Featured
-
-
Article
| Open Access A chromatinized origin reduces the mobility of ORC and MCM through interactions and spatial constraint
A chromatinized origin reduces the mobility of ORC and MCM through interactions and spatial constraint
Here the authors investigate the impact of chromatinizing origins of replication on ORC and MCM is at the single-molecule level. They find mobility of ORC reduced, but not its binding to the origin. MCM is both efficiently recruited and spatially confined to the origin.
- Humberto Sánchez
- , Zhaowei Liu
- & Nynke H. Dekker
-
Article
| Open Access NFIB facilitates replication licensing by acting as a genome organizer
NFIB facilitates replication licensing by acting as a genome organizer
The precise rule of replication origin selection and activation in metazoans remains unclear. Here, the authors identify NFIB as a genome organizer and replication pioneer by facilitating nucleosome remodeling and chromatin assembly of the pre-RC.
- Wenting Zhang
- , Yue Wang
- & Yongfeng Shang
-
Article
| Open Access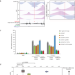 Dimeric G-quadruplex motifs-induced NFRs determine strong replication origins in vertebrates
Dimeric G-quadruplex motifs-induced NFRs determine strong replication origins in vertebrates
A program regulating replication origins ensures the exact duplication of vertebrate genomes. The authors identify a combination of guanine-rich motifs, known to form secondary DNA structures, which are sufficient to assemble efficient replication origins.
- Jérémy Poulet-Benedetti
- , Caroline Tonnerre-Doncarli
- & Marie-Noëlle Prioleau
-
Article
| Open Access Nucleosome-directed replication origin licensing independent of a consensus DNA sequence
Nucleosome-directed replication origin licensing independent of a consensus DNA sequence
Most eukaryotes do not use a consensus DNA sequence as binding sites for the origin recognition complex (ORC) to initiate DNA replication, however budding yeast do. Here the authors show S. cerevisiae ORC can bind nucleosomes near nucleosome-free regions and recruit replicative helicases to form a pre-replication complex independent of the DNA sequence.
- Sai Li
- , Michael R. Wasserman
- & Shixin Liu
-
Article
| Open Access A plasmid system with tunable copy number
A plasmid system with tunable copy number
The range of available copy numbers for cloning vectors is largely restricted to the handful of ORIs that have been isolated from plasmids found in nature. Here the authors introduce a plasmid system that allow for the continuous, finely-tuned control of plasmid copy number between 1 and 800 copies per cell.
- Miles V. Rouches
- , Yasu Xu
- & Guillaume Lambert
-
Article
| Open Access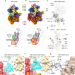 A mechanism of origin licensing control through autoinhibition of S. cerevisiae ORC·DNA·Cdc6
A mechanism of origin licensing control through autoinhibition of S. cerevisiae ORC·DNA·Cdc6
Cryo-EM structures of S. cerevisiae ORC bound to DNA and Cdc6 reveal an autoinhibited conformation and suggest a mechanism of origin licensing control in response to CDK phosphorylation.
- Jan Marten Schmidt
- , Ran Yang
- & Franziska Bleichert
-
Article
| Open Access Unwinding of a DNA replication fork by a hexameric viral helicase
Unwinding of a DNA replication fork by a hexameric viral helicase
Replicative hexameric helicases are fundamental components of replisomes. Here the authors resolve a cryo-EM structure of the E1 helicase from papillomavirus bound to a DNA replication fork, providing insights into the mechanism of DNA unwinding by these hexameric enzymes.
- Abid Javed
- , Balazs Major
- & Elena V. Orlova
-
Article
| Open Access Dynamics of replication origin over-activation
Dynamics of replication origin over-activation
DNA replication processes are often dysregulated in cancer. Here the authors analyse DNA synthesis patterns in cancer cells undergoing partial genome re-replication to reveal that re-replication exhibits aberrant replication fork dynamics and a skewed distribution of replication initiation that over-duplicates early-replicating genomic regions.
- Haiqing Fu
- , Christophe E. Redon
- & Mirit I. Aladjem
-
Article
| Open Access DNA replication origins retain mobile licensing proteins
DNA replication origins retain mobile licensing proteins
Eukaryotic DNA replication is regulated to ensure copying of the genome (only) once per cell cycle. Here the authors, using optical trapping and confocal microscopy, demonstrate the dynamics of the origin recognition complex and subsequent intermediates that lead up to the loading of an MCM helicase onto DNA.
- Humberto Sánchez
- , Kaley McCluskey
- & Nynke H. Dekker
-
Article
| Open Access Evolution of DNA replication origin specification and gene silencing mechanisms
Evolution of DNA replication origin specification and gene silencing mechanisms
Contrary to most eukaryotes that lack sequence-specific origins of replication, S. cerevisiae origins are defined by specific DNA sequence motifs. Here the authors reveal that multiple subunits of ORC, including Orc2 and Orc4, contribute to the sequence-specificity of origins in S. cerevisiae.
- Y. Hu
- , A. Tareen
- & B. Stillman
-
Article
| Open Access Involvement of G-quadruplex regions in mammalian replication origin activity
Involvement of G-quadruplex regions in mammalian replication origin activity
Origins of replications are associated with potential G quadruplexes forming structures (G4s). Here the authors reveal the functional role of G4 elements in DNA replication initiation.
- Paulina Prorok
- , Marie Artufel
- & Marcel Méchali
-
Article
| Open Access The ORC ubiquitin ligase OBI1 promotes DNA replication origin firing
The ORC ubiquitin ligase OBI1 promotes DNA replication origin firing
DNA replication is initiated at defined genomic sites called origins of replication following ORC pre-replicative complex assembly. Here the authors identify a protein ubiquitylating ORC that is involved in origin activation and may act as a selector of origins to be fired.
- Philippe Coulombe
- , Joelle Nassar
- & Marcel Méchali
-
Article
| Open Access Allele-specific analysis of DNA replication origins in mammalian cells
Allele-specific analysis of DNA replication origins in mammalian cells
DNA sequences contribute to the location and timing of replication origin firings. Here by allele-specific analysis, the authors show that replication asynchrony is associated with small cumulative variations in the initiation efficiency of origins, rather than with the activation of dormant origins.
- Boris Bartholdy
- , Rituparna Mukhopadhyay
- & Eric E. Bouhassira

 Dormant origin firing promotes head-on transcription-replication conflicts at transcription termination sites in response to BRCA2 deficiency
Dormant origin firing promotes head-on transcription-replication conflicts at transcription termination sites in response to BRCA2 deficiency