Featured
-
-
Article
| Open Access ProTInSeq: transposon insertion tracking by ultra-deep DNA sequencing to identify translated large and small ORFs
ProTInSeq: transposon insertion tracking by ultra-deep DNA sequencing to identify translated large and small ORFs
Identifying small proteins is challenging. ProTInSeq uses modified transposons to express markers inserted in-frame to protein-coding genes. This method identifies 153 unannotated small proteins in M. pneumoniae and additional proteomic information.
- Samuel Miravet-Verde
- , Rocco Mazzolini
- & Luis Serrano
-
Article
| Open Access Alt-RPL36 downregulates the PI3K-AKT-mTOR signaling pathway by interacting with TMEM24
Alt-RPL36 downregulates the PI3K-AKT-mTOR signaling pathway by interacting with TMEM24
Many alternative ORFs are co-encoded with characterized proteins, but their function is often not understood. Here, the authors discover that ribosomal protein L36 is co-encoded with alternative protein, which they identify as an upstream regulator of PI3K-AKT-mTOR signaling.
- Xiongwen Cao
- , Alexandra Khitun
- & Sarah A. Slavoff
-
Article
| Open Access Full-length transcriptome reconstruction reveals a large diversity of RNA and protein isoforms in rat hippocampus
Full-length transcriptome reconstruction reveals a large diversity of RNA and protein isoforms in rat hippocampus
It is challenging to characterize diverse transcript isoforms by short-read sequencing. Here the authors report full-length transcriptomes in rat hippocampus by hybrid-sequencing, predict isoform-specific translational status, and reconstruct open reading frames validated by mass spectrometry.
- Xi Wang
- , Xintian You
- & Wei Chen
-
Article |
 Maintenance of protein synthesis reading frame by EF-P and m1G37-tRNA
Maintenance of protein synthesis reading frame by EF-P and m1G37-tRNA
Slippery mRNA sequences CC[C/U]-[C/U] are prone to +1 frameshift (+1FS) errors during mRNA translation. Here, the authors show that +1FS errors occur predominantly when CC[C/U]-[C/U] are placed at the second sense codon, and that error suppression requires m1G37-tRNA and the translation factor EF-P.
- Howard B. Gamper
- , Isao Masuda
- & Ya-Ming Hou

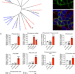 Illuminating the function of the orphan transporter, SLC22A10, in humans and other primates
Illuminating the function of the orphan transporter, SLC22A10, in humans and other primates