Featured
-
-
Letter |
 Phosphorylation of MLL by ATR is required for execution of mammalian S-phase checkpoint
Phosphorylation of MLL by ATR is required for execution of mammalian S-phase checkpoint
Cell cycle checkpoints, such as the S-phase checkpoint, delay cell division to give the cell time to repair any damaged DNA. Here it is shown that the MLL gene — frequently disrupted in leukaemia — is part of the S-phase checkpoint. When DNA is damaged, MLL is phosphorylated by the ATR protein, causing MLL to accumulate on chromatin and methylate histone H3 on lysine 4. This delays DNA replication. MLL translocations, such as those that occur in leukaemia, disrupt this pathway and cause genomic instability.
- Han Liu
- , Shugaku Takeda
- & James J.-D. Hsieh
-
Letter |
 Mechanism of the ATP-dependent DNA end-resection machinery from Saccharomyces cerevisiae
Mechanism of the ATP-dependent DNA end-resection machinery from Saccharomyces cerevisiae
When double-strand breaks occur in DNA, the broken ends must undergo processing to prepare them for repair. Here, and in an accompanying study, this processing reaction has now been replicated in vitro using yeast proteins. Processing minimally requires the activities of a helicase, a nuclease and a single-strand-binding protein, although the reaction is enhanced by the addition of three factors that help to target the core complex and stimulate the unwinding activity.
- Hengyao Niu
- , Woo-Hyun Chung
- & Patrick Sung
-
Letter |
 Genome-wide measurement of RNA secondary structure in yeast
Genome-wide measurement of RNA secondary structure in yeast
Experimental determination of the secondary structure of RNA molecules has usually been carried out on a case-by-case basis. Now, however, a deep-sequencing approach has been used to profile the secondary structure of 3,000 distinct messenger RNA transcripts from Saccharomyces cerevisiae. The results provide interesting hints about the role of secondary structure in protein translation, and set the stage for the examination of how such structures can change in response to environmental conditions.
- Michael Kertesz
- , Yue Wan
- & Eran Segal
-
Letter |
 DNA end resection by Dna2–Sgs1–RPA and its stimulation by Top3–Rmi1 and Mre11–Rad50–Xrs2
DNA end resection by Dna2–Sgs1–RPA and its stimulation by Top3–Rmi1 and Mre11–Rad50–Xrs2
When double-strand breaks occur in DNA, the broken ends must undergo processing to prepare them for repair. Here, and in an accompanying study, this processing reaction has now been replicated in vitro using yeast proteins. Processing minimally requires the activities of a helicase, a nuclease and a single-strand-binding protein, although the reaction is enhanced by the addition of three factors that help to target the core complex and stimulate the unwinding activity.
- Petr Cejka
- , Elda Cannavo
- & Stephen C. Kowalczykowski
-
Letter |
 Selectivity mechanism of the nuclear pore complex characterized by single cargo tracking
Selectivity mechanism of the nuclear pore complex characterized by single cargo tracking
Nuclear pore complexes selectively transport cargos across the nuclear envelope. Here, a nuclear transport assay has been developed that allows the movement of single cargo proteins to be followed in real time. A succession of transport substeps is observed, and the NPC is found to be functionally asymmetric to importing cargos. The study provides insight into the mechanism of selective transport through the NPC.
- Alan R. Lowe
- , Jake J. Siegel
- & Jan T. Liphardt
-
Letter |
 Identification of a quality-control mechanism for mRNA 5′-end capping
Identification of a quality-control mechanism for mRNA 5′-end capping
Following their synthesis, eukaryotic messenger RNAs have a 7-methylguanosine cap added to their 5′ ends to protect the mRNAs from degradation. Here it is shown that, in vitro and in yeast, caps lacking a methyl group are recognized by the Rai1 protein, which clips off the incomplete cap. The data provide evidence that Rai1 is part of a quality-control mechanism that monitors, and promotes the digestion of, aberrant mRNAs that might arise during stress conditions.
- Xinfu Jiao
- , Song Xiang
- & Megerditch Kiledjian
-
Letter |
 Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification
Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification
TET1 is an enzyme that catalyses the conversion of 5-methylcytosine of DNA to 5-hydroxymethylcytosine, raising the possibility that it is involved in mediating DNA demethylation. These authors show that Tet1 is involved in mouse embryonic stem cell maintenance and specification of the inner cell mass. It is required to maintain both the expression of Nanog in embryonic stem cells and the Nanog promoter in a hypomethylated state, supporting a role for Tet1 in regulating DNA methylation.
- Shinsuke Ito
- , Ana C. D’Alessio
- & Yi Zhang
-
Article |
 Heterochromatin silencing of p53 target genes by a small viral protein
Heterochromatin silencing of p53 target genes by a small viral protein
Adenovirus E1B-55k targets transcription factor p53 for degradation and is thought to be critical for p53 inactivation during adenovirus replication. Indeed, mutant viruses lacking E1B-55k have been tested as viral cancer therapies selective for p53-positive tumours. These authors find another adenoviral protein, E4-ORF3, to inactivate p53 independently of E1B-55k by means of a chromatin-silencing mechanism that prevents access of p53 to its DNA target sites.
- Conrado Soria
- , Fanny E. Estermann
- & Clodagh C. O’Shea
-
Letter |
 NRMT is an α-N-methyltransferase that methylates RCC1 and retinoblastoma protein
NRMT is an α-N-methyltransferase that methylates RCC1 and retinoblastoma protein
α-N-methylation is an unusual post-translational modification in which the amino-terminal residues of proteins are methylated. One example is the Ran guanine nucleotide-exchange factor, RCC1, which requires methylation for its association with chromatin. These authors describe the first α-N-methyltransferase, named N-terminal RCC1 methyltransferase (NRMT). They identify the NRMT recognition sequence and several new methylation targets, and demonstrate the importance of α-N-methylation for normal bipolar spindle formation and chromosome segregation.
- Christine E. Schaner Tooley
- , Janusz J. Petkowski
- & Ian G. Macara
-
Article |
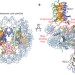 Structure of RCC1 chromatin factor bound to the nucleosome core particle
Structure of RCC1 chromatin factor bound to the nucleosome core particle
The small GTPase Ran regulates nuclear transport and cell division by creating a gradient of RanGTP around chromosomes. The RCC1 protein recruits Ran to nucleosomes and activates Ran's nucleotide exchange activity. Here, the crystal structure of the complex between RCC1 and the nucleosome core particle is revealed. It provides an atomic view of how a chromatin protein interacts with the histone and DNA components of the nucleosome.
- Ravindra D. Makde
- , Joseph R. England
- & Song Tan
-
Letter |
 The structure of (CENP-A–H4)2 reveals physical features that mark centromeres
The structure of (CENP-A–H4)2 reveals physical features that mark centromeres
The centromeres of chromosomes are specified epigenetically, and the histone H3 variant CENP-A is assembled into the chromatin of all active centromeres. Here, the crystal structure of CENP-A in a tetrameric complex with histone H4 reveals the physical features of centromeric chromatin. CENP-A seems to mark the centromere by altering nucleosome structure from within its folded histone core.
- Nikolina Sekulic
- , Emily A. Bassett
- & Ben E. Black
-
Letter |
 Phosphorylation of the CPC by Cdk1 promotes chromosome bi-orientation
Phosphorylation of the CPC by Cdk1 promotes chromosome bi-orientation
The chromosomal passenger complex (CPC) coordinates several processes during cell division, including chromosome bi-orientation and cytokinesis, and its proper localization is crucial. These authors provide a mechanism for its localization to the inner centromere. Cdk1–cyclin-B-dependent phosphorylation of the CPC promotes binding to shugoshin, which the authors define as a conserved centromeric adaptor of the CPC. This mechanism is conserved between fission yeast and human cells and highlights a crucial role of Cdk1–cyclin B in chromosome bi-orientation.
- Tatsuya Tsukahara
- , Yuji Tanno
- & Yoshinori Watanabe
-
Article |
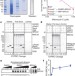 Purified human BRCA2 stimulates RAD51-mediated recombination
Purified human BRCA2 stimulates RAD51-mediated recombination
The two hereditary breast cancer susceptibility genes, BRCA1 and BRCA2, have roles in responding to DNA damage. When they are mutated or absent, genomic instability, a contributory factor to cancer development, results. Studies of BRCA2 have been hampered by its large size, which makes purification of the full-length protein challenging. These authors report the first in vitro characterization of full-length BRCA2 and delineate the different ways by which BRCA2 facilitates RAD51-mediated homologous recombination.
- Ryan B. Jensen
- , Aura Carreira
- & Stephen C. Kowalczykowski
-
Letter |
 Trans-acting small RNA determines dominance relationships in Brassica self-incompatibility
Trans-acting small RNA determines dominance relationships in Brassica self-incompatibility
A diploid organism has two copies of each gene, one inherited from each parent. The expression levels of the two alleles can be biased by dominant/recessive relationships. In Brassica, self-incompatibility in pollen is determined by dominance relationships between the two alleles of the gene SP11; the recessive allele is methylated and hence silenced. Here it is shown that such methylation is controlled by a small non-coding RNA encoded in the flanking region of the dominant allele.
- Yoshiaki Tarutani
- , Hiroshi Shiba
- & Seiji Takayama
-
Article |
 Non-canonical inhibition of DNA damage-dependent ubiquitination by OTUB1
Non-canonical inhibition of DNA damage-dependent ubiquitination by OTUB1
When double-strand breaks occur in eukaryotic DNA, the chromatin that protects and organizes the genome must be removed from the vicinity of the break to allow repair factors to bind. Such chromatin displacement involves the addition of ubiquitin groups to histone proteins near the break by the ubiquitin ligases RNF8 and RNF168. Here it is shown that the enzyme OTUB1 prevents RNF168-dependent poly-ubiquitination. Pharmacological targeting of this process might enhance the DNA damage response.
- Shinichiro Nakada
- , Ikue Tai
- & Daniel Durocher
-
News & Views |
 Blocking ubiquitin transfer
Blocking ubiquitin transfer
The protein OTUB1 inhibits DNA repair without using its enzymatic activity. Instead, it sequesters a protein that is required for the assembly of certain forms of ubiquitin chain, which function as key signals during repair.
- April Rose
- & Christian Schlieker
-
Article |
 Mediator and cohesin connect gene expression and chromatin architecture
Mediator and cohesin connect gene expression and chromatin architecture
Gene activation may involve the formation of a DNA loop that connects enhancer-bound transcription factors with the transcription apparatus at the core promoter. But this process is not well understood. Here, two proteins, mediator and cohesin, are shown to connect the enhancers and core promoters of active genes in embryonic stem cells. These proteins seem to generate cell-type-specific DNA loops linked to the gene expression program of each cell.
- Michael H. Kagey
- , Jamie J. Newman
- & Richard A. Young
-
Letter |
 Dynamics and mechanism of repair of ultraviolet-induced (6–4) photoproduct by photolyase
Dynamics and mechanism of repair of ultraviolet-induced (6–4) photoproduct by photolyase
The repair enzyme (6–4) photolyase uses light energy to cleave the ultraviolet-induced bond between pyrimidine dimers. These authors use ultrafast spectroscopy to examine the detailed electron and proton movements during the catalytic photocycle. Histidine 364 is identified as the crucial residue involved in the rate-limiting step.
- Jiang Li
- , Zheyun Liu
- & Dongping Zhong
-
Article |
 Mammalian microRNAs predominantly act to decrease target mRNA levels
Mammalian microRNAs predominantly act to decrease target mRNA levels
MicroRNAs are known to affect the levels of both messenger RNA (mRNA) and protein. But as protein production is dependent on the presence of mRNA, it was not clear what the relative contributions of microRNA-mediated mRNA cleavage and translational repression were. These authors have parsed out the two mechanisms, and unexpectedly find that microRNAs function primarily by affecting mRNA levels rather than their translation. This suggests a reassessment of many previous conclusions is necessary.
- Huili Guo
- , Nicholas T. Ingolia
- & David P. Bartel
-
Letter |
 Structure of the LexA–DNA complex and implications for SOS box measurement
Structure of the LexA–DNA complex and implications for SOS box measurement
Normally, expression of bacterial DNA damage repair genes is repressed by the binding of LexA protein to SOS ‘boxes’ in their operators. DNA damage activates the RecA protein, which promotes autocleavage of LexA such that its repression is relieved and repair proteins are expressed. These authors solve several structures of LexA dimer bound to SOS box DNA, and find that the orientation of the DNA-binding wings can account for the strict intersite spacing.
- Adrianna P. P. Zhang
- , Ying Z. Pigli
- & Phoebe A. Rice
-
Research Highlights |
Genetics: Long and the short of it
-
Letter |
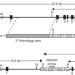 Production of p53 gene knockout rats by homologous recombination in embryonic stem cells
Production of p53 gene knockout rats by homologous recombination in embryonic stem cells
The rat is a animal model widely used for studying human physiology and disease, but functional genomics and genetic research have been stifled by the limited availability of gene targeting tools. These authors have established gene targeting by homologous recombination in rat embryonic stem cells, and have generated p53 gene knockout rats for the first time.
- Chang Tong
- , Ping Li
- & Qi-Long Ying
-
Letter |
 OncomiR addiction in an in vivo model of microRNA-21-induced pre-B-cell lymphoma
OncomiR addiction in an in vivo model of microRNA-21-induced pre-B-cell lymphoma
One model for cancer development posits that the proliferating cells in a tumour can become 'addicted' to activating mutations in an oncogene. With the realization that certain microRNAs promote tumorigenesis, it has been proposed that tumours may also become dependent on such 'oncomiRs'. Here, evidence is provided that the gene encoding microRNA-21 is an oncogene, and that in its absence, tumours undergo apoptosis and regress. Thus tumours can indeed become addicted to oncomiRs.
- Pedro P. Medina
- , Mona Nolde
- & Frank J. Slack
-
Letter |
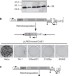 Epigenetic silencing of engineered L1 retrotransposition events in human embryonic carcinoma cells
Epigenetic silencing of engineered L1 retrotransposition events in human embryonic carcinoma cells
The ability of retrotransposons to mobilize and insert into genes presents a challenge to a cell needing to maintain its genomic integrity. These authors have studied retrotransposition in embryonic carcinoma-derived cells. On insertion into DNA, the retrotransposon is quickly silenced, but the retrotransposon-specificity of this process implies that multiple silencing mechanisms may exist. Once cells differentiate, the ability to silence newly introduced retrotransposons is lost but previously inactivated retrotransposons remain inactive.
- Jose L. Garcia-Perez
- , Maria Morell
- & John V. Moran
-
Research Highlights |
Cancer biology: Blood vessel regulator
-
Letter |
 A ribosome-associating factor chaperones tail-anchored membrane proteins
A ribosome-associating factor chaperones tail-anchored membrane proteins
Tail-anchored proteins have a single transmembrane domain at their carboxy termini and are post-translationally targeted to the endoplasmic reticulum via the cytosolic ATPase TRC40. These authors identify a conserved protein complex called Bat3 complex that is recruited to ribosomes, interacts with the transmembrane domain of newly released tail-anchored proteins and transfers them to TRC40 for subsequent targeting to the endoplasmic reticulum.
- Malaiyalam Mariappan
- , Xingzhe Li
- & Ramanujan S. Hegde
-
Letter |
 New class of gene-termini-associated human RNAs suggests a novel RNA copying mechanism
New class of gene-termini-associated human RNAs suggests a novel RNA copying mechanism
In the course of characterizing short RNAs from human cells using single-molecule high-throughput sequencing, these authors identify a new short RNA species. The presence of non-genomically encoded poly(U) residues at their 5' ends implies the existence of an unknown RNA copying mechanism in human cells.
- Philipp Kapranov
- , Fatih Ozsolak
- & Patrice M. Milos
-
Letter |
 Pathogenic LRRK2 negatively regulates microRNA-mediated translational repression
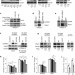
Pathogenic LRRK2 negatively regulates microRNA-mediated translational repression
LRRK2 activity is dysregulated in Parkinson's disease, but how it contributes to the pathogenesis is unknown. These authors show that Drosophila LRRK2 interacts with the Argonaute component (dAgo1) of the RNA-induced silencing complex. This is associated with reduced levels of dAgo1, interaction between phospho-4E-BP1 and hAgo2, and impairment of microRNA-mediated repression. This leads to overexpression of the cell cycle genes e2f1 and dp, and consequent degeneration of dopaminergic neurons.
- Stephan Gehrke
- , Yuzuru Imai
- & Bingwei Lu
-
News |
 Mystery RNA spawns gene-activating peptides
Mystery RNA spawns gene-activating peptides
Short peptides that regulate fruitfly development are produced from 'junk' RNA.
- Heidi Ledford
-
Article |
 Ribosome dynamics and tRNA movement by time-resolved electron cryomicroscopy
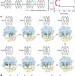
Ribosome dynamics and tRNA movement by time-resolved electron cryomicroscopy
During protein synthesis within the ribosome, transfer RNAs (tRNAs) move sequentially through different sites as their attached amino acids are transferred onto the growing protein chain. Large conformational movements accompany this process. Here, a staggering 1.9 million electron cryomicroscopy images of the ribosome have been processed to visualize these changes. The results reveal that the ribosome functions as a Brownian machine that couples spontaneous changes driven by thermal energy to directed movement.
- Niels Fischer
- , Andrey L. Konevega
- & Holger Stark
-
News & Views |
 Translocation in slow motion
Translocation in slow motion
Time-resolved electron microscopy can capture structural changes in active macromolecular complexes, but detailed imaging is essential. The dynamics of one step in protein synthesis has been deduced from two million images.
- Måns Ehrenberg
-
Letter |
 Regulation of heterochromatic DNA replication by histone H3 lysine 27 methyltransferases
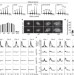
Regulation of heterochromatic DNA replication by histone H3 lysine 27 methyltransferases
DNA replication occurs only once per cell cycle, and numerous pathways prevent re-replication. Here it is shown that mutations in ARABIDOPSIS TRITHORAX-RELATED PROTEIN5 (ATXR5) and ATXR6 — which encode histone methyltransferases — lead to re-replication of specific genomic locations, notably those corresponding to transposons and other repetitive and silenced elements. ATXR5 and ATXR6 are proposed to be components of a pathway that prevents over-replication of heterochromatin in Arabidopsis.
- Yannick Jacob
- , Hume Stroud
- & Steven E. Jacobsen
-
Letter |
 A novel pathway regulates memory and plasticity via SIRT1 and miR-134
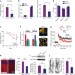
A novel pathway regulates memory and plasticity via SIRT1 and miR-134
The deacetylase SIRT1 has been suggested to function in normal brain physiology, but it is not known whether it participates in higher-order brain functions. These authors demonstrate a role for SIRT1 in synaptic plasticity and memory formation, with activation enhancing synaptic strength and memory formation. These effects were regulated through a post-transcriptional mechanism involving CREB activation and miR-134 production. This interplay represents another mechanism of plasticity regulation with behavioural consequences.
- Jun Gao
- , Wen-Yuan Wang
- & Li-Huei Tsai
-
Letter |
 PHF8 mediates histone H4 lysine 20 demethylation events involved in cell cycle progression
PHF8 mediates histone H4 lysine 20 demethylation events involved in cell cycle progression
These authors show that the JmjC domain-containing protein PHF8 has histone demethylase activity against H4K20me1 and is linked to two distinct events during cell cycle progression. PHF8 is recruited to the promoters of genes involved in the G1–S phase transition, where it removes H4K20me1 and contributes to gene activation, whereas dissociation of PHF8 from chromatin in prophase allows H4K20me1 to accumulate during mitosis.
- Wen Liu
- , Bogdan Tanasa
- & Michael G. Rosenfeld
-
Letter |
 Convergent evolution of chicken Z and human X chromosomes by expansion and gene acquisition
Convergent evolution of chicken Z and human X chromosomes by expansion and gene acquisition
Birds and mammals have distinct sex chromosomes: in birds, males are ZZ and females ZW; in mammals, males are XY and females XX. By sequencing the chicken Z chromosome and comparing it with the human X chromosome, these authors overturn the currently held view that these chromosomes have diverged little from their autosomal progenitors. The Z and X chromosomes seem to have followed convergent evolutionary trajectories, despite evolving with opposite systems of heterogamety.
- Daniel W. Bellott
- , Helen Skaletsky
- & David C. Page
-
Letter |
 Mechanism and regulation of acetylated histone binding by the tandem PHD finger of DPF3b
Mechanism and regulation of acetylated histone binding by the tandem PHD finger of DPF3b
The lysine residues of histone proteins can be acetylated or methylated, with important effects on gene expression. Until recently the protein modules that bind acetyl-lysine have been limited to bromodomains. However, the tandem plant homeodomain (PHD) finger of human DPF3b — which is involved in gene activation — has also been reported to bind to acetylated histones. Here, three-dimensional solution structures of DPF3b offer mechanistic insight into how this protein recognizes acetylation marks.
- Lei Zeng
- , Qiang Zhang
- & Ming-Ming Zhou
-
Letter |
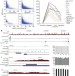 Conserved role of intragenic DNA methylation in regulating alternative promoters
Conserved role of intragenic DNA methylation in regulating alternative promoters
The methylation of DNA in 5′ promoter regions suppresses gene expression, but what is the role of DNA methylation in the bodies of genes? Here, a map of DNA methylation is generated from human brain tissue; it is found that most methylated CpG islands are within intragenic and intergenic regions, rather than within promoters. It is proposed that intragenic methylation regulates the expression of alternative gene transcripts in different tissues and cell types.
- Alika K. Maunakea
- , Raman P. Nagarajan
- & Joseph F. Costello
-
Article |
 Striatal microRNA controls cocaine intake through CREB signalling
Striatal microRNA controls cocaine intake through CREB signalling
Extended cocaine taking triggers several structural and functional changes in the brain that may lead to compulsive drug seeking, but the mechanisms that regulate the process are unclear. Here, a microRNA — miR-212 — is identified that is upregulated in the striatum of rats with a history of extended access to cocaine. The authors suggest that miR-212 protects against the development of compulsive drug taking, and that it may act through the CREB protein, a known regulator of the rewarding effects of cocaine.
- Jonathan A. Hollander
- , Heh-In Im
- & Paul J. Kenny
-
News & Views |
 MicroRNA knocks down cocaine
MicroRNA knocks down cocaine
Cocaine abuse results in increased craving for the drug. But in the long run, cocaine intake induces the expression of a microRNA in the brain, and this seems to limit further drug intake.
- Marina R. Picciotto
-
Letter |
 Subnanometre single-molecule localization, registration and distance measurements
Subnanometre single-molecule localization, registration and distance measurements
These authors have developed a method that enables them to observe single-molecule fluorescent probes with subnanometre precision and accuracy using conventional far-field fluorescence imaging. The improved resolution will enable researchers to characterize single 'molecules' of large, multisubunit biological complexes in biologically relevant environments.
- Alexandros Pertsinidis
- , Yunxiang Zhang
- & Steven Chu
-
Letter |
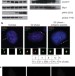 SCFCyclin F controls centrosome homeostasis and mitotic fidelity through CP110 degradation
SCFCyclin F controls centrosome homeostasis and mitotic fidelity through CP110 degradation
Cyclin F is the founding member of the F-box protein family but its functions are unknown; unlike most cyclins, it does not bind or activate cyclin-dependent kinases. Here the authors identify CP110, a protein essential for centrosome duplication, as a substrate of Cyclin F. CP110 and Cyclin F associate on centrioles during the cell cycle, and Cyclin F is proposed to limit centrosome duplication by targeting CP110 for degradation.
- Vincenzo D’Angiolella
- , Valerio Donato
- & Michele Pagano
-
Article |
 Chromatin regulation by Brg1 underlies heart muscle development and disease
Chromatin regulation by Brg1 underlies heart muscle development and disease
Cardiac hypertrophy is associated with a decrease in expression of the adult isoform of the molecular motor myosin heavy chain (α-MHC) and the induction of expression of its fetal isoform (β-MHC). Here the authors reveal the mechanism regulating this switch in expression, which impairs heart function. Cardiac stress in adult hearts reactivates the developmental chromatin-modifying complex Brg1/BAF, which interacts with histone deacetylase and poly (ADP ribose) polymerase to induce a pathological α-MHC-to-β-MHC shift.
- Calvin T. Hang
- , Jin Yang
- & Ching-Pin Chang
-
Letter |
 The male mouse pheromone ESP1 enhances female sexual receptive behaviour through a specific vomeronasal receptor
The male mouse pheromone ESP1 enhances female sexual receptive behaviour through a specific vomeronasal receptor
Although pheromones and their detection by the vomeronasal organ are known to govern social behaviour in mice, specific chemical signals have rarely been linked to selective behavioural responses. Here the authors show that the ESP1 peptide secreted in male tears makes females sexually receptive, and identify its specific vomeronasal receptor and the sex-specific neuronal circuits activated during the behavioural response.
- Sachiko Haga
- , Tatsuya Hattori
- & Kazushige Touhara
-
Letter |
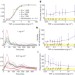 Single-cell NF-κB dynamics reveal digital activation and analogue information processing
Single-cell NF-κB dynamics reveal digital activation and analogue information processing
Multicellular organisms, particularly their immune systems, rely on complex cell-to-cell communication, mediated by signalling molecules that form spatiotemporal concentration gradients. Here, high-throughput microfluidic cell culture and fluorescence microscopy, together with quantitative gene expression analysis and mathematical modelling, have been used to investigate how mammalian cells respond to different levels of TNF-α and signal to NF-κB. Both digital and analogue responses are revealed.
- Savaş Tay
- , Jacob J. Hughey
- & Markus W. Covert
-
Letter |
 Activation of autophagy during cell death requires the engulfment receptor Draper
Activation of autophagy during cell death requires the engulfment receptor Draper
- Christina K. McPhee
- , Mary A. Logan
- & Eric H. Baehrecke
-
Article |
 Structure and mechanism of human DNA polymerase η
Structure and mechanism of human DNA polymerase η
Ultraviolet radiation causes damage to DNA in skin cells, blocking DNA replication and causing mutations that can lead to cancer. One way in which the cell deals with such damage involves specialized DNA polymerases, such as Polη, that can bypass lesions. Here the crystal structure of Polη is presented at four consecutive steps during DNA synthesis through thymine dimers. Polη acts like a 'molecular splint' to stabilize damaged DNA, and accommodates the thymine dimer in an atypically large active site.
- Christian Biertümpfel
- , Ye Zhao
- & Wei Yang
-
Letter |
 Sphingosine-1-phosphate is a missing cofactor for the E3 ubiquitin ligase TRAF2
Sphingosine-1-phosphate is a missing cofactor for the E3 ubiquitin ligase TRAF2
Engagement of the tumour-necrosis factor (TNF) receptor results in the assembly of multi-component signalling complexes by adaptor proteins that include TNF receptor-associated factor 2 (TRAF2). Genetic evidence indicates that TRAF2 is needed for the polyubiquitination of receptor interacting protein 1 (RIP1), but direct evidence has been lacking. Here it is shown that the lipid sphingosine-1-phosphate is a co-factor needed for this ubiquitination activity of TRAF2.
- Sergio E. Alvarez
- , Kuzhuvelil B. Harikumar
- & Sarah Spiegel
-
Article |
 A coding-independent function of gene and pseudogene mRNAs regulates tumour biology
A coding-independent function of gene and pseudogene mRNAs regulates tumour biology
The canonical role of messenger RNA (mRNA) is in protein coding and synthesis. But could mRNAs also have a role that is related to their ability to compete for microRNA binding? Here, the functional relationship between the mRNAs produced by the PTEN tumour suppressor gene and its pseudogene PTENP1 is investigated. The results suggest that pseudogenes have a biological function, in sequestering microRNAs and so affecting their regulation of gene expression.
- Laura Poliseno
- , Leonardo Salmena
- & Pier Paolo Pandolfi
-
News & Views |
 How to accurately bypass damage
How to accurately bypass damage
Ultraviolet radiation can cause cancer through DNA damage — specifically, by linking adjacent thymine bases. Crystal structures show how the enzyme DNA polymerase η accurately bypasses such lesions, offering protection.
- Suse Broyde
- & Dinshaw J. Patel
Browse broader subjects
Browse narrower subjects
- Cell division
- Chromatin
- Chromosomes
- CRISPR-Cas systems
- DNA damage and repair
- DNA metabolism
- DNA recombination
- DNA replication
- Epigenetics
- Non-coding RNAs
- Nuclear organization
- Post-translational modifications
- Protein folding
- Proteolysis
- Proteomics
- Riboswitches
- Ribozymes
- RNA metabolism
- RNAi
- Single-molecule biophysics
- Transcription
- Transcriptomics
- Translation
- Transposition

 HAATI survivors replace canonical telomeres with blocks of generic heterochromatin
HAATI survivors replace canonical telomeres with blocks of generic heterochromatin