Article
|
Open Access
Featured
-
-
Article
| Open Access DeepDOF-SE: affordable deep-learning microscopy platform for slide-free histology
DeepDOF-SE: affordable deep-learning microscopy platform for slide-free histology
Histopathology can be limited by the time-consuming and labour-intensive preparation of slides from resected tissue. Here, the authors report DeepDOF-SE, a deep-learning-enabled microscope to rapidly scan intact tissue at cellular resolution without the need for physical sectioning.
- Lingbo Jin
- , Yubo Tang
- & Ashok Veeraraghavan
-
Article
| Open Access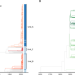 The recent rapid expansion of multidrug resistant Ural lineage Mycobacterium tuberculosis in Moldova
The recent rapid expansion of multidrug resistant Ural lineage Mycobacterium tuberculosis in Moldova
Chitwood et al. report on the rapid expansion of a Ural-lineage multidrug resistant strain of Mycobacterium tuberculosis in Moldova. This strain has an estimated reproduction number more than two times greater than otherwise similar drug susceptible strains.
- Melanie H. Chitwood
- , Caroline Colijn
- & Benjamin Sobkowiak
-
Article
| Open Access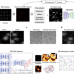 Context-aware deep learning enables high-efficacy localization of high concentration microbubbles for super-resolution ultrasound localization microscopy
Context-aware deep learning enables high-efficacy localization of high concentration microbubbles for super-resolution ultrasound localization microscopy
Ultrasound localisation microscopy enables deep tissue microvascular imaging. Here, authors introduce LOCA-ULM, a deep learning pipeline enhancing localisation accuracy in high microbubble concentrations. LOCA-ULM reveals dense cerebrovascular networks and enhances the sensitivity of functional ULM.
- YiRang Shin
- , Matthew R. Lowerison
- & Pengfei Song
-
Article
| Open Access Genomic language model predicts protein co-regulation and function
Genomic language model predicts protein co-regulation and function
A gene’s function is governed by its sequence, structure and context. Here, the authors develop a genomic language model that learns contextualized functional representations from diverse and large-scale metagenomic datasets.
- Yunha Hwang
- , Andre L. Cornman
- & Peter R. Girguis
-
Article
| Open Access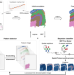 Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Recognising spatial spots’ biological identity in spatial transcriptomics remains a challenge. Here, authors introduce Pianno, a tool that helps annotate the biological structures or cell-type constructions across diverse tissues, offering new perspectives on understanding spatial transcriptomics.
- Yuqiu Zhou
- , Wei He
- & Ying Zhu
-
Article
| Open Access Early detection of emerging viral variants through analysis of community structure of coordinated substitution networks
Early detection of emerging viral variants through analysis of community structure of coordinated substitution networks
Rise of new viral strains is a major public health challenge, demanding advanced detection and forecasting methods. This study shows how examining communities within networks of viral mutations enables early detection of emerging strains.
- Fatemeh Mohebbi
- , Alex Zelikovsky
- & Pavel Skums
-
Article
| Open Access Reconstructing the evolution history of networked complex systems
Reconstructing the evolution history of networked complex systems
Evolution processes of complex networked systems in biology and social sciences, and their underlying mechanisms, still need better understanding. The authors propose a machine learning approach to reconstruct the evolution history of complex networks.
- Junya Wang
- , Yi-Jiao Zhang
- & Yanqing Hu
-
Article
| Open Access Data-driven identification of predictive risk biomarkers for subgroups of osteoarthritis using interpretable machine learning
Data-driven identification of predictive risk biomarkers for subgroups of osteoarthritis using interpretable machine learning
Osteoarthritis can be caused by multiple biological mechanisms but the drivers of disease risk are not well understood. Here, the authors use data from UK Biobank in machine learning models to identify clinical and biological markers associated with development of osteoarthritis and identify sub-groups with different risk profiles.
- Rikke Linnemann Nielsen
- , Thomas Monfeuga
- & Ramneek Gupta
-
Article
| Open Access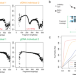 FinaleMe: Predicting DNA methylation by the fragmentation patterns of plasma cell-free DNA
FinaleMe: Predicting DNA methylation by the fragmentation patterns of plasma cell-free DNA
DNA methylation from cell-free DNA (cfDNA) can be profiled using whole genome bisulfite sequencing (WGBS). Here, the authors develop a computational method, FinaleMe, that predicts DNA methylation and tissues of-origin in cfDNA and validate its performance using paired deep and shallow-coverage whole-genome sequencing (WGS) and WGBS data.
- Yaping Liu
- , Sarah C. Reed
- & Manolis Kellis
-
Article
| Open Access PLMSearch: Protein language model powers accurate and fast sequence search for remote homology
PLMSearch: Protein language model powers accurate and fast sequence search for remote homology
Homologous protein search is one of the most commonly used methods for protein analysis. Here, authors propose PLMSearch, a search method that takes only sequences as input and can search millions of protein pairs in seconds while maintaining sensitivity comparable to SOTA structure search methods.
- Wei Liu
- , Ziye Wang
- & Shanfeng Zhu
-
Article
| Open Access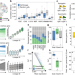 Gene-expression memory-based prediction of cell lineages from scRNA-seq datasets
Gene-expression memory-based prediction of cell lineages from scRNA-seq datasets
Combining experimental lineage tracing with single cell transcriptomics is technically demanding. Here, authors present GEMLI, a computational tool to annotate cell lineages in single cell RNA sequencing data solely based on gene expression.
- A. S. Eisele
- , M. Tarbier
- & D. M. Suter
-
Article
| Open Access DELVE: feature selection for preserving biological trajectories in single-cell data
DELVE: feature selection for preserving biological trajectories in single-cell data
Characteristic genes or proteins driving continuous biological processes are difficult to uncover from noisy single-cell data. Here, authors present DELVE, an unsupervised feature selection method to identify core molecular features driving cell fate decisions.
- Jolene S. Ranek
- , Wayne Stallaert
- & Jeremy E. Purvis
-
Article
| Open Access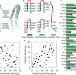 Hairpin trimer transition state of amyloid fibril
Hairpin trimer transition state of amyloid fibril
Amyloid fibrils are ordered protein assemblies implicated in neurodegenerative disease. Here the authors show that hairpin trimers can be transition states of fibril nucleation, explaining how different fibril isoforms may arise from alternative nucleation sites.
- Levent Sari
- , Sofia Bali
- & Milo M. Lin
-
Article
| Open Access Mapping cell-to-tissue graphs across human placenta histology whole slide images using deep learning with HAPPY
Mapping cell-to-tissue graphs across human placenta histology whole slide images using deep learning with HAPPY
Placenta histopathology for maternal and newborn health is highly specialised and time consuming. Here, authors present a deep learning pipeline for quantifying cells and tissues in placenta whole slide images, revealing biological and clinical insights.
- Claudia Vanea
- , Jelisaveta Džigurski
- & Christoffer Nellåker
-
Article
| Open Access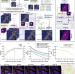 SpiDe-Sr: blind super-resolution network for precise cell segmentation and clustering in spatial proteomics imaging
SpiDe-Sr: blind super-resolution network for precise cell segmentation and clustering in spatial proteomics imaging
Imaging mass cytometry (IMC) is a powerful single-cell resolution platform for targeted spatial proteomics, but it can be constrained by imaging noise and resolution. Here, the authors propose SpiDe-Sr, a super-resolution network embedded with a denoising module for IMC spatial resolution enhancement.
- Rui Chen
- , Jiasu Xu
- & Xianting Ding
-
Article
| Open Access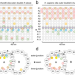 Uncovering structural themes across cilia microtubule inner proteins with implications for human cilia function
Uncovering structural themes across cilia microtubule inner proteins with implications for human cilia function
The inside surface of microtubules contains so-called microtubule inner proteins, but little is known about their identity. Here the authors use bioinformatics to identify structural motifs within this class of proteins and potential new members.
- Jens S. Andersen
- , Aaran Vijayakumaran
- & Kenneth Bødtker Schou
-
Article
| Open Access High-throughput prediction of protein conformational distributions with subsampled AlphaFold2
High-throughput prediction of protein conformational distributions with subsampled AlphaFold2
Protein dynamics, crucial for life, are difficult and expensive to predict. This study shows that AI-based structure prediction methods can be modified for rapidly predicting the conformational landscapes of proteins, with strong correlations with experimentally-measured relative state populations.
- Gabriel Monteiro da Silva
- , Jennifer Y. Cui
- & Brenda M. Rubenstein
-
Article
| Open Access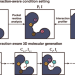 3D molecular generative framework for interaction-guided drug design
3D molecular generative framework for interaction-guided drug design
Designing a molecule that favorably binds to a protein pocket is a keystone of drug discovery. Zhung et al. devise DeepICL, which leverages the generalizable features of non-covalent protein-ligand interactions on a 3D molecular generative model, improving the quality of AI-designed molecules.
- Wonho Zhung
- , Hyeongwoo Kim
- & Woo Youn Kim
-
Article
| Open Access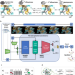 A dual diffusion model enables 3D molecule generation and lead optimization based on target pockets
A dual diffusion model enables 3D molecule generation and lead optimization based on target pockets
Structure-based generative chemistry is crucial in computer-aided drug discovery. Here, authors propose PMDM, a conditional generative model for 3D molecule generation tailored to specific targets. Extensive experiments demonstrate that PMDM can effectively generate rational bioactive molecules
- Lei Huang
- , Tingyang Xu
- & Hengtong Zhang
-
Article
| Open Access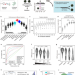 Precise prediction of phase-separation key residues by machine learning
Precise prediction of phase-separation key residues by machine learning
Understanding intracellular phase separation is essential for transcriptional control, cell fate, and disease. Here the authors report PSPHunter which accurately predicts key residues, aiding in disease-associated protein identification and mechanistic insights.
- Jun Sun
- , Jiale Qu
- & Junjun Ding
-
Article
| Open Access Predicting and improving complex beer flavor through machine learning
Predicting and improving complex beer flavor through machine learning
Perception and appreciation of food flavour depends on many factors, posing a challenge for effective prediction. Here, the authors combine extensive chemical and sensory analyses of 250 commercial Belgian beers to train machine learning models that enable flavour and consumer appreciation prediction.
- Michiel Schreurs
- , Supinya Piampongsant
- & Kevin J. Verstrepen
-
Article
| Open Access Diverging co-translational protein complex assembly pathways are governed by interface energy distribution
Diverging co-translational protein complex assembly pathways are governed by interface energy distribution
Protein complex assembly can occur co-translationally. Here, the authors uncover diverging assembly pathways and hotspot disruptions in N-terminal acetyltransferases, enzymes implicated in neurodegenerative diseases. Their model predicts co-translational assembly based on interface energy distribution.
- Johannes Venezian
- , Hagit Bar-Yosef
- & Ayala Shiber
-
Article
| Open Access Developmental progression of DNA double-strand break repair deciphered by a single-allele resolution mutation classifier
Developmental progression of DNA double-strand break repair deciphered by a single-allele resolution mutation classifier
DNA double-strand breaks (DSBs) are repaired by a hierarchically regulated network of pathways. Here, authors develop ICP for deciphering somatic DSB repair patterns in multicellular organisms and discover developmental regulation in flies and mosquitoes, enabling tracking of mutant alleles and interhomolog copying of gene cassettes.
- Zhiqian Li
- , Lang You
- & Ethan Bier
-
Article
| Open Access Multi-omic integration of microbiome data for identifying disease-associated modules
Multi-omic integration of microbiome data for identifying disease-associated modules
Here, Muller et al. introduce MintTea, a method for analyzing multi-omic microbiome data and identifying disease-associated modules comprising mixed sets of features that collectively shift in disease, offering insights into microbiome-disease interactions.
- Efrat Muller
- , Itamar Shiryan
- & Elhanan Borenstein
-
Article
| Open Access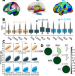 The genetic architecture of multimodal human brain age
The genetic architecture of multimodal human brain age
The biological basis of brain aging is not well understood, but it has implications for human health. Here, the authors explore the genetic basis of human brain aging, finding genetic variants, genes and potential causal relationships with disease.
- Junhao Wen
- , Bingxin Zhao
- & Christos Davatzikos
-
Article
| Open Access InPACT: a computational method for accurate characterization of intronic polyadenylation from RNA sequencing data
InPACT: a computational method for accurate characterization of intronic polyadenylation from RNA sequencing data
Intronic polyadenylation (IPA) can produce transcripts with truncated coding regions and has been implicated in diverse biological processes and diseases. Here, the authors present a computational method for the accurate delineation of IPA events using RNA-sequencing data.
- Xiaochuan Liu
- , Hao Chen
- & Yang Yang
-
Article
| Open Access Predicting nuclear G-quadruplex RNA-binding proteins with roles in transcription and phase separation
Predicting nuclear G-quadruplex RNA-binding proteins with roles in transcription and phase separation
RNA G-quadruplexes are important regulatory elements, yet our knowledge of their structure-based interactions is at present limited. Here the authors combine experimental and computational methods to develop a predictive tool, G4-FUNNIES, to estimate proteins’ RNA G4-binding propensities.
- Johanna Luige
- , Alexandros Armaos
- & Ulf Andersson Vang Ørom
-
Article
| Open Access Allele-specific transcriptional effects of subclonal copy number alterations enable genotype-phenotype mapping in cancer cells
Allele-specific transcriptional effects of subclonal copy number alterations enable genotype-phenotype mapping in cancer cells
Quantifying the impact of copy-number alterations (CNAs) on gene expression at the subclone level in cancer remains a challenge. Here, the authors develop TreeAlign, a method that integrates sample-matched single-cell DNA and RNA sequencing data to infer the impact of CNAs on subclonal gene expression.
- Hongyu Shi
- , Marc J. Williams
- & Sohrab P. Shah
-
Article
| Open Access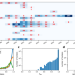 Strong positive selection biases identity-by-descent-based inferences of recent demography and population structure in Plasmodium falciparum
Strong positive selection biases identity-by-descent-based inferences of recent demography and population structure in Plasmodium falciparum
Identity-by-descent (IBD) is used to infer malaria parasite population demography, but positive selection imposed by drug pressure can bias IBD estimates. This study shows that strong selection distorts IBD distributions impacting downstream inferences and presents an approach to correct these biases.
- Bing Guo
- , Victor Borda
- & Shannon Takala-Harrison
-
Article
| Open Access Balancing competing effects of tissue growth and cytoskeletal regulation during Drosophila wing disc development
Balancing competing effects of tissue growth and cytoskeletal regulation during Drosophila wing disc development
The authors integrate computational and quantitative approaches to elucidate how organ shape arises through the interplay between multiple growth pathways through regulation of both proliferation and the cytoskeleton.
- Nilay Kumar
- , Jennifer Rangel Ambriz
- & Mark Alber
-
Article
| Open Access Tradeoffs in alignment and assembly-based methods for structural variant detection with long-read sequencing data
Tradeoffs in alignment and assembly-based methods for structural variant detection with long-read sequencing data
Long-read sequencing can greatly improve detection of genomic structural variants (SVs), and numerous methods have been developed to identify SVs using long-read data. Here the authors compare the performance of these methods and provide guidelines to aid users in selecting the most suitable tools for various scenarios.
- Yichen Henry Liu
- , Can Luo
- & Xin Maizie Zhou
-
Article
| Open Access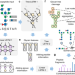 Prediction of glycopeptide fragment mass spectra by deep learning
Prediction of glycopeptide fragment mass spectra by deep learning
Deep learning has achieved a notable success in proteomics and is now emerging in glycoproteomics. Here, the authors develop a neural network-based method to predict mass spectra of intact glycopeptides and demonstrate its potential in data-dependent and data-independent acquisition glycoproteomics.
- Yi Yang
- & Qun Fang
-
Article
| Open Access The bone ecosystem facilitates multiple myeloma relapse and the evolution of heterogeneous drug resistant disease
The bone ecosystem facilitates multiple myeloma relapse and the evolution of heterogeneous drug resistant disease
Here, the authors develop a hybrid agent-based model to quantify the contributions of intrinsic cellular mechanisms and bone ecosystem factors to therapy resistance in multiple myeloma. They show that intrinsic mechanisms are essential for resistance, and that the bone microenvironment provides a protective niche that increases the likelihood.
- Ryan T. Bishop
- , Anna K. Miller
- & David Basanta
-
Article
| Open Access Accurate and rapid antibiotic susceptibility testing using a machine learning-assisted nanomotion technology platform
Accurate and rapid antibiotic susceptibility testing using a machine learning-assisted nanomotion technology platform
Sturm et. al developed a 2 to 4 h antibiotic susceptibility test based on bacterial vibrations. This diagnostic test applies to the most frequently found gram-negative bacteria in bloodstream infections and demonstrates its potential in contributing to faster treatment decisions.
- Alexander Sturm
- , Grzegorz Jóźwiak
- & Danuta Cichocka
-
Article
| Open Access Data-driven prediction of colonization outcomes for complex microbial communities
Data-driven prediction of colonization outcomes for complex microbial communities
Predicting the colonization of exogenous species in complex communities is a challenge in ecology. Here, the authors propose a data-driven approach to predict colonization outcomes and perform validation experiments in human gut microbial communities.
- Lu Wu
- , Xu-Wen Wang
- & Lei Dai
-
Article
| Open Access Local prediction-learning in high-dimensional spaces enables neural networks to plan
Local prediction-learning in high-dimensional spaces enables neural networks to plan
The task of planning a sequence of actions, and dynamically adjusting the plan in dependence of unforeseen circumstances, remains challenging for artificial intelligence frameworks. The authors introduce a learning approach inspired by cognitive functions, that demonstrates high flexibility and generalization capability in planning tasks, suitable for on-chip learning.
- Christoph Stöckl
- , Yukun Yang
- & Wolfgang Maass
-
Article
| Open Access Charting cellular differentiation trajectories with Ricci flow
Charting cellular differentiation trajectories with Ricci flow
When stem cells develop into tissues intracellular signalling is rewired, errors in this process lead to cancer. Here, authors applied tools from differential geometry made by Albert Einstein’s General Relativity to understand and predict biological network rewiring in health and disease.
- Anthony Baptista
- , Ben D. MacArthur
- & Christopher R. S. Banerji
-
Article
| Open Access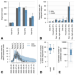 The decline of the 2022 Italian mpox epidemic: Role of behavior changes and control strategies
The decline of the 2022 Italian mpox epidemic: Role of behavior changes and control strategies
Mpox cases in Italy rapidly declined following a peak in summer 2022. Here, the authors investigate potential reasons for the decline in cases using an individual-based model of a sexual contact network of men who have sex with men.
- Giorgio Guzzetta
- , Valentina Marziano
- & Stefano Merler
-
Article
| Open Access Structure-based prediction and characterization of photo-crosslinking in native protein–RNA complexes
Structure-based prediction and characterization of photo-crosslinking in native protein–RNA complexes
Feng et al. developed a computational method PxR3D-map to jointly analyze crosslinked nucleotides and amino acids in protein-RNA complexes, which revealed key structural features underlying photocrosslinking of protein and RNA in cells.
- Huijuan Feng
- , Xiang-Jun Lu
- & Chaolin Zhang
-
Article
| Open Access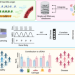 Cell type signatures in cell-free DNA fragmentation profiles reveal disease biology
Cell type signatures in cell-free DNA fragmentation profiles reveal disease biology
Deconvolution of cfDNA fragmentation benefits from cell type-specific reference data. Here, the authors create a disease agnostic cfDNA cell type of origin analysis and show it can successfully predict cell types of origin from plasma samples.
- Kate E. Stanley
- , Tatjana Jatsenko
- & Joris Robert Vermeesch
-
Article
| Open Access Systematic review and meta-analysis for a Global Patient co-Owned Cloud (GPOC)
Systematic review and meta-analysis for a Global Patient co-Owned Cloud (GPOC)
Use of cloud-based personal health records are increasing globally. Here, authors introduce the Global Patient co-Owned Cloud (GPOC) concept. The systematic review and meta-analysis examine factors like data security, efficiency, privacy, and cost. It aims to establish a scientific basis for a GPOC, which may disseminate global artificial intelligence for healthcare.
- Niklas Lidströmer
- , Joe Davids
- & Eric Herlenius
-
Article
| Open Access BASALT refines binning from metagenomic data and increases resolution of genome-resolved metagenomic analysis
BASALT refines binning from metagenomic data and increases resolution of genome-resolved metagenomic analysis
Binning is an essential step in genome-resolved metagenomic analysis in which assembled contigs originating from the same source population are clustered. However it is challenging, especially for low abundance microbial species. Here the authors introduce a toolkit that integrates multiple prominent binning tools and AI for efficient and high-resolution recovery of non-redundant bins from short- and long-read metagenomic sequencing datasets.
- Zhiguang Qiu
- , Li Yuan
- & Ke Yu
-
Article
| Open Access Enabling large-scale screening of Barrett’s esophagus using weakly supervised deep learning in histopathology
Enabling large-scale screening of Barrett’s esophagus using weakly supervised deep learning in histopathology
Diagnosis of Barrett’s esophagus depends on pathologist assessment of stained slides. Here, the authors utilise a deep learning approach to prioritize potential cases using diagnostic labels in two datasets, with the aim to improve Barrett’s screening capacity.
- Kenza Bouzid
- , Harshita Sharma
- & Javier Alvarez-Valle
-
Article
| Open Access AlphaPept: a modern and open framework for MS-based proteomics
AlphaPept: a modern and open framework for MS-based proteomics
Mass spectrometry-based proteomics faces the challenge of processing vast data amounts. Here, the authors introduce AlphaPept, an open-source, Python-based framework that offers high speed analysis and easy integration for large-scale proteome analysis.
- Maximilian T. Strauss
- , Isabell Bludau
- & Matthias Mann
-
Article
| Open Access Drivers and impact of the early silent invasion of SARS-CoV-2 Alpha
Drivers and impact of the early silent invasion of SARS-CoV-2 Alpha
The SARS-CoV-2 Alpha variant of concern emerged in the UK in late 2020 but spread internationally before it was detected. Here, the authors reconstruct the dynamics of dissemination of this variant out of the UK by combining extent of genomic sequencing, travel volume, and local epidemic dynamics in a Bayesian model.
- Benjamin Faucher
- , Chiara E. Sabbatini
- & Chiara Poletto
-
Article
| Open Access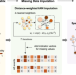 Interrogations of single-cell RNA splicing landscapes with SCASL define new cell identities with physiological relevance
Interrogations of single-cell RNA splicing landscapes with SCASL define new cell identities with physiological relevance
RNA splicing serves as a critical layer of gene expression regulation. Here, authors introduce SCASL for investigating the heterogeneity of RNA splicing landscapes at single-cell resolution, offering a novel scheme for classifying cell identities with physiological relevance.
- Xianke Xiang
- , Yao He
- & Xuerui Yang
-
Article
| Open Access The defensome of complex bacterial communities
The defensome of complex bacterial communities
Bacteria have evolved numerous innate and adaptive defence mechanisms. Here, Beavogui et al characterise the impact of biogeography, genetic mobility, and clustering in defense islands, on the defence systems of soil, marine, and human gut bacterial populations genomes.
- Angelina Beavogui
- , Auriane Lacroix
- & Pedro H. Oliveira
-
Article
| Open Access Viscosity-dependent control of protein synthesis and degradation
Viscosity-dependent control of protein synthesis and degradation
Xenopus egg extracts constitute a cell-like system for studying biochemical reactions. Here Chen and co-workers show that extract protein synthesis and degradation are differently affected by cytoplasmic concentration and viscosity.
- Yuping Chen
- , Jo-Hsi Huang
- & James E. Ferrell Jr.
-
Article
| Open Access Machine learning predictor PSPire screens for phase-separating proteins lacking intrinsically disordered regions
Machine learning predictor PSPire screens for phase-separating proteins lacking intrinsically disordered regions
Here the authors report a machine learning model, PSPire, which integrates both residue-level and structure-level features and outperforms tools in identifying phase-separating proteins lacking intrinsically disordered regions.
- Shuang Hou
- , Jiaojiao Hu
- & Yong Zhang
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 De novo diploid genome assembly using long noisy reads
De novo diploid genome assembly using long noisy reads