Article
|
Open Access
Featured
-
-
Article
| Open Access High-throughput Pore-C reveals the single-allele topology and cell type-specificity of 3D genome folding
High-throughput Pore-C reveals the single-allele topology and cell type-specificity of 3D genome folding
Here the authors establish a high throughput Pore-C (HiPore-C) method increasing the sequencing output of multi-way chromatin contacts and reveal single-allele topology diversity and cell type-specificity of 3D genome folding.
- Jia-Yong Zhong
- , Longjian Niu
- & Chuan-Le Xiao
-
Article
| Open Access Systematic characterization of chromodomain proteins reveals an H3K9me1/2 reader regulating aging in C. elegans
Systematic characterization of chromodomain proteins reveals an H3K9me1/2 reader regulating aging in C. elegans
Chromodomains mainly function as histone methyl-lysine readers to regulate gene expression. Here the authors delineate a functional map of chromodomain proteins and identify an H3K9me1/2 writer, MET-2, and reader, CEC-5, that are required for the normal lifespan of C. elegans.
- Xinhao Hou
- , Mingjing Xu
- & Xuezhu Feng
-
Article
| Open Access Histone H2A monoubiquitination marks are targeted to specific sites by cohesin subunits in Arabidopsis
Histone H2A monoubiquitination marks are targeted to specific sites by cohesin subunits in Arabidopsis
How histone H2A monoubiquitination (H2Aub1) is established at specific genomic locations remains unclear. Here, the authors report that Arabidopsis cohesin subunits SCC3 and SYN4 are involved in H2Aub1 through their direct or indirect interaction with BMI1A/B/C subunits of PRC1, the E3 ligases in PRC1 for H2Aub1.
- Yu Zhang
- , Min Ma
- & Yuda Fang
-
Article
| Open Access Widespread contribution of transposable elements to the rewiring of mammalian 3D genomes
Widespread contribution of transposable elements to the rewiring of mammalian 3D genomes
Here the authors show transposable elements, formerly considered junk DNA, are a source of CTCF binding sites that contribute to species-specific 3D-genome structure, and may impact gene regulation during the course of mammalian evolution.
- Mayank N. K. Choudhary
- , Kara Quaid
- & Ting Wang
-
Article
| Open Access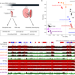 Mechanisms and function of de novo DNA methylation in placental development reveals an essential role for DNMT3B
Mechanisms and function of de novo DNA methylation in placental development reveals an essential role for DNMT3B
DNA methylation is a repressive modification that is essential for development. Here the authors reveal a critical role for DNA methylation in placental development during pregnancy. Failure to properly establish placental DNA methylation patterns compromises not only placental function, but embryo survival.
- Simon Andrews
- , Christel Krueger
- & Courtney W. Hanna
-
Article
| Open Access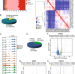 Epigenomic charting and functional annotation of risk loci in renal cell carcinoma
Epigenomic charting and functional annotation of risk loci in renal cell carcinoma
The epigenomic landscape of renal cell carcinoma (RCC) remains to be explored. Here, integrative epigenomic analysis of primary human RCC samples and RCC GWAS risk SNPs identifies transcription-factor specific subtypes and enrichment of risk variants in allelically-imbalanced peaks.
- Amin H. Nassar
- , Sarah Abou Alaiwi
- & Matthew L. Freedman
-
Article
| Open Access Mapping nucleolus-associated chromatin interactions using nucleolus Hi-C reveals pattern of heterochromatin interactions
Mapping nucleolus-associated chromatin interactions using nucleolus Hi-C reveals pattern of heterochromatin interactions
Here the authors developed a nucleolus Hi-C technique (nHi-C) for enriching nucleolus-associated interactions, and revealed specific heterochromatin interaction patterns within and around nucleoli in human cells at high resolution.
- Ting Peng
- , Yingping Hou
- & Cheng Li
-
Article
| Open Access Semi-quantitative detection of pseudouridine modifications and type I/II hypermodifications in human mRNAs using direct long-read sequencing
Semi-quantitative detection of pseudouridine modifications and type I/II hypermodifications in human mRNAs using direct long-read sequencing
Pseudouridine (psi) is an RNA modification that can affect its physiology, including increased half-life. Here the authors identify sites of psi modification in the human transcriptome using direct RNA sequencing and provide a “ground truth” list of psi sites, sites of high psi occupancy, and transcripts that may be modified at multiple sites.
- Sepideh Tavakoli
- , Mohammad Nabizadeh
- & Sara H. Rouhanifard
-
Article
| Open Access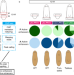 Wnt/β-catenin signalling is required for pole-specific chromatin remodeling during planarian regeneration
Wnt/β-catenin signalling is required for pole-specific chromatin remodeling during planarian regeneration
Any planarian fragment regenerates the missing head and tail in the proper end. Early activation of the Wnt/β-catenin signaling pathway changes the chromatin accessibility of the cells of the posterior-facing wound to regenerate a tail.
- Eudald Pascual-Carreras
- , Marta Marín-Barba
- & Teresa Adell
-
Article
| Open Access Comparative analysis of genome-scale, base-resolution DNA methylation profiles across 580 animal species
Comparative analysis of genome-scale, base-resolution DNA methylation profiles across 580 animal species
DNA methylation is involved in regulatory processes throughout the animal kingdom. Here, the authors map DNA methylation in 535 vertebrates and 45 invertebrates, establishing a reference dataset for cross-species analysis and exploring epigenetic variation across vertebrate evolution.
- Johanna Klughammer
- , Daria Romanovskaia
- & Christoph Bock
-
Article
| Open Access A comparison of the genes and genesets identified by GWAS and EWAS of fifteen complex traits
A comparison of the genes and genesets identified by GWAS and EWAS of fifteen complex traits
Genome-wide and epigenome-wide association studies both link genomic regions to human traits, but here the authors demonstrate that these study types are capturing different genes and biological aspects of complex traits.
- Thomas Battram
- , Tom R. Gaunt
- & Gibran Hemani
-
Article
| Open Access High-throughput robust single-cell DNA methylation profiling with sciMETv2
High-throughput robust single-cell DNA methylation profiling with sciMETv2
Despite the importance of DNA methylation, accessible and high-throughput methods to profile methylation at the single-cell level are lacking. Here, the authors present sciMETv2, a high-throughput workflow that provides high-quality single-cell methylomes in a robust and simple workflow.
- Ruth V. Nichols
- , Brendan L. O’Connell
- & Andrew C. Adey
-
Article
| Open Access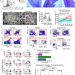 The 3D enhancer network of the developing T cell genome is shaped by SATB1
The 3D enhancer network of the developing T cell genome is shaped by SATB1
Here the authors analyze the 3D genome structure of murine thymocytes and show that SATB1, a predominantly T-cell specific protein, helps to establish a regulatory, finer-scale organizational layer built upon a pre-existing chromatin scaffold mediated by other architectural proteins, such as CTCF.
- Tomas Zelenka
- , Antonios Klonizakis
- & Charalampos Spilianakis
-
Article
| Open Access Transposable elements orchestrate subgenome-convergent and -divergent transcription in common wheat
Transposable elements orchestrate subgenome-convergent and -divergent transcription in common wheat
How subgenome-divergent and -convergent transcription is mediated and harmonized in hexaploid common wheat genome remains unclear. Here, via characterizing the cistrome maps, the authors reveal that transposon elements with transcription factor binding ability have the potential to make the contribution.
- Yuyun Zhang
- , Zijuan Li
- & Yijing Zhang
-
Article
| Open Access Epigenetic control of chromosome-associated lncRNA genes essential for replication and stability
Epigenetic control of chromosome-associated lncRNA genes essential for replication and stability
Heskett et al. describe several members of a class of long non-coding RNAs, known as ASARs, which show distinct epigenetic regulation between subclonal lineages and are essential for normal DNA replication timing and stability of human autosomes.
- Michael B. Heskett
- , Athanasios E. Vouzas
- & Mathew J. Thayer
-
Article
| Open Access The immune factors driving DNA methylation variation in human blood
The immune factors driving DNA methylation variation in human blood
Many studies assess epigenetic marks in white blood cells, but it is unclear how much immune factors affect the epigenome. Here, the authors show that fine-scale blood cell composition and cytomegalovirus infection affect the DNA methylome of adults.
- Jacob Bergstedt
- , Sadoune Ait Kaci Azzou
- & Lluis Quintana-Murci
-
Article
| Open Access Autophagy induction promoted by m6A reader YTHDF3 through translation upregulation of FOXO3 mRNA
Autophagy induction promoted by m6A reader YTHDF3 through translation upregulation of FOXO3 mRNA
The role of eiptranscriptomic modifications in autophagy is unclear. Here, the authors show that the m6A reader YTHDF3 functions as a nutrient responder to recognize upregulated m6A modification, promoting FOXO3 translation to subsequently initiate autophagy.
- WeiChao Hao
- , MeiJuan Dian
- & Dong Xiao
-
Article
| Open Access Modeling tissue-specific breakpoint proximity of structural variations from whole-genomes to identify cancer drivers
Modeling tissue-specific breakpoint proximity of structural variations from whole-genomes to identify cancer drivers
Identifying structural variants (SVs) under positive selection in cancer is challenging. Here, the authors develop CSVDriver, a method that computes SV breakpoint proximity and the contribution of elements such as topologically associating domains, and identifies loci that show signs of positive selection and contain known and putative drivers.
- Alexander Martinez-Fundichely
- , Austin Dixon
- & Ekta Khurana
-
Article
| Open Access Multi-omic brain and behavioral correlates of cell-free fetal DNA methylation in macaque maternal obesity models
Multi-omic brain and behavioral correlates of cell-free fetal DNA methylation in macaque maternal obesity models
In animal models, maternal obesity is associated with development of neurodevelopmental disorder like phenotypes. Here the authors show in a macaque model that in obese dams, cell-free fetal DNA methylation, inflammatory cytokines, and metabolites correlated with infant brain DNA methylation, lipids, and metabolites.
- Benjamin I. Laufer
- , Yu Hasegawa
- & Janine M. LaSalle
-
Article
| Open Access Intrinsic bias estimation for improved analysis of bulk and single-cell chromatin accessibility profiles using SELMA
Intrinsic bias estimation for improved analysis of bulk and single-cell chromatin accessibility profiles using SELMA
Genome-wide profiling of chromatin accessibility by DNase-seq or ATAC-seq has been widely used to identify regulatory DNA elements and transcription factor binding sites. Here the authors develop a computational model, SELMA, to estimate and correct enzymatic cleavage biases in chromatin accessibility profiling data.
- Shengen Shawn Hu
- , Lin Liu
- & Chongzhi Zang
-
Article
| Open Access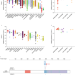 Epigenetic activation of the FLT3 gene by ZNF384 fusion confers a therapeutic susceptibility in acute lymphoblastic leukemia
Epigenetic activation of the FLT3 gene by ZNF384 fusion confers a therapeutic susceptibility in acute lymphoblastic leukemia
Different molecular subtypes defined by specific gene rearrangements have been described for acute lymphoblastic leukaemia (ALL). Here, the authors show that ZNF384 fusion activates FLT3 expression conferring a therapeutic vulnerability for ZNF384- rearranged ALL subtype.
- Xujie Zhao
- , Ping Wang
- & Jun J. Yang
-
Article
| Open Access Multiple parameters shape the 3D chromatin structure of single nuclei at the doc locus in Drosophila
Multiple parameters shape the 3D chromatin structure of single nuclei at the doc locus in Drosophila
Here the authors applied their recently developed multiplexed DNA-FISH Hi-M method to dissect the sources of heterogeneity in topologically associating domain (TAD)-like organization during Drosophila embryogenesis. This single-nucleus analysis allows them to reveal that multiple parameters contribute to shaping the trace of the chromatin path from a single nucleus.
- Markus Götz
- , Olivier Messina
- & Marcelo Nollmann
-
Article
| Open Access Clonal hematopoiesis of indeterminate potential, DNA methylation, and risk for coronary artery disease
Clonal hematopoiesis of indeterminate potential, DNA methylation, and risk for coronary artery disease
Clonal hematopoiesis, often caused by mutations in DNMT3A and TET2, is associated with blood cancer and coronary artery disease. Here, the authors conduct an epigenome-wide association study, finding that clonal hematopoiesis caused by DNMT3A vs. TET2 mutations has directionally opposing changes in DNA methylation profiles, with both promoting stem cell self-renewal.
- M d Mesbah Uddin
- , Ngoc Quynh H. Nguyen
- & Karen N. Conneely
-
Article
| Open Access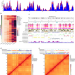 Regulation associated modules reflect 3D genome modularity associated with chromatin activity
Regulation associated modules reflect 3D genome modularity associated with chromatin activity
Here the authors report histone modifications show a modular pattern referred to as ‘regulation associated modules’ (RAMs) that reflect the spatial modularity of chromatin. They find enhancer-promoter interactions and extrachromosomal DNAs (ecDNAs) occur more often within the same RAMs than within the same TADs, indicating stronger insulation of the RAM boundaries.
- Lina Zheng
- & Wei Wang
-
Article
| Open Access KLF4 recruits SWI/SNF to increase chromatin accessibility and reprogram the endothelial enhancer landscape under laminar shear stress
KLF4 recruits SWI/SNF to increase chromatin accessibility and reprogram the endothelial enhancer landscape under laminar shear stress
Here the authors studied pulmonary arterial endothelial cells (PAEC) under laminar shear stress and show that this physiologic condition markedly changes chromatin accessibility at regulatory regions, when compared to cells grown in a static state. They find that KLF4 organizes chromatin by interacting with the SWI/SNF nucleosome remodeling complex to regulate vasculo-protective gene expression.
- Jan-Renier Moonen
- , James Chappell
- & Marlene Rabinovitch
-
Article
| Open Access Transcription-coupled and epigenome-encoded mechanisms direct H3K4 methylation
Transcription-coupled and epigenome-encoded mechanisms direct H3K4 methylation
Methylation of histone H3 lysine 4 is associated with transcription. Here the authors describe three Arabidopsis methyltransferases that direct H3K4me1 and propose that one methyltransferase works co-transcriptionally while the others are recruited to genomic and epigenomic features.
- Satoyo Oya
- , Mayumi Takahashi
- & Soichi Inagaki
-
Article
| Open Access Integrated methylome and phenome study of the circulating proteome reveals markers pertinent to brain health
Integrated methylome and phenome study of the circulating proteome reveals markers pertinent to brain health
Characterising associations between the methylome, proteome and phenome may provide insight into biological pathways governing brain health. Here, blood protein markers of brain health are integrated with omics data to reveal DNA methylation differences that associate with these protein markers.
- Danni A. Gadd
- , Robert F. Hillary
- & Riccardo E. Marioni
-
Article
| Open Access Whole blood DNA methylation analysis reveals respiratory environmental traits involved in COVID-19 severity following SARS-CoV-2 infection
Whole blood DNA methylation analysis reveals respiratory environmental traits involved in COVID-19 severity following SARS-CoV-2 infection
Genetic associations with severe COVID-19 have been discovered, but epigenetic associations are not as well described. Here, the authors perform a genome-wide epigenetic analysis of COVID-19 patients, discovering an interaction between environmental exposure, genetics, and epigenetics which might play a role in severe disease.
- Guillermo Barturen
- , Elena Carnero-Montoro
- & Marta E. Alarcón-Riquelme
-
Article
| Open Access TP53-dependent toxicity of CRISPR/Cas9 cuts is differential across genomic loci and can confound genetic screening
TP53-dependent toxicity of CRISPR/Cas9 cuts is differential across genomic loci and can confound genetic screening
Toxicity of CRISPR/Cas9 induced DNA breaks depends on their repair mechanism, and on the chromatin environment at the cut site. Here the authors show that edits in active genes or regulatory elements can incur a higher toxicity via a TP53-dependent mechanism.
- Miguel M. Álvarez
- , Josep Biayna
- & Fran Supek
-
Article
| Open Access Maternal SMCHD1 regulates Hox gene expression and patterning in the mouse embryo
Maternal SMCHD1 regulates Hox gene expression and patterning in the mouse embryo
Parents transmit both genetic and epigenetic information to their offspring, with maternal effect genes being critical regulators of the offspring epigenome. Here they show that maternally deposited SMCHD1 has long-lasting effects on Hox gene expression and vertebral patterning during post-implantation development.
- Natalia Benetti
- , Quentin Gouil
- & Marnie E. Blewitt
-
Article
| Open Access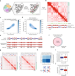 Polycomb-lamina antagonism partitions heterochromatin at the nuclear periphery
Polycomb-lamina antagonism partitions heterochromatin at the nuclear periphery
Here the authors developed ‘Lamina-Inducible Methylation and Hi-C’ (LIMe-Hi-C) to simultaneously measure chromosome conformation, DNA methylation, and nuclear lamina positioning. Application of the method revealed dynamic changes upon PRC2 inhibition and an essential function of H3K27me3 in regulating sub-compartments and lamina association.
- Allison P. Siegenfeld
- , Shelby A. Roseman
- & Brian B. Liau
-
Article
| Open Access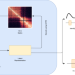 Learning representations of chromatin contacts using a recurrent neural network identifies genomic drivers of conformation
Learning representations of chromatin contacts using a recurrent neural network identifies genomic drivers of conformation
Despite the availability of chromatin conformation capture experiments, discerning the relationship between the 1D genome and 3D conformation remains a challenge. Here, the authors propose a method that produces low-dimensional latent representations that summarize intra-chromosomal Hi-C contacts.
- Kevin B. Dsouza
- , Alexandra Maslova
- & Maxwell W. Libbrecht
-
Article
| Open Access Large-scale multi-omics analysis suggests specific roles for intragenic cohesin in transcriptional regulation
Large-scale multi-omics analysis suggests specific roles for intragenic cohesin in transcriptional regulation
Cohesin complex is essential for transcriptional regulation and chromosome folding. Here the authors provide a large-scale multi-omics analysis suggesting that cohesin bound on gene bodies has a specific role in transcriptional regulation.
- Jiankang Wang
- , Masashige Bando
- & Ryuichiro Nakato
-
Article
| Open Access Integrating 3D genomic and epigenomic data to enhance target gene discovery and drug repurposing in transcriptome-wide association studies
Integrating 3D genomic and epigenomic data to enhance target gene discovery and drug repurposing in transcriptome-wide association studies
Transcriptome-wide association studies can be used to test the effects of predicted gene expression in a cohort of individuals based on genetic data. Here, the authors developed a transcriptome-wide association method that integrates 3D genomic and epigenomic data with expression quantitative trait loci to improve gene expression predictions.
- Chachrit Khunsriraksakul
- , Daniel McGuire
- & Dajiang J. Liu
-
Article
| Open Access Integrative epigenomic and transcriptomic analysis reveals the requirement of JUNB for hematopoietic fate induction
Integrative epigenomic and transcriptomic analysis reveals the requirement of JUNB for hematopoietic fate induction
Here they perform an integrative analysis of the epigenetic landscape of human pluripotent stem cell hematopoietic differentiation and show that JUNB is an indispensable transcription factor for hemogenic endothelium development.
- Xia Chen
- , Peiliang Wang
- & Jie Na
-
Article
| Open Access Single-cell chromatin profiling of the primitive gut tube reveals regulatory dynamics underlying lineage fate decisions
Single-cell chromatin profiling of the primitive gut tube reveals regulatory dynamics underlying lineage fate decisions
The primitive gut tube gives rise to all major internal organs, while underlying regulatory mechanisms are unclear. Here, the authors analyze its chromatin landscape at the single-cell level and define the epigenetic regulation of lineage fate decisions and plasticity in organ development and homeostasis.
- Ryan J. Smith
- , Hongpan Zhang
- & Tae-Hee Kim
-
Article
| Open Access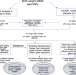 Placental multi-omics integration identifies candidate functional genes for birthweight
Placental multi-omics integration identifies candidate functional genes for birthweight
The placenta plays key roles in fetal development and subsequent health. Here, the authors integrate placental methylation and transcriptome data with genetic loci associated with birthweight to identify functional genes underpinning fetal growth regulation.
- Fasil Tekola-Ayele
- , Xuehuo Zeng
- & Ronald Wapner
-
Article
| Open Access Inducible and reversible RNA N6-methyladenosine editing
Inducible and reversible RNA N6-methyladenosine editing
RNA modifications, including N6-methyladenosine (m6A), have been reported to regulate fundamental RNA processes and properties, and directly linked to various human diseases. Here, the authors develop a chemically inducible and reversible RNA m6A modification editing platform integrating chemically induced proximity (CIP) and CRISPR methods.
- Huaxia Shi
- , Ying Xu
- & Fu-Sen Liang
-
Article
| Open Access Multidimensional chromatin profiling of zebrafish pancreas to uncover and investigate disease-relevant enhancers
Multidimensional chromatin profiling of zebrafish pancreas to uncover and investigate disease-relevant enhancers
Alterations in cis-regulatory elements (CREs) can contribute to pancreatic diseases. Here the authors combine chromatin profiling and interaction points with in vivo reporter assays in zebrafish to uncover functionally equivalent human CREs, helping to predict disease-relevant enhancers.
- Renata Bordeira-Carriço
- , Joana Teixeira
- & José Bessa
-
Article
| Open Access Lupus enhancer risk variant causes dysregulation of IRF8 through cooperative lncRNA and DNA methylation machinery
Lupus enhancer risk variant causes dysregulation of IRF8 through cooperative lncRNA and DNA methylation machinery
The functional effects of genetic loci associated with autoimmune disease are not well understood. By dissecting an autoimmune disease genetic locus, the authors define an immune cell-type-specific enhancer and the molecular mechanisms underlying the dysregulation of IRF8 expression by lupus risk variants.
- Tian Zhou
- , Xinyi Zhu
- & Nan Shen
-
Article
| Open Access DNA sequence-dependent formation of heterochromatin nanodomains
DNA sequence-dependent formation of heterochromatin nanodomains
The ability to predict epigenetic regulation is an important challenge in biology. Here the authors describe heterochromatin nanodomains (HNDs) and compare four different types of H3K9me2/3-marked HNDs in mouse embryonic stem cells. They further develop a computational framework to predict genome-wide HND maps from DNA sequence and protein concentrations, at single-nucleotide resolution.
- Graeme J. Thorn
- , Christopher T. Clarkson
- & Vladimir B. Teif
-
Article
| Open Access Single-cell Atlas of common variable immunodeficiency shows germinal center-associated epigenetic dysregulation in B-cell responses
Single-cell Atlas of common variable immunodeficiency shows germinal center-associated epigenetic dysregulation in B-cell responses
Common variable immunodeficiency (CVID) is the most prevalent primary immunodeficiency. Here the authors perform single-cell omics analyses in CVID-discordant monozygotic twins and show epigenetic and transcriptional alterations associated with activation in memory B cells.
- Javier Rodríguez-Ubreva
- , Anna Arutyunyan
- & Esteban Ballestar
-
Article
| Open Access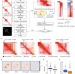 Stripenn detects architectural stripes from chromatin conformation data using computer vision
Stripenn detects architectural stripes from chromatin conformation data using computer vision
Chromosome conformation capture techniques have recently revealed features beyond chromatin loops such as architectural stripes. Here the authors present their stripe detection tool ‘Stripenn’ to detect and quantitate stripes from any type of chromatin conformation capture data. They show that architectural stripes are enriched at transcriptionally active and accessible genomic regions.
- Sora Yoon
- , Aditi Chandra
- & Golnaz Vahedi
-
Article
| Open Access Oncogenic gene expression and epigenetic remodeling of cis-regulatory elements in ASXL1-mutant chronic myelomonocytic leukemia
Oncogenic gene expression and epigenetic remodeling of cis-regulatory elements in ASXL1-mutant chronic myelomonocytic leukemia
‘Mutations in the chromatin remodeler ASXL1 (ASXL1MT) are associated with poor clinical outcome, however, their impact on chromatin dynamics remains unexplored. Here the authors use a multi-omics approach for chronic myelomonocytic leukemia (CMML) and investigate the transcriptome and chromatin landscape of ASXL1MT CMML.
- Moritz Binder
- , Ryan M. Carr
- & Mrinal M. Patnaik
-
Article
| Open Access Bacterial N4-methylcytosine as an epigenetic mark in eukaryotic DNA
Bacterial N4-methylcytosine as an epigenetic mark in eukaryotic DNA
Eukaryotic DNA can be methylated as 5-methylcytosine and N6-methyladenine, but whether other forms of DNA methylation occur has been controversial. Here the authors show that a bacterial DNA methyltransferase was acquired >60 Mya in bdelloid rotifers that catalyzes N4-methylcytosine addition and is involved in suppression of transposon proliferation.
- Fernando Rodriguez
- , Irina A. Yushenova
- & Irina R. Arkhipova
-
Article
| Open Access GLI transcriptional repression is inert prior to Hedgehog pathway activation
GLI transcriptional repression is inert prior to Hedgehog pathway activation
GLI repression has been presumed to be the default transcriptional state and important for pre-patterning tissues. Challenging current models, the authors show that GLI3 repression is inert in the limb bud before the onset of Hedgehog signaling.
- Rachel K. Lex
- , Weiqiang Zhou
- & Steven A. Vokes
-
Article
| Open Access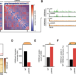 A Mediator-cohesin axis controls heterochromatin domain formation
A Mediator-cohesin axis controls heterochromatin domain formation
The link between 3D genome architecture and gene expression is still far from resolved. Here the authors show that loss of the CDK catalytic subunit of the Mediator complex results in heterochromatic silencing, which can be rescued by stabilization of cohesin on chromatin.
- Judith H. I. Haarhuis
- , Robin H. van der Weide
- & Elzo de Wit
-
Article
| Open Access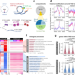 Pre-configuring chromatin architecture with histone modifications guides hematopoietic stem cell formation in mouse embryos
Pre-configuring chromatin architecture with histone modifications guides hematopoietic stem cell formation in mouse embryos
Here, the authors employed Hi-C and low-input itChIP-seq in four rare populations of the hematopoietic stem cell (HSC) ontogeny trajectory across early arterial endothelial cells (eAECs), hemogenic endothelial cells (HECs), pre-HSCs, and long-term HSCs (LT-HSCs) from mouse embryos to show that active histone modifications are largely set up in eAECs and 3D genome is then subsequently configured.
- Chen C. Li
- , Guangyu Zhang
- & Aibin He
-
Article
| Open Access Epigenetic loss of heterogeneity from low to high grade localized prostate tumours
Epigenetic loss of heterogeneity from low to high grade localized prostate tumours
High tumour heterogeneity hinders the identification of molecular subtypes in prostate cancer. Here, the authors integrate single-cell chromatin accessibility data with multiplex imaging and reveal distinct chromatin features and transcriptional factor binding signatures in high- and low-grade prostate tumours.
- Sebnem Ece Eksi
- , Alex Chitsazan
- & Andrew C. Adey

 MYC reshapes CTCF-mediated chromatin architecture in prostate cancer
MYC reshapes CTCF-mediated chromatin architecture in prostate cancer