Featured
-
-
Review Article
| Open Access Neuropathogenesis-on-chips for neurodegenerative diseases
Neuropathogenesis-on-chips for neurodegenerative diseases
This review focuses on recent advances in on-chip platforms for patient-like in vitro modeling of the pathology of neurodegenerative diseases, including Alzheimer’s, Parkinson’s, and Huntington’s diseases as well as Amyotrophic lateral sclerosis. The authors advocate for broader usage of these human-relevant models in the academic and pharmaceutical fields.
- Sarnai Amartumur
- , Huong Nguyen
- & Chaejeong Heo
-
Article
| Open Access Characterising the RNA-binding protein atlas of the mammalian brain uncovers RBM5 misregulation in mouse models of Huntington’s disease
Characterising the RNA-binding protein atlas of the mammalian brain uncovers RBM5 misregulation in mouse models of Huntington’s disease
RNA-Binding Proteins (RBPs) are critical regulators of RNA biology. Here, the authors describe the Brain-pCLAP methodology, uncover the RBP atlas of the mouse brain and demonstrate the differential binding of the splicing factor RBM5 to Huntington’s disease relevant transcripts in R6/2 mice.
- Meeli Mullari
- , Nicolas Fossat
- & Michael L. Nielsen
-
Article
| Open Access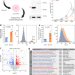 Genome-wide screening in pluripotent cells identifies Mtf1 as a suppressor of mutant huntingtin toxicity
Genome-wide screening in pluripotent cells identifies Mtf1 as a suppressor of mutant huntingtin toxicity
Huntington’s disease is caused by mutation in the HTT gene. Here, the authors screen for suppressors of mutant HTT-induced toxicity, identifying Mtf1. Mtf1 reduced oxidative stress and cell death in stem cells, and motor defects and protein aggregates in mouse models.
- Giorgia Maria Ferlazzo
- , Anna Maria Gambetta
- & Graziano Martello
-
Article
| Open Access CryoET reveals organelle phenotypes in huntington disease patient iPSC-derived and mouse primary neurons
CryoET reveals organelle phenotypes in huntington disease patient iPSC-derived and mouse primary neurons
Huntington’s disease (HD) is a fatal neurodegenerative disorder caused by a genetic mutation in the huntingtin gene (HTT). Here, cryo electron tomography provides insights into the morphology of the cells derived from patients with HD and mouse models of the disease.
- Gong-Her Wu
- , Charlene Smith-Geater
- & Wah Chiu
-
Article
| Open Access Transcriptional vulnerabilities of striatal neurons in human and rodent models of Huntington’s disease
Transcriptional vulnerabilities of striatal neurons in human and rodent models of Huntington’s disease
In human and mouse models of Huntington’s disease, Matsushima, Pineda et al. show, using snRNAsequencing, the two axes defining identities of striatal projection neurons are multiplexed and differentially compromised, calling for distinct therapies.
- Ayano Matsushima
- , Sergio Sebastian Pineda
- & Ann M. Graybiel
-
Article
| Open Access Huntington disease oligodendrocyte maturation deficits revealed by single-nucleus RNAseq are rescued by thiamine-biotin supplementation
Huntington disease oligodendrocyte maturation deficits revealed by single-nucleus RNAseq are rescued by thiamine-biotin supplementation
Here the authors evaluate single cell gene expression from mouse and human Huntington’s disease brains, finding incomplete oligodendrocyte maturation and pathways involved. Treating mice with thiamine/biotin ameliorates molecular pathology.
- Ryan G. Lim
- , Osama Al-Dalahmah
- & Leslie M. Thompson
-
Article
| Open Access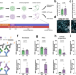 An alternative splicing modulator decreases mutant HTT and improves the molecular fingerprint in Huntington’s disease patient neurons
An alternative splicing modulator decreases mutant HTT and improves the molecular fingerprint in Huntington’s disease patient neurons
Krach et al. dissect the molecular mechanism of the alternative splicing modulator Branaplam in Huntington’s disease. They show that the drug lowers mutant HTT protein levels and ameliorates alternative splicing pathology in an iPSC disease model.
- Florian Krach
- , Judith Stemick
- & Juergen Winkler
-
Article
| Open Access Chemical engineering of therapeutic siRNAs for allele-specific gene silencing in Huntington’s disease models
Chemical engineering of therapeutic siRNAs for allele-specific gene silencing in Huntington’s disease models
Chemically modified siRNAs distinguish between mutant and normal huntingtin based on a single nucleotide difference and lower mutant huntingtin specifically in patient derived cells and in a mouse model of Huntington’s disease.
- Faith Conroy
- , Rachael Miller
- & Edith L. Pfister
-
Article
| Open Access Identification of a HTT-specific binding motif in DNAJB1 essential for suppression and disaggregation of HTT
Identification of a HTT-specific binding motif in DNAJB1 essential for suppression and disaggregation of HTT
Ayala Mariscal et al have identified and characterized the interface of pathogenic Huntingtin and the molecular chaperone DNAJB1. Histidine-244 of the C-terminal domain of DNAJB1 is a key residues for binding to the poly-proline region of HTT. This binding site is specific for the interaction with Huntingtin.
- S. M. Ayala Mariscal
- , M. L. Pigazzini
- & J. Kirstein
-
Article
| Open Access An orally available, brain penetrant, small molecule lowers huntingtin levels by enhancing pseudoexon inclusion
An orally available, brain penetrant, small molecule lowers huntingtin levels by enhancing pseudoexon inclusion
Huntington’s disease (HD) results from the abnormal expansion of CAG repeats in exon 1 of the HTT gene. Here, the authors show that orally available, brain penetrant molecule branaplam lowers HTT transcript by promoting inclusion of a poison exon or pseudoexon.
- Caroline Gubser Keller
- , Youngah Shin
- & Rajeev Sivasankaran
-
Article
| Open Access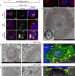 Nuclear and cytoplasmic huntingtin inclusions exhibit distinct biochemical composition, interactome and ultrastructural properties
Nuclear and cytoplasmic huntingtin inclusions exhibit distinct biochemical composition, interactome and ultrastructural properties
The mechanisms underlying Huntingtin protein (Htt) aggregation are not fully understood. Here the authors perform a detailed investigation of the ultrastructural and biochemical properties of huntingtin cytoplasmic and nuclear inclusions, and reveal that they form via distinct mechanisms and exert their toxicity via different pathways.
- Nathan Riguet
- , Anne-Laure Mahul-Mellier
- & Hilal A. Lashuel
-
Article
| Open Access Small-molecule suppression of calpastatin degradation reduces neuropathology in models of Huntington’s disease
Small-molecule suppression of calpastatin degradation reduces neuropathology in models of Huntington’s disease
Mitochondrial dysfunction is a common hallmark of neurological disorders. Here, the authors identify CHIR99021 as a potent enhancer of mitochondrial function, which improved mitochondrial phenotypes in Huntington’s disease models. CHIR99021 was shown to stabilize calpastatin, which suppressed calpain activation and Drp1-induced mitochondrial fragmentation.
- Di Hu
- , Xiaoyan Sun
- & Xin Qi
-
Article
| Open Access Mutant Huntingtin stalls ribosomes and represses protein synthesis in a cellular model of Huntington disease
Mutant Huntingtin stalls ribosomes and represses protein synthesis in a cellular model of Huntington disease
Huntington disease (HD) is a neurodegenerative disorder caused by the expansion of a polyglutamine tract in the huntingtin (mHtt) protein. Here the authors suggest that mHtt promotes ribosome stalling and inhibits protein synthesis.
- Mehdi Eshraghi
- , Pabalu P. Karunadharma
- & Srinivasa Subramaniam
-
Article
| Open Access Age-related and disease locus-specific mechanisms contribute to early remodelling of chromatin structure in Huntington’s disease mice
Age-related and disease locus-specific mechanisms contribute to early remodelling of chromatin structure in Huntington’s disease mice
The dynamics of chromatin and transcriptional changes underlying Huntington’s disease remain poorly understood. Here the authors use a Huntington’s mouse model to profile the striatal chromatin landscape, finding that the Huntington’s mutation accelerates age-dependent epigenetic and transcriptional changes, and locally affects 3D chromatin organization.
- Rafael Alcalá-Vida
- , Jonathan Seguin
- & Karine Merienne
-
Article
| Open Access DNA methylation study of Huntington’s disease and motor progression in patients and in animal models
DNA methylation study of Huntington’s disease and motor progression in patients and in animal models
Although Huntington’s disease (HD) is a well-studied genetic disorder, less is known about the epigenetic changes underlying it. Here, the authors characterize DNA methylation levels in tissues from patients, a mouse huntingtin (Htt) gene knock-in model, and a transgenic HTT sheep model, and provide evidence that HD is accompanied by DNA methylation changes in these three species.
- Ake T. Lu
- , Pritika Narayan
- & Steve Horvath
-
Article
| Open Access Truncation of mutant huntingtin in knock-in mice demonstrates exon1 huntingtin is a key pathogenic form
Truncation of mutant huntingtin in knock-in mice demonstrates exon1 huntingtin is a key pathogenic form
The mechanisms by which mutant Huntington protein Htt leads to selective neurodegeneration are not fully understood. Here, using gene editing in HD140Q knock-in mice, the authors show that exon1 Htt is a critical pathological form of the protein.
- Huiming Yang
- , Su Yang
- & Xiao-Jiang Li
-
Article
| Open Access Gene therapy conversion of striatal astrocytes into GABAergic neurons in mouse models of Huntington’s disease
Gene therapy conversion of striatal astrocytes into GABAergic neurons in mouse models of Huntington’s disease
In vivo reprogramming of reactive glia using transfection of a single transcription factor has been described before by these authors and applied to models of neurodegeneration. Here the authors use this procedure in the R6/2 mouse model of Huntington’s disease, targeting astrocytes in the striatum, converting them to GABAergic neurons.
- Zheng Wu
- , Matthew Parry
- & Gong Chen
-
Article
| Open Access MHC matching fails to prevent long-term rejection of iPSC-derived neurons in non-human primates
MHC matching fails to prevent long-term rejection of iPSC-derived neurons in non-human primates
Matching iPSC donors’ and patients’ HLA haplotypes has been proposed as a way to generate cell therapy products with enhanced immunological compatibility. Here the authors show that MHC matching alone is insufficient to grant long-term survival of neuronal grafts in the lesioned brain of non-human primates.
- Romina Aron Badin
- , Aurore Bugi
- & Anselme L. Perrier
-
Article
| Open Access Galectin-3 is required for the microglia-mediated brain inflammation in a model of Huntington’s disease
Galectin-3 is required for the microglia-mediated brain inflammation in a model of Huntington’s disease
The authors show that Galectin-3 is up–regulated in brain tissues from patients and a mouse model of Huntington’s disease (HD) and correlates with disease severity. Galectin-3 accumulates at damaged lysosomes in HD microglia, prevents the clearance of damaged lysosomes, and promotes inflammation.
- Jian Jing Siew
- , Hui-Mei Chen
- & Yijuang Chern
-
Article
| Open Access ATAD3A oligomerization causes neurodegeneration by coupling mitochondrial fragmentation and bioenergetics defects
ATAD3A oligomerization causes neurodegeneration by coupling mitochondrial fragmentation and bioenergetics defects
Huntington’s disease leads to mitochondrial fragmentation and bioenergetic failure, although how the two events are connected is poorly understood. Here, Zhao et al. identify ATAD3A as a molecular linker and show that a peptide inhibitor of ATAD3A oligomerization suppresses HD phenotypes.
- Yuanyuan Zhao
- , Xiaoyan Sun
- & Xin Qi
-
Article
| Open Access Modulation of AMPA receptor surface diffusion restores hippocampal plasticity and memory in Huntington’s disease models
Modulation of AMPA receptor surface diffusion restores hippocampal plasticity and memory in Huntington’s disease models
Cognitive decline in Huntington’s disease (HD) may be due to impaired hippocampal synaptic plasticity. In this study the authors show that AMPA receptor surface diffusion, a key player in synaptic plasticity, is deregulated in multiple HD mouse models as a result of impaired BDNF signalling that underlies the memory deficits, and can be pharmacologically rescued.
- Hongyu Zhang
- , Chunlei Zhang
- & Daniel Choquet
-
Article
| Open Access Deubiquitinase Usp12 functions noncatalytically to induce autophagy and confer neuroprotection in models of Huntington’s disease
Deubiquitinase Usp12 functions noncatalytically to induce autophagy and confer neuroprotection in models of Huntington’s disease
Abnormal accumulations of toxic proteins are often found in degenerating neurons. Here, Aron and colleagues show that non-enzymatic function of deubiquitinase Usp12 can mitigate neuronal cell death caused by mutant Huntingtin by inducing neuronal autophagic function.
- Rebecca Aron
- , Pasquale Pellegrini
- & Steven Finkbeiner
-
Article
| Open Access Regulatory mechanisms of incomplete huntingtin mRNA splicing
Regulatory mechanisms of incomplete huntingtin mRNA splicing
Incomplete splicing of HTT results in the production of the highly pathogenic exon 1 HTT protein. Here the authors identify the necessary intronic regions and the underlying mechanisms that contribute to this process.
- Andreas Neueder
- , Anaelle A. Dumas
- & Gillian P. Bates
-
Article
| Open Access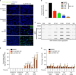 The ubiquitin ligase UBR5 suppresses proteostasis collapse in pluripotent stem cells from Huntington’s disease patients
The ubiquitin ligase UBR5 suppresses proteostasis collapse in pluripotent stem cells from Huntington’s disease patients
Induced pluripotent stem cells (iPSCs) suppress the aggregation of Huntington’s disease (HD) polyQ-expanded huntingtin (HTT). Here the authors show that proteasome activity determines the levels of mutant HTT in HD-iPSCs and find that UBR5 is a modulator of super-vigilant proteostasis of iPSCs.
- Seda Koyuncu
- , Isabel Saez
- & David Vilchez
-
Article
| Open Access Pyk2 modulates hippocampal excitatory synapses and contributes to cognitive deficits in a Huntington’s disease model
Pyk2 modulates hippocampal excitatory synapses and contributes to cognitive deficits in a Huntington’s disease model
Several kinases regulate spine morphology and plasticity. Here, the authors show that the tyrosine kinase Pyk2 regulates hippocampal plasticity and contributes to cognitive and hippocampal plasticity deficits in a mouse model of Huntington’s disease.
- Albert Giralt
- , Veronica Brito
- & Jean-Antoine Girault
-
Article
| Open Access Fibril polymorphism affects immobilized non-amyloid flanking domains of huntingtin exon1 rather than its polyglutamine core
Fibril polymorphism affects immobilized non-amyloid flanking domains of huntingtin exon1 rather than its polyglutamine core
Huntington's disease is caused by a polyglutamine stretch expansion in the first exon of huntingtin. Here, the authors use infrared spectroscopy and solid-state NMR and show that polymorphic huntingtin exon1 fibres differ in their flanking regions but not their core polyglutamine amyloid structures.
- Hsiang-Kai Lin
- , Jennifer C. Boatz
- & Patrick C. A. van der Wel
-
Article
| Open Access Abnormal degradation of the neuronal stress-protective transcription factor HSF1 in Huntington’s disease
Abnormal degradation of the neuronal stress-protective transcription factor HSF1 in Huntington’s disease
Huntington’s disease (HD) is caused by misfolding of mutant Htt protein. The authors find that in HD models, the decreased expression of heat shock transcription factor 1 that usually protects against protein misfolding, is in part caused by elevated CK2α’ kinase and Fbxw7 E3 ligase expression.
- Rocio Gomez-Pastor
- , Eileen T. Burchfiel
- & Dennis J. Thiele
-
Article
| Open Access Human glia can both induce and rescue aspects of disease phenotype in Huntington disease
Human glia can both induce and rescue aspects of disease phenotype in Huntington disease
The contribution of glia to Huntington's disease is unclear. The authors show that human glial progenitor cells (GPCs) expressing mutant huntingtin impair motor performance when engrafted into wild type mice, and wild type human GPCs ameliorate disease phenotypes when engrafted into an HD mouse model.
- Abdellatif Benraiss
- , Su Wang
- & Steven A. Goldman
-
Article
| Open Access Real-time imaging of glutamate clearance reveals normal striatal uptake in Huntington disease mouse models
Real-time imaging of glutamate clearance reveals normal striatal uptake in Huntington disease mouse models
Huntington disease (HD) has been linked via biochemical uptake assays to impaired glutamate clearance and resultant excitotoxicity. Here, utilizing a fluorescent reporter, the authors measure real-time glutamate dynamics in mouse model HD brain slices and find normal or even accelerated glutamate clearance.
- Matthew P. Parsons
- , Matthieu P. Vanni
- & Lynn A. Raymond
-
Article |
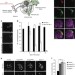 Prion-like transmission of neuronal huntingtin aggregates to phagocytic glia in the Drosophila brain
Prion-like transmission of neuronal huntingtin aggregates to phagocytic glia in the Drosophila brain
Phagocytic glia can internalize protein aggregates in vitro. Here Pearce et al. show in Drosophila that glia clear mutant huntingtin (Htt) aggregates in a scavenger receptor Draper-dependent manner in vivo, and that internalized Htt aggregates induce the prion-like conversion of wild-type Htt expressed in the glial cytoplasm.
- Margaret M. P. Pearce
- , Ellen J. Spartz
- & Ron R. Kopito
-
Article
| Open Access A failure in energy metabolism and antioxidant uptake precede symptoms of Huntington’s disease in mice
A failure in energy metabolism and antioxidant uptake precede symptoms of Huntington’s disease in mice
Defective ascorbic acid flux is a sign of metabolic failure associated with Huntington’s disease. Here, Acuña et al.show that reduction in ascorbic acid flux from astrocytes precedes the symptoms of Huntington’s disease in mice and impairs ascorbic acid uptake in neurons.
- Aníbal I. Acuña
- , Magdalena Esparza
- & Maite A. Castro
-
Article |
 Soluble forms of polyQ-expanded huntingtin rather than large aggregates cause endoplasmic reticulum stress
Soluble forms of polyQ-expanded huntingtin rather than large aggregates cause endoplasmic reticulum stress
Large insoluble aggregates of huntingtin are believed to contribute to the onset of Huntington’s disease, partly by interfering with endoplasmic reticulum-associated protein degradation. Leitman et al.show that huntingtin interferes with protein degradation before it forms insoluble aggregates.
- Julia Leitman
- , F. Ulrich Hartl
- & Gerardo Z. Lederkremer
-
Article |
 Translation of HTT mRNA with expanded CAG repeats is regulated by the MID1–PP2A protein complex
Translation of HTT mRNA with expanded CAG repeats is regulated by the MID1–PP2A protein complex
Expansion of CAG repeats in messenger RNAs is a common feature of various neurodegenerative disorders, including Huntington’s disease. Krauß et al.show that messenger RNAs with expanded CAG repeats bind to a protein complex that regulates translation and promotes overproduction of such aberrant proteins.
- Sybille Krauß
- , Nadine Griesche
- & Susann Schweiger

 Splice modulators target PMS1 to reduce somatic expansion of the Huntington’s disease-associated CAG repeat
Splice modulators target PMS1 to reduce somatic expansion of the Huntington’s disease-associated CAG repeat