Article
|
Open Access
Featured
-
-
Article
| Open Access A comprehensive platform for analyzing longitudinal multi-omics data
A comprehensive platform for analyzing longitudinal multi-omics data
The analysis of longitudinal bulk and single-cell multi-omics data is a highly complex task. Here, the authors introduce PALMO, a software platform with five modules to analyse longitudinal bulk and single-cell multi-omics data, which is extensively tested in external datasets that include multiple omics modalities.
- Suhas V. Vasaikar
- , Adam K. Savage
- & Xiao-jun Li
-
Article
| Open Access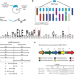 Genome mining unveils a class of ribosomal peptides with two amino termini
Genome mining unveils a class of ribosomal peptides with two amino termini
RiPP discovery has expanded the scope of post-translational modification chemistry, but genome mining of RiPP classes remains an unsolved challenge. Here, the authors employed bioinformatics and synthetic biology approaches to discover and characterize an unknown class of RiPPs, defined by an unusual amino-modified C-terminus.
- Hengqian Ren
- , Shravan R. Dommaraju
- & Huimin Zhao
-
Article
| Open Access Integrated analysis of genomic and transcriptomic data for the discovery of splice-associated variants in cancer
Integrated analysis of genomic and transcriptomic data for the discovery of splice-associated variants in cancer
Analysing the regulatory consequences of mutations and splice variants at large scale in cancer requires efficient computational tools. Here, the authors develop RegTools, a software package that can identify splice-associated variants from large-scale genomics and transcriptomics data with efficiency and flexibility.
- Kelsy C. Cotto
- , Yang-Yang Feng
- & Malachi Griffith
-
Article
| Open Access A comprehensive benchmarking with practical guidelines for cellular deconvolution of spatial transcriptomics
A comprehensive benchmarking with practical guidelines for cellular deconvolution of spatial transcriptomics
This study comprehensively benchmarks 18 state-of-the-art methods for cellular deconvolution of spatial transcriptomics and provide decision-tree-style guidelines and recommendations for method selection.
- Haoyang Li
- , Juexiao Zhou
- & Xin Gao
-
Article
| Open Access Benchmarking integration of single-cell differential expression
Benchmarking integration of single-cell differential expression
Integration of single-cell RNA sequencing data between different samples has been a major challenge for analyzing cell populations. Here the authors benchmark 46 workflows for differential expression analysis of single-cell data with multiple batches and suggest several high-performance methods under different conditions based on simulation and real data analyses.
- Hai C. T. Nguyen
- , Bukyung Baik
- & Dougu Nam
-
Article
| Open Access Data integration across conditions improves turnover number estimates and metabolic predictions
Data integration across conditions improves turnover number estimates and metabolic predictions
The construction of protein-constrained genome-scale metabolic models depends on the integration of organism-specific enzyme turnover numbers. Here, the authors show that correction of turnover numbers by simultaneous consideration of proteomics and physiological data leads to improved predictions of condition-specific growth rates.
- Philipp Wendering
- , Marius Arend
- & Zoran Nikoloski
-
Article
| Open Access Identification of a physiologic vasculogenic fibroblast state to achieve tissue repair
Identification of a physiologic vasculogenic fibroblast state to achieve tissue repair
Here, the authors report on the discovery of physiological vasculogenic fibroblasts capable of forming functional blood vessels. In vivo tissue reprogramming triggered by topical tissue nanotransfection (TNT) of a single anti-miR-200b oligonucleotide achieved therapeutic tissue vascularization.
- Durba Pal
- , Subhadip Ghatak
- & Chandan K. Sen
-
Article
| Open Access Batch alignment of single-cell transcriptomics data using deep metric learning
Batch alignment of single-cell transcriptomics data using deep metric learning
The increasing scale of single-cell RNA-seq studies presents new challenge for integrating datasets from different batches. Here, the authors develop scDML, a tool that simultaneously removes batch effects, improves clustering performance, recovers true cell types, and scales well to large datasets.
- Xiaokang Yu
- , Xinyi Xu
- & Xiangjie Li
-
Article
| Open Access Single-cell RNA sequencing reveals the effects of chemotherapy on human pancreatic adenocarcinoma and its tumor microenvironment
Single-cell RNA sequencing reveals the effects of chemotherapy on human pancreatic adenocarcinoma and its tumor microenvironment
The role of therapy in shaping the tumor microenvironment in pancreatic ductal adenocarcinoma (PDAC) remains to be explored. Here, the authors perform single-cell RNA sequencing in PDAC samples before and after chemotherapy and suggest that chemotherapy may promote resistance to immunotherapy.
- Gregor Werba
- , Daniel Weissinger
- & Diane M. Simeone
-
Article
| Open Access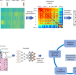 Cartography of Genomic Interactions Enables Deep Analysis of Single-Cell Expression Data
Cartography of Genomic Interactions Enables Deep Analysis of Single-Cell Expression Data
Existing genomic data analysis methods tend to not take full advantage of underlying biological characteristics. Here, the authors leverage the inherent interactions of scRNA-seq data and develop a cartography strategy to contrive the data into a spatially configured genomap for accurate deep pattern discovery.
- Md Tauhidul Islam
- & Lei Xing
-
Article
| Open Access Reanalysis of ribosome profiling datasets reveals a function of rocaglamide A in perturbing the dynamics of translation elongation via eIF4A
Reanalysis of ribosome profiling datasets reveals a function of rocaglamide A in perturbing the dynamics of translation elongation via eIF4A
The compound Rocaglamide A (RocA) is known for repressing translation initiation. Here the authors identify a dual mode of action for RocA in blocking translation initiation and elongation via eIF4A using previous datasets and new analyses.
- Fajin Li
- , Jianhuo Fang
- & Xuerui Yang
-
Article
| Open Access Decision level integration of unimodal and multimodal single cell data with scTriangulate
Decision level integration of unimodal and multimodal single cell data with scTriangulate
Single-cell genomics has expanded to measure diverse molecular modalities within the same cell. Here the authors provide a computational framework called scTriangulate to integrate cluster annotations from diverse independent sources, algorithms, and modalities to define statistically stable populations.
- Guangyuan Li
- , Baobao Song
- & Nathan Salomonis
-
Article
| Open Access Topological identification and interpretation for single-cell gene regulation elucidation across multiple platforms using scMGCA
Topological identification and interpretation for single-cell gene regulation elucidation across multiple platforms using scMGCA
A major challenge in analyzing scRNA-seq data arises from challenges related to dimensionality and the prevalence of dropout events. Here the authors develop a deep graph learning method called scMGCA based on a graph-embedding autoencoder that simultaneously learns cell-cell topology representation and cluster assignments, outperforming other state-of-the-art models across multiple platforms.
- Zhuohan Yu
- , Yanchi Su
- & Xiangtao Li
-
Article
| Open Access scMoMaT jointly performs single cell mosaic integration and multi-modal bio-marker detection
scMoMaT jointly performs single cell mosaic integration and multi-modal bio-marker detection
Many methods for single cell data integration have been developed, though mosaic integration remains challenging. Here the authors present scMoMaT, a mosaic integration method for single cell multi-modality data from multiple batches, that jointly learns cell representations and marker features across modalities for different cell clusters, to interpret the cell clusters from different modalities.
- Ziqi Zhang
- , Haoran Sun
- & Xiuwei Zhang
-
Article
| Open Access Probabilistic embedding, clustering, and alignment for integrating spatial transcriptomics data with PRECAST
Probabilistic embedding, clustering, and alignment for integrating spatial transcriptomics data with PRECAST
Methods that perform data integration are needed to analyse spatial transcriptomics data from multiple tissue slides. Here, the authors present PRECAST, an efficient data integration method for multiple spatial transcriptomics datasets with complex batch or biological effects between slides.
- Wei Liu
- , Xu Liao
- & Jin Liu
-
Article
| Open Access Simultaneous profiling of histone modifications and DNA methylation via nanopore sequencing
Simultaneous profiling of histone modifications and DNA methylation via nanopore sequencing
The interplay between histone modifications and DNA methylation plays a crucial role in establishing and maintaining the epigenomic landscape. Here, the authors develop a nanopore sequencing based method for mapping histone modifications and DNA methylation from native, long, single DNA molecules.
- Xue Yue
- , Zhiyuan Xie
- & Yimeng Yin
-
Matters Arising
| Open AccessReply to: A balanced measure shows superior performance of pseudobulk methods in single-cell RNA-sequencing analysis
- Kip D. Zimmerman
- , Ciaran Evans
- & Carl D. Langefeld
-
Matters Arising
| Open Access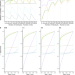 A balanced measure shows superior performance of pseudobulk methods in single-cell RNA-sequencing analysis
A balanced measure shows superior performance of pseudobulk methods in single-cell RNA-sequencing analysis
- Alan E. Murphy
- & Nathan G. Skene
-
Article
| Open Access Deep transfer learning enables lesion tracing of circulating tumor cells
Deep transfer learning enables lesion tracing of circulating tumor cells
Liquid biopsy offers great promise for noninvasive cancer diagnostics, while the lack of adequate target characterization and analysis hinders its wide application. Here, the authors design a transfer learning-based algorithm to transfer lesion labels from the primary cancer cell atlas to circulating tumor cells.
- Xiaoxu Guo
- , Fanghe Lin
- & Jia Song
-
Article
| Open Access AlphaPeptDeep: a modular deep learning framework to predict peptide properties for proteomics
AlphaPeptDeep: a modular deep learning framework to predict peptide properties for proteomics
Deep learning (DL) has been frequently used in mass spectrometry-based proteomics but there is still a lot of potential. Here, the authors develop a framework that enables building DL models to predict arbitrary peptide properties with only a few lines of code.
- Wen-Feng Zeng
- , Xie-Xuan Zhou
- & Matthias Mann
-
Article
| Open Access Library adaptors with integrated reference controls improve the accuracy and reliability of nanopore sequencing
Library adaptors with integrated reference controls improve the accuracy and reliability of nanopore sequencing
Adding library adaptors to DNA samples is an essential step in preparing samples for next-generation sequencing. Here, Gunter et al. describe the development of Control Library Adaptors (CAPTORs), that correct sequencing errors and normalise quantitative biases in Nanopore libraries.
- Helen M. Gunter
- , Scott E. Youlten
- & Tim R. Mercer
-
Article
| Open Access Online single-cell data integration through projecting heterogeneous datasets into a common cell-embedding space
Online single-cell data integration through projecting heterogeneous datasets into a common cell-embedding space
Integrative analyses of single-cell datasets are facing new challenges as data size and complexity grow. Here the authors present SCALEX, which projects cells from different datasets into a common latent space, allowing accurate online integration as well as cross-referencing with atlas-scale data.
- Lei Xiong
- , Kang Tian
- & Qiangfeng Cliff Zhang
-
Article
| Open Access Elucidating tumor heterogeneity from spatially resolved transcriptomics data by multi-view graph collaborative learning
Elucidating tumor heterogeneity from spatially resolved transcriptomics data by multi-view graph collaborative learning
Multi-view graph approaches could enhance the analysis of tissue heterogeneity in spatial transcriptomics. Here, the authors develop the Spatial Transcriptomics data analysis by Multiple View Collaborative-learning - stMVC - framework, and apply it to detect spatial domains and cell states in brain and tumor tissues.
- Chunman Zuo
- , Yijian Zhang
- & Luonan Chen
-
Article
| Open Access Robust data storage in DNA by de Bruijn graph-based de novo strand assembly
Robust data storage in DNA by de Bruijn graph-based de novo strand assembly
DNA data storage is a rapidly developing technology with great potential due to its high density, long-term durability, and low maintenance cost. Here the authors present a strand assembly algorithm (DBGPS) using de Bruijn graph and greedy path search.
- Lifu Song
- , Feng Geng
- & Ying-Jin Yuan
-
Article
| Open Access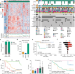 Molecular characterization of colorectal cancer related peritoneal metastatic disease
Molecular characterization of colorectal cancer related peritoneal metastatic disease
Colorectal cancer can lead to the development of peritoneal metastases, which are associated with worse disease outcome. Here, the authors characterize peritoneal metastases from 52 patients using RNA-seq and mutational sequencing and show a distinct molecular subtype.
- Kristiaan J. Lenos
- , Sander Bach
- & Louis Vermeulen
-
Article
| Open Access TidyMass an object-oriented reproducible analysis framework for LC–MS data
TidyMass an object-oriented reproducible analysis framework for LC–MS data
Reproducibility, traceability, and transparency have been long-standing issues in metabolomics data analysis. Here, the authors present tidyMass, an R-based computational framework that allows designing traceable, shareable, and reproducible data processing and analysis workflows for untargeted metabolomics.
- Xiaotao Shen
- , Hong Yan
- & Michael P. Snyder
-
Article
| Open Access An expanded reference map of the human gut microbiome reveals hundreds of previously unknown species
An expanded reference map of the human gut microbiome reveals hundreds of previously unknown species
Here, Leviatan et al. produce 241,118 genome assemblies to produce a new human gut microbiome reference set of 3,594 species genomes, of which 310 represent previously undescribed species, making the catalog a valuable resource for further research.
- Sigal Leviatan
- , Saar Shoer
- & Eran Segal
-
Article
| Open Access Biomarkers of nanomaterials hazard from multi-layer data
Biomarkers of nanomaterials hazard from multi-layer data
Nanomaterials have a range of potential applications, however, toxicity remains a concern, limiting application and requiring extensive testing. Here, the authors report on a predictive framework made using a range of tests linking materials properties with toxicity, allowing the prediction of toxicity from physiochemical and biological properties.
- Vittorio Fortino
- , Pia Anneli Sofia Kinaret
- & Dario Greco
-
Article
| Open Access ChIP-Hub provides an integrative platform for exploring plant regulome
ChIP-Hub provides an integrative platform for exploring plant regulome
A comprehensive data portal to explore plant regulomes is still unavailable. Here, the authors develop a web-based platform ChIP-Hub in the ENCODE standards and demonstrate its applications in the identification of hierarchical regulatory network, tissue-specific chromatin dynamics, putative enhancers and chromatin states.
- Liang-Yu Fu
- , Tao Zhu
- & Dijun Chen
-
Article
| Open Access Deep learning of a bacterial and archaeal universal language of life enables transfer learning and illuminates microbial dark matter
Deep learning of a bacterial and archaeal universal language of life enables transfer learning and illuminates microbial dark matter
Computational methods to analyse microbial systems rely on reference databases which do not capture their full functional diversity. Here the authors develop a deep learning model and apply it using transfer learning, creating biologically useful models for multiple different tasks.
- A. Hoarfrost
- , A. Aptekmann
- & Y. Bromberg
-
Article
| Open Access Single-cell transcriptomics identifies Mcl-1 as a target for senolytic therapy in cancer
Single-cell transcriptomics identifies Mcl-1 as a target for senolytic therapy in cancer
Cell senescence remains a barrier to tumor elimination in many cancers. Here, the authors use single cell RNA-seq to identify a role for Mcl-1 in senescent cell survival, and show that Mcl-1 inhibition may be an effective therapeutic strategy.
- Martina Troiani
- , Manuel Colucci
- & Andrea Alimonti
-
Article
| Open Access Normalizing and denoising protein expression data from droplet-based single cell profiling
Normalizing and denoising protein expression data from droplet-based single cell profiling
Current single cell protein expression profiling approaches come with substantial measurement noise. Here the authors discover the sources of this noise and develop a denoising algorithm that improves data quality and downstream applications.
- Matthew P. Mulè
- , Andrew J. Martins
- & John S. Tsang
-
Article
| Open Access PyUUL provides an interface between biological structures and deep learning algorithms
PyUUL provides an interface between biological structures and deep learning algorithms
While artificial intelligence (AI) is quickly becoming ubiquitous, biology still suffers from the lack of interfaces connecting biological structures and modern AI methods. Here, the authors report PyUUL, a library to translate biological structures into 3D differentiable tensorial representations.
- Gabriele Orlando
- , Daniele Raimondi
- & Frederic Rousseau
-
Article
| Open Access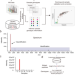 SMAP is a pipeline for sample matching in proteogenomics
SMAP is a pipeline for sample matching in proteogenomics
Sample mix-up is a potential problem in large-scale omic studies due to the complexity of sample processing. Here, the authors present a pipeline for sample matching in proteogenomics to verify sample identity and ensure data integrity.
- Ling Li
- , Mingming Niu
- & Xusheng Wang
-
Article
| Open Access Single cell transcriptomic landscape of diabetic foot ulcers
Single cell transcriptomic landscape of diabetic foot ulcers
Diabetic foot ulcers (DFUs) remain a complication of diabetes that are difficult to heal and lead to disability. Here the authors use single-cell RNA-sequencing and spatial transcriptomics to characterize the DFU cellular landscape and identify a population of fibroblasts that is associated with successful wound closure.
- Georgios Theocharidis
- , Beena E. Thomas
- & Manoj Bhasin
-
Article
| Open Access RNA modifications detection by comparative Nanopore direct RNA sequencing
RNA modifications detection by comparative Nanopore direct RNA sequencing
Nanopore direct RNA Sequencing data contain information about the presence of RNA modifications, but their detection poses substantial challenges. Here the authors introduce Nanocompore, a new methodology for modification detection from Nanopore data.
- Adrien Leger
- , Paulo P. Amaral
- & Tony Kouzarides
-
Article
| Open Access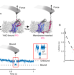 Monitoring the binding and insertion of a single transmembrane protein by an insertase
Monitoring the binding and insertion of a single transmembrane protein by an insertase
The insertion and folding nascent or fully synthesized polypeptides into membranes is assisted by insertases. Here, the authors use a range of biophysical approaches to provide molecular details of how the transmembrane insertase YidC facilitates the insertion a protein into a phospholipid membrane.
- Pawel R. Laskowski
- , Kristyna Pluhackova
- & Daniel J. Müller
-
Article
| Open Access Identification of the cross-strand chimeric RNAs generated by fusions of bi-directional transcripts
Identification of the cross-strand chimeric RNAs generated by fusions of bi-directional transcripts
Gene fusion, trans-splicing or transcription read-through contributes to generation of chimeric RNA. Here the authors develop a pipeline to identify non-canonical type of chimeric RNAs called cross-strand chimeric RNA (cscRNA), which are fused between two precursor RNAs transcribed from the opposite DNA strands.
- Yuting Wang
- , Qin Zou
- & Xuerui Yang
-
Article
| Open Access Development of a quantitative prediction algorithm for target organ-specific similarity of human pluripotent stem cell-derived organoids and cells
Development of a quantitative prediction algorithm for target organ-specific similarity of human pluripotent stem cell-derived organoids and cells
Quantitative methods to assess the quality of hPSC-derived organoids have not been developed. Here they present a prediction algorithm to assess the transcriptomic similarity between hPSC-derived organoids and the corresponding human target organs and perform validation on lung bud organoids, antral gastric organoids, and cardiomyocytes.
- Mi-Ok Lee
- , Su-gi Lee
- & Hyun-Soo Cho
-
Article
| Open Access Strainberry: automated strain separation in low-complexity metagenomes using long reads
Strainberry: automated strain separation in low-complexity metagenomes using long reads
Existing long-read de novo assembly methods can partially, but not completely, separate strains. Here, the authors develop Strainberry, a metagenome assembly bioinformatic pipeline that exclusively uses longread data to accurately separate and reconstruct strain genomes from single-sample low-complexity microbiomes.
- Riccardo Vicedomini
- , Christopher Quince
- & Rayan Chikhi
-
Article
| Open Access CRISPECTOR provides accurate estimation of genome editing translocation and off-target activity from comparative NGS data
CRISPECTOR provides accurate estimation of genome editing translocation and off-target activity from comparative NGS data
The control of off-target activity is a challenge for adapting CRISPR to therapeutic use. Here the authors present CRISPECTOR, a software tool to detect, evaluate and quantify editing activity, including translocations, from NGS data.
- Ido Amit
- , Ortal Iancu
- & Zohar Yakhini
-
Article
| Open Access Osteocyte transcriptome mapping identifies a molecular landscape controlling skeletal homeostasis and susceptibility to skeletal disease
Osteocyte transcriptome mapping identifies a molecular landscape controlling skeletal homeostasis and susceptibility to skeletal disease
Osteocytes are the master regulatory cells within the skeleton. Here, the authors map the transcriptome of osteocytes from diverse skeletal sites, ages and between sexes and identify an osteocyte transcriptome signature associated with rare skeletal disorders and common complex skeletal diseases.
- Scott E. Youlten
- , John P. Kemp
- & Peter I. Croucher
-
Article
| Open Access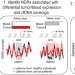 Tissue-specific cell-free DNA degradation quantifies circulating tumor DNA burden
Tissue-specific cell-free DNA degradation quantifies circulating tumor DNA burden
Circulating tumour DNA (ctDNA) represents a non-invasive option to monitor cancer progression. Here, the authors perform deep sequencing of plasma cell-free DNA, and find that nucleosome-dependent cfDNA degradation at 6 specific regulatory regions is predictive of ctDNA burden.
- Guanhua Zhu
- , Yu A. Guo
- & Anders J. Skanderup
-
Article
| Open Access Identifying transposable element expression dynamics and heterogeneity during development at the single-cell level with a processing pipeline scTE
Identifying transposable element expression dynamics and heterogeneity during development at the single-cell level with a processing pipeline scTE
How transposable elements (TE) contribute to cell fate changes is unclear. Here, the authors generate a pipeline to quantify TE expression from single cell data. They show the dynamic expression of TEs from gastrulation to somatic cell reprogramming and human disease
- Jiangping He
- , Isaac A. Babarinde
- & Jiekai Chen
-
Article
| Open Access Comprehensive analysis of single cell ATAC-seq data with SnapATAC
Comprehensive analysis of single cell ATAC-seq data with SnapATAC
Single cell analysis of transposase-accessible chromatin is deepening our understanding on the origins of cellular diversity, yet methods are limited by data sparsity. Here, the authors introduce SnapATAC, a pipeline to resolve cellular heterogeneity and reveal candidate regulatory elements across different cell populations.
- Rongxin Fang
- , Sebastian Preissl
- & Bing Ren
-
Article
| Open Access Uniform genomic data analysis in the NCI Genomic Data Commons
Uniform genomic data analysis in the NCI Genomic Data Commons
The Genomic Data Commons repository contains genomic, epigenomic, proteomic and clinical data from the TCGA and TARGET datasets. Here, the authors describe the analysis methods for how these divergent datasets were integrated together.
- Zhenyu Zhang
- , Kyle Hernandez
- & Robert L. Grossman
-
Article
| Open Access The molecular basis of socially mediated phenotypic plasticity in a eusocial paper wasp
The molecular basis of socially mediated phenotypic plasticity in a eusocial paper wasp
Connecting genotypes to complex social behaviour is challenging. Taylor et al. use machine learning to show a strong response of caste-associated gene expression to queen loss, wherein individual wasp’s expression profiles become intermediate between queen and worker states, even in the absence of behavioural changes.
- Benjamin A. Taylor
- , Alessandro Cini
- & Seirian Sumner
-
Article
| Open Access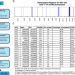 AutoMap is a high performance homozygosity mapping tool using next-generation sequencing data
AutoMap is a high performance homozygosity mapping tool using next-generation sequencing data
Homozygosity mapping is a useful tool for identifying candidate mutations in recessive conditions, however application to next generation sequencing data has been sub-optimal. Here, the authors present AutoMap, which efficiently identifies runs of homozygosity in whole exome/genome sequencing data.
- Mathieu Quinodoz
- , Virginie G. Peter
- & Carlo Rivolta
-
Article
| Open Access Cellular Heterogeneity–Adjusted cLonal Methylation (CHALM) improves prediction of gene expression
Cellular Heterogeneity–Adjusted cLonal Methylation (CHALM) improves prediction of gene expression
Here, the authors introduce Cell Heterogeneity–Adjusted cLonal Methylation (CHALM) as a methylation quantification method that considers the heterogeneity of sequenced bulk cells. They apply CHALM to methylation datasets to detect differentially methylated genes that exhibit distinct biological functions supporting underlying mechanisms.
- Jianfeng Xu
- , Jiejun Shi
- & Wei Li

 Characterization of genome-wide STR variation in 6487 human genomes
Characterization of genome-wide STR variation in 6487 human genomes