Featured
-
-
Letter |
 Genome-wide detection of DNase I hypersensitive sites in single cells and FFPE tissue samples
Genome-wide detection of DNase I hypersensitive sites in single cells and FFPE tissue samples
A DNase sequencing method termed scDNase-seq detects DNase I hypersensitive sites genome-wide in single cells and pools of cells dissected from cancer biopsies.
- Wenfei Jin
- , Qingsong Tang
- & Keji Zhao
-
Letter |
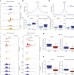 Integrator mediates the biogenesis of enhancer RNAs
Integrator mediates the biogenesis of enhancer RNAs
This study demonstrates a role for the Integrator complex in the stimulus-dependent induction of eRNAs and their 3′ processing; together with previously known roles of Integrator in transcription elongation and RNA processing, these results indicate that Integrator has broad functions in the regulation of eukaryotic gene expression.
- Fan Lai
- , Alessandro Gardini
- & Ramin Shiekhattar
-
Letter |
 Single-cell chromatin accessibility reveals principles of regulatory variation
Single-cell chromatin accessibility reveals principles of regulatory variation
A single-cell method for probing genome-wide chromatin accessibility has been developed; the results provide insight into the relationship between cell-to-cell variation associated with specific trans-factors and cis-elements, as well insights into the relationship between chromatin accessibility and three-dimensional genome organization.
- Jason D. Buenrostro
- , Beijing Wu
- & William J. Greenleaf
-
Letter
| Open Access Human body epigenome maps reveal noncanonical DNA methylation variation
Human body epigenome maps reveal noncanonical DNA methylation variation
As part of the Epigenome Roadmap Project, genome-wide maps of DNA methylation and transcriptomes together with genomic DNA sequencing of 18 different primary human tissue types from 4 individuals are presented; analysis reveals widespread differential methylation of CG sites between tissues, and the presence of non-CG methylation in adult tissues.
- Matthew D. Schultz
- , Yupeng He
- & Joseph R. Ecker
-
Article
| Open Access Integrative analysis of 111 reference human epigenomes
Integrative analysis of 111 reference human epigenomes
This study describes the integrative analysis of 111 reference human epigenomes, profiled for histone modification patterns, DNA accessibility, DNA methylation and RNA expression; the results annotate candidate regulatory elements in diverse tissues and cell types, their candidate regulators, and the set of human traits for which they show genetic variant enrichment, providing a resource for interpreting the molecular basis of human disease.
- Anshul Kundaje
- , Wouter Meuleman
- & Manolis Kellis
-
Letter
| Open Access Integrative analysis of haplotype-resolved epigenomes across human tissues
Integrative analysis of haplotype-resolved epigenomes across human tissues
As part of the Epigenome Roadmap project, this study uses a chromosome-spanning haplotype reconstruction strategy to construct haplotype-resolved epigenomic maps for a diverse set of human tissues; the maps reveal extensive allelic biases in chromatin state and transcription, which vary across individuals due to genetic backgrounds.
- Danny Leung
- , Inkyung Jung
- & Bing Ren
-
Article
| Open Access Transcription factor binding dynamics during human ES cell differentiation
Transcription factor binding dynamics during human ES cell differentiation
Lineage-specific transcription factors and signalling pathways cooperate with pluripotency regulators to control the transcriptional networks that drive cell specification and exit from an embryonic stem cell state; here, we report genome-wide binding data for 38 transcription factors combined with analysis of epigenomic and gene expression data during the differentiation of human embryonic stem cells into the three germ layers.
- Alexander M. Tsankov
- , Hongcang Gu
- & Alexander Meissner
-
Article
| Open Access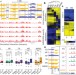 Chromatin architecture reorganization during stem cell differentiation
Chromatin architecture reorganization during stem cell differentiation
An analysis of genome-wide chromatin interactions during human embryonic stem cell differentiation reveals changes in chromatic organization and simultaneously identifies allele-resolved chromatin structure and differences in gene expression during differentiation.
- Jesse R. Dixon
- , Inkyung Jung
- & Bing Ren
-
Letter |
 Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation
Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation
Genome-wide localization and activity analysis of the de novo DNA methyltransferases DNMT3A and DNMT3B in mouse embryonic stem cells identifies overlapping and individual targeting preferences to the genome, including a role for DNMT3B in gene body methylation.
- Tuncay Baubec
- , Daniele F. Colombo
- & Dirk Schübeler
-
Letter |
 Dissecting neural differentiation regulatory networks through epigenetic footprinting
Dissecting neural differentiation regulatory networks through epigenetic footprinting
The integrative analysis of epigenetic footprints along consecutive stages of neural progenitors derived from human ES cells reveals regulatory mechanisms that orchestrate stage-specific differentiation.
- Michael J. Ziller
- , Reuven Edri
- & Alexander Meissner
-
Article
| Open Access A comparative encyclopedia of DNA elements in the mouse genome
A comparative encyclopedia of DNA elements in the mouse genome
The Mouse ENCODE Consortium has mapped transcription, DNase I hypersensitivity, transcription factor binding, chromatin modifications and replication domains throughout the mouse genome in diverse cell and tissue types; these data were compared with those from human to confirm substantial conservation in the newly annotated potential functional sequences and to reveal pronounced divergence of other sequences involved in transcriptional regulation, chromatin state and higher order chromatin organization.
- Feng Yue
- , Yong Cheng
- & Bing Ren
-
Article |
 Genetic and epigenetic fine mapping of causal autoimmune disease variants
Genetic and epigenetic fine mapping of causal autoimmune disease variants
Genome-wide association studies combined with data from epigenomic maps for immune cells have been used to fine-map causal variants for 21 autoimmune diseases; disease risk tends to be linked to single nucleotide polymorphisms in cell-type-specific enhancers, often in regions adjacent to transcription factor binding motifs.
- Kyle Kai-How Farh
- , Alexander Marson
- & Bradley E. Bernstein
-
Letter
| Open Access Comparative analysis of metazoan chromatin organization
Comparative analysis of metazoan chromatin organization
A large collection of new modENCODE and ENCODE genome-wide chromatin data sets from cell lines and developmental stages in worm, fly and human are analysed; this reveals many conserved features of chromatin organization among the three organisms, as well as notable differences in the composition and locations of repressive chromatin.
- Joshua W. K. Ho
- , Youngsook L. Jung
- & Peter J. Park
-
Letter |
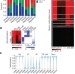 DNA methylation dynamics of the human preimplantation embryo
DNA methylation dynamics of the human preimplantation embryo
Genome-scale DNA methylation maps over early human embryogenesis and embryonic stem cell derivation provide insights into shared and unique modes of regulation when compared to the mouse model, including relationships to gene expression, transposable element activity, and maternal-specific methylation.
- Zachary D. Smith
- , Michelle M. Chan
- & Alexander Meissner
-
Outlook |
 Food science: Fat chance
Food science: Fat chance
For children with epilepsy whose condition is resistant to medication, a high-fat, low-carbohydrate diet may help bring their seizures under control.
- Rachel Brazil
-
Letter |
 Chromatin connectivity maps reveal dynamic promoter–enhancer long-range associations
Chromatin connectivity maps reveal dynamic promoter–enhancer long-range associations
A chromatin interaction analysis with paired-end tagging (ChIA-PET) approach is used to delineate chromatin interactions mediated by RNA polymerase II in several different stem-cell populations; putative long-range promoter–enhancer interactions are inferred, indicating that linear juxtaposition does not necessarily guide enhancer target selection and prevalent cell-specific enhancer usage.
- Yubo Zhang
- , Chee-Hong Wong
- & Chia-Lin Wei
-
Letter |
 Histone deacetylase 3 coordinates commensal-bacteria-dependent intestinal homeostasis
Histone deacetylase 3 coordinates commensal-bacteria-dependent intestinal homeostasis
This work identifies a role for intestinal epithelial cell (IEC)-intrinsic expression of histone deacetylase 3 in regulating commensal-bacteria-dependent gene expression and intestinal homeostasis; IEC-specific HDAC3 deficiency gives rise to Paneth cell abnormalities, impaired intestinal barrier function, and increased DSS-induced intestinal inflammation in commensal-bacteria-containing, but not germ-free, mice.
- Theresa Alenghat
- , Lisa C. Osborne
- & David Artis
-
Letter |
 A high-resolution map of the three-dimensional chromatin interactome in human cells
A high-resolution map of the three-dimensional chromatin interactome in human cells
A novel approach to analyse high-depth Hi-C data provides a comprehensive chromatin interaction map at approximately 5–10 kb resolution in human fibroblasts; this reveals that TNF-α-responsive enhancers are already in contact with target promoters before signalling and that this chromatin looping is a strong predictor of gene induction.
- Fulai Jin
- , Yan Li
- & Bing Ren
-
Article |
 Single-cell Hi-C reveals cell-to-cell variability in chromosome structure
Single-cell Hi-C reveals cell-to-cell variability in chromosome structure
A novel genomic technique, single-cell Hi-C, detects thousands of simultaneous chromatin contacts in a single cell; this is used to show that individual chromosomes maintain domain organization at the megabase scale, but that chromosome structures vary from cell to cell at larger scales.
- Takashi Nagano
- , Yaniv Lubling
- & Peter Fraser
-
Letter |
 Charting a dynamic DNA methylation landscape of the human genome
Charting a dynamic DNA methylation landscape of the human genome
Whole-genome bisulphite sequencing data from diverse human cell and tissue types shows that only about 22% of CpGs change their methylation state across these cell types; most of these CpGs are located at gene regulatory elements, particularly enhancers and transcription-factor-binding sites, and these selected regions with dynamic DNA methylation patterns could help to define putative regulatory elements further.
- Michael J. Ziller
- , Hongcang Gu
- & Alexander Meissner
-
Letter |
 The haplotype-resolved genome and epigenome of the aneuploid HeLa cancer cell line
The haplotype-resolved genome and epigenome of the aneuploid HeLa cancer cell line
Haplotype-resolved whole-genome sequencing of the HeLa CCL-2 strain shows that HeLa is relatively stable in terms of point variation; integration of several data sets reveals strong, haplotype-specific activation of the proto-oncogene MYC by the human papilloma virus type 18 genome, and enables the relationship between gene dosage and expression to be examined.
- Andrew Adey
- , Joshua N. Burton
- & Jay Shendure
-
Article
| Open Access Patterns of population epigenomic diversity
Patterns of population epigenomic diversity
A population epigenomic analysis of wild Arabidopsis thaliana accessions is presented, obtained by sequencing their whole genomes, methylomes and transcriptomes; thousands of DNA methylation variants are identified, some of which are associated with methylation quantitative trait loci.
- Robert J. Schmitz
- , Matthew D. Schultz
- & Joseph R. Ecker
-
News |
 Archived blood spots could be epigenetic jackpot
Archived blood spots could be epigenetic jackpot
But privacy concerns could complicate their use for research.
- Cassandra Willyard
-
Letter |
 RNF12 initiates X-chromosome inactivation by targeting REX1 for degradation
RNF12 initiates X-chromosome inactivation by targeting REX1 for degradation
The pluripotency factor REX1 is a key target of RNF12 during X-chromosome inactivation; degradation of REX1 by RNF12 leads to relief of its inhibitory action on X-chromosome inactivation.
- Cristina Gontan
- , Eskeatnaf Mulugeta Achame
- & Joost Gribnau
-
Outlook |
 Genetics: Seeking a gene genie
Genetics: Seeking a gene genie
Rare gene variants could be key to unlocking the underlying genetics of allergy, now that whole genome sequencing and other technologies have sharpened the focus of epidemiology.
- Erica Westly
-
News |
 Europe to map the human epigenome
Europe to map the human epigenome
DNA-modification studies get a multi-million euro boost.
- Alison Abbott
-
Article |
 Comprehensive analysis of the chromatin landscape in Drosophila melanogaster
Comprehensive analysis of the chromatin landscape in Drosophila melanogaster
As part of the modENCODE initiative, which aims to characterize functional DNA elements in D. melanogaster and C. elegans, this study presents a genome-wide chromatin landscape of the fruitfly, based on 18 histone modifications. Nine prevalent chromatin states are described. Integrating these analyses with other data types reveals individual characteristics of different genomic elements. The work provides a resource of unprecedented scale for future experimental investigations.
- Peter V. Kharchenko
- , Artyom A. Alekseyenko
- & Peter J. Park
-
News |
 Epigenome effort makes its mark
Epigenome effort makes its mark
Major release of maps charting non-genetic modifications goes beyond DNA in a bid to beat complex human disease.
- Alla Katsnelson
-
News |
 Genomics goes beyond DNA sequence
Genomics goes beyond DNA sequence
A technology that simultaneously reads a DNA sequence and its crucial modifications makes its debut.
- Alla Katsnelson
-
Letter
| Open Access Genome, epigenome and RNA sequences of monozygotic twins discordant for multiple sclerosis
Genome, epigenome and RNA sequences of monozygotic twins discordant for multiple sclerosis
Studies of identical twins are widely used to dissect the contributions of genes and the environment to human diseases. In multiple sclerosis, an autoimmune demyelinating disease, identical twins often show differences. This might suggest that environmental effects are most significant in this case, but genetic and epigenetic differences between identical twins have been described. Here, however, studies of identical twins show no evidence for genetic, epigenetic or transcriptome differences that could explain disease discordance.
- Sergio E. Baranzini
- , Joann Mudge
- & Stephen F. Kingsmore
-
Correspondence |
Questions over the scientific basis of epigenome project
- Mark Ptashne
- , Oliver Hobert
- & Eric Davidson
-
News |
 Project set to map marks on genome
Project set to map marks on genome
Consortium sets sights on the differences that make us different.
- Alison Abbott

 Genome-wide nucleosome specificity and function of chromatin remodellers in ES cells
Genome-wide nucleosome specificity and function of chromatin remodellers in ES cells