species with specialized nitrogen-fixing cells.
- Jesús A. G. Ochoa de Alda
- , Rocío Esteban
- & Jean Houmard
-
Article
| Open Access A robust SNP barcode for typing Mycobacterium tuberculosis complex strains
A robust SNP barcode for typing Mycobacterium tuberculosis complex strains
Genetic variation in Mycobacterium tuberculosiscomplex (MTBC) bacteria is responsible for differences in factors such as virulence and transmissibility. Here, the authors analyse the genomes of 1,601 MTBC isolates from diverse geographic locations and identify 62 SNPs that may be used to resolve lineages and sublineages of these strains.
- Francesc Coll
- , Ruth McNerney
- & Taane G. Clark
-
Article |
 Dynamic reassortments and genetic heterogeneity of the human-infecting influenza A (H7N9) virus
Dynamic reassortments and genetic heterogeneity of the human-infecting influenza A (H7N9) virus
H7N9 influenza A viruses capable of infecting humans have recently emerged in China. Here, the authors show that these viruses remain genetically diverse, suggesting that they are still in the process of adapting to human hosts.
- Lunbiao Cui
- , Di Liu
- & George F. Gao
-
Article
| Open Access Extraordinary phylogenetic diversity and metabolic versatility in aquifer sediment
Extraordinary phylogenetic diversity and metabolic versatility in aquifer sediment
Turnover of sediment organic matter contributes to global carbon cycling, yet the microorganisms involved are largely unknown. Castelleet al.reveal that an aquifer sediment core hosts a ‘zoo’ of organisms, including representatives of a previously undescribed phylum (Zixibacteria).
- Cindy J. Castelle
- , Laura A. Hug
- & Jillian F. Banfield

 HIV transmission dynamics and population-wide drug resistance in rural South Africa
HIV transmission dynamics and population-wide drug resistance in rural South Africa