Abstract
The domain architecture of a large RNA can help explain and/or predict folding, function, biogenesis and evolution. We offer a formal and general definition of an RNA domain and use that definition to experimentally characterize the rRNA of the ribosomal small subunit. Here the rRNA comprising a domain is compact, with a self-contained system of molecular interactions. A given rRNA helix or stem-loop must be allocated uniquely to a single domain. Local changes such as mutations can give domain-wide effects. Helices within a domain have interdependent orientations, stabilities and interactions. With these criteria we identify a core domain (domain A) of small subunit rRNA. Domain A acts as a hub, linking the four peripheral domains and imposing orientational and positional restraints on the other domains. Experimental characterization of isolated domain A, and mutations and truncations of it, by methods including selective 2′OH acylation analyzed by primer extension and circular dichroism spectroscopy are consistent with our architectural model. The results support the utility of the concept of an RNA domain. Domain A, which exhibits structural similarity to tRNA, appears to be an essential core of the small ribosomal subunit.
Similar content being viewed by others
Introduction
The ribosome is a ribonucleoprotein complex that conducts one of life’s universal processes, which is the synthesis of proteins. The large ribosomal subunit (LSU) contains the peptidyl transferase center (PTC) and catalyzes transpeptidation. The small ribosomal subunit (SSU) contains the decoding center and reads messenger RNA (mRNA). Much of ribosomal function is performed by ribosomal RNAs (rRNAs)1,2 while the ribosomal proteins act primarily as structural stabilizers3. Our understanding of translation has advanced over the last decade and a half with the explosion in sequences and by the determination of three-dimensional structures2,4,5,6. X-ray crystallography and cryo-electron microscopy (Cryo-EM) have provided atomic resolution structures of ribosomes from all three primary branches of the tree of life2,6,7,8.
In ancient stepwise evolutionary processes, the ribosome acquired capabilities for RNA folding, catalysis, subunit association, correlated subunit evolution, decoding, and energy transduction9. Understanding function and evolution of the ribosome requires defining and recognizing secondary elements, motifs and domains. RNA domains are ruled by a special logic; folding is driven by complementary sidechains10, that form small, independent folds, primarily stem-loops. The RNA backbone is self-repulsive; phosphate interactions are mediated by cationic cofactors. By contrast, protein domains are large integrated units that fold cooperatively11, using favorable interactions between backbone atoms to create hydrophilic surfaces and hydrophobic cores12,13.
To understand and explain the ribosome and its domain structure we use the following general definition. (i) The RNA comprising a domain is compact and modular, with a self-contained and integrated system of molecular interactions. (ii) Any given RNA helix or stem-loop is contained uniquely within a single domain. (iii) Local changes such as mutations or metal binding can have domain-wide effects. (iv) Molecular interactions between stem-loops within an RNA domain dictate their orientations, stabilities and interactions to be interdependent. (v) An RNA domain has the capacity to fold autonomously when excised from the surrounding RNA.
The accepted canonical secondary structure of the SSU rRNA is based upon covariation14, chemical modification and RNase digestion15 and three dimensional structures16,17; here we are not proposing changes to the SSU rRNA historical secondary structure. We present an altered domain model for the SSU rRNA with a revised allocation of secondary structural elements to domains. We propose that an organizational hub (‘domain A’, Fig. 1) links to peripheral domains (the central domain, 3′M domain, 3′m domain, and the 5′ domain). Each peripheral domain connects to domain A by a spoke. The revised domain model differs from the historical domain model, in which the peripheral domains link directly to each other at a common origin and several helices participate in multiple domains8,15,17. In our revised model, with domain A as a nexus, each helix is allocated to a single domain. Here we experimentally test predictions of this domain model.
(a) Secondary structure of the T. thermophilus SSU rRNA colored by domains. Domain A is black, the 5′ domain is yellow, the central domain is red, the 3′ major domain is blue, and the 3′ minor domain is green. (b) Three-dimensional structure of the SSU rRNA (PDB ID 4V51)6. The rRNA is represented in ribbon, except for domain A, which is in space filling representation. The domains in the three-dimensional representation are colored by same scheme as in the secondary structure. (c) Space filling representation of domain A alone. Helix numbers are indicated. (d-f) A series of 90° rotations of the SSU rRNA.
This domain model has utility, and explains some dynamical properties of the SSU. The spokes are relatively flexible, allowing the domains to move relative to each other during initiation and translocation18,19. Helix 3 is the spoke linking domain A to the 5′ domain. Helix 19 is the spoke linking the central domain while helix 28 is the spoke linking the 3′ major domain. The 3′ end of domain A is the spoke linking 3′ minor domain (Figs 1, 2, 3). Domain A incorporates the central pseudoknot (CPK)20,21,22,23 and consists of helices 1, 2, 3, 19, 27, and 28 (Fig. 3).
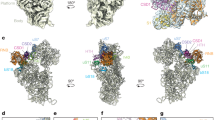
(a) Three-dimensional ribbon representation structure of T. thermophilus :SSU rRNA colored by domains. Domain A is black, the 5′ domain is yellow, the 3′ minor domain is green, the 3′ major domain is red, and the central domain is blue. (b) Three dimensional structure of domain A showing the orientations of the helical spokes that radiate from it. The two perpendicular arcs indicated by the arrows, show how domain A acts a hub that organizes the SSU. (c) Ribbon representation showing close association between secondary elements within the domain A. Nucleotides are indicated as red sticks. (d) Stick representation showing some of the molecular interactions that maintain the integrity of domain A. Hydrogen bonding between the nucleotides is shown as dotted lines. Nucleotides are labeled as in panels (c,e). (e) Secondary structure of domain A illustrating how the other domains radiate from it. The colored arrows correspond to linkages to the peripheral domains. Helices (black) and nucleotide numbers (salmon) are indicated. The box around the red nucleotides in panel (e) indicates nucleotides highlighted in panels (d,c). The black dot in helix 27 of panel (e) corresponds the black dot at the head of the arrow in panel (b). This portion of helix 27 contains the bridge B2c that contacts the large ribosomal subunit. Panels (a–d) are rotated 180° relative to panels (b,c) of Fig. 1.
(a) Secondary structure of domain A. Domain A is black. The linkers that connect the domain A fragments to form a single RNA polymer are pink while the remainder of the :SSU rRNA is grey. (b) Three dimensional ribbon representation of domain AISO model colored as in panel (a). Helix numbers are indicated.
Domain A imposes orientational and positional restraints on the other domains, which are depicted by arcs in (Figure 2). Helices 3 and 19 of domain A form one arc and helices 27 and 28 form another arc. These two orthogonal arcs intersect within the central pseudoknot (Fig. 2b). The intersecting arcs position the four peripheral domains. Nucleotides at the 5′ end of :SSU rRNA (nucleotides 9 to 13) interact with both arcs and stabilize their relative orientation. The molecular interactions that stabilize the intersecting arcs relative to each other are illustrated in Fig. 2c,d. Universally conserved nucleotides are shown in Supplementary Fig. 6. Small changes in domain A are propagated into larger motions of the peripheral domains during translocation19.
One goal here is to test this domain model. Therefore we isolated domain A from the rest of the :SSU rRNA. We refer to isolated domain A as “domain AISO” (Fig. 3). To form domain AISO as a single RNA polymer, we linked rRNA fragments together with three stem-loops (rGGCGUAAGCC), within helices 3, 19, and 28 (Fig. 3). The stem loops replace the connections between domain A and the four peripheral domains. The stem loops are intended to render domain A independent of the surrounding RNA without influencing its structure, especially its tertiary structure. We characterized domain AISO and mutations and truncations of domain AISO by methods including selective 2′OH acylation analyzed by primer extension (SHAPE) and circular dichroism (CD) spectroscopy. In addition, we observe that the three-dimensional structure of domain A has analogy in other biological RNAs.
Results
Folding of domain AISO
Domain A appears to satisfy the criteria of an RNA domain. Domain AISO is characterized here by SHAPE reactivity and CD spectroscopy. We determine effects of mutations and of added Mg2+. We compare the SHAPE reactivity of domain AISO with that of the same rRNA elements within the intact SSU, previously published by Weeks and coworkers24.
Three-dimensional and secondary structures can be probed with SHAPE. Paired nucleotides, in double-stranded regions, are less reactive to the SHAPE reagent than unpaired nucleotides in loops, bulges and single strands25. Nucleotides involved in tertiary and Mg2+ interactions change reactivity upon the addition of Mg2+26,27,28,29. The data suggest that in the presence of Na+ alone, domain AISO forms helices 1, 2, 3, 19, 27 and 28 (Fig. 4a). For helices 1, 3 and 19, the duplex regions are unreactive and the loop regions are reactive. High reactivity of nucleotide C31 suggests a defect near the loop of helix 3. Helix 27 shows the same anomalous pattern of reactivity in domain AISO as in the intact SSU (Supplementary Fig. 8).
Base pairs predicted from the secondary structure of the intact SSU are indicated by black lines. Base pairs in the linkers are indicated by red lines. Helix and nucleotide numbers are indicated. (a) Absolute SHAPE reactivity of domain AISO in the presence of Na+ only (250 mM). The red circles indicate high reactivity while the blue circles indicate low reactivity. The color scale is shown in the outbox. (b) Difference in SHAPE reactivity upon addition of Mg2+ (2 mM) (c) Difference in SHAPE reactivity upon mutation of C18 to A18. (d) Difference in SHAPE reactivity upon excision of helix 28. For panels b-c, red indicates a increase in reactivity, while blue indicates a decrease. Green indicates no change. The coloring scheme is shown in the outbox. Shape data for mutant and truncated domain AISO were acquired in the presence of both Na+ and Mg2+. Data were not obtained for the uncolored nucleotides. The primer binding tail is omitted for clarity. The full sequence of the construct is shown in the Supplementary Information (Supplementary Fig. 7).
Helices 2 and 28 are anomalously reactive in domain AISO, consistent with their anomalous reactivity in the intact SSU24,30. Nucleotides involved in base triples in the intact SSU (nucleotides G9, U20, and G22) show suppressed reactivity in domain AISO. The 5′ terminus of domain AISO (which is also the terminus of the :SSU rRNA) shows elevated SHAPE reactivity as expected of unstructured RNA. Similarly, the single-stranded nucleotides between stems 3 and 19 (A45, U46, U47) have higher reactivity than the flanking stems.
Mg2+ ions appear to stabilize domain AISO and facilitate folding to the native state. Monovalent cations generally allow RNAs to form secondary structures and a subset of tertiary interactions. Divalent cations are required for complete folding to the native state31,32. Here we used CD spectroscopy along with SHAPE to characterize the effects of divalent cations (Fig. 5). The addition of Mg2+ to domain AISO increases the intensity of the diagnostic CD band at 265 nm. The intensity increases over the range of [Mg2+] from 0 to 700 μM after which it plateaus. These Mg2+ effects on domain AISO are similar to those of well-characterized globular RNAs such as tRNA33 and P4-P6 of the Tetrahymena group I ribozyme34.
(a) Mg2+ titration of domain AISO rRNA (solid black), the C18A mutant of domain AISO (dashed blue), and domain AISO rRNA with helix 28 excised (dotted red). Mg2+ concentration is plotted versus the intensity of the diagnostic CD peak (265 nm). (b) CD spectra of the same series of RNAs in the presence of 1.0 mM Mg2+. The outbox shows a close-up of the 265 nm peak. Initial rRNA samples were depleted in Mg2+ ions.
The CD results are consistent with SHAPE reactivities. Mg2+ has subtle but widely distributed effects on the SHAPE reactivity of domain AISO. Mg2+ is expected to influence SHAPE reactivities of nucleotides that directly contact Mg2+ or are involved in Mg2+-dependent tertiary interactions. This pattern of Mg2+-dependent SHAPE reactivity has previously been observed for tRNA, RNase P, the P4-P6 domain of the Tetrahymena Group I intron and Domain III of the 23S rRNA27,28,29,35,36. Upon the addition of Mg2+, nucleotides in domain AISO show slight overall decreases in SHAPE reactivity while some loop regions and bulges show increases (Fig. 4b and Supplementary Fig. 1). Reactivity of nucleotides A16 and C31 drop upon addition of Mg2+ suggesting that correct folding of Helix 3 requires Mg2+. Based on the intact SSU, A16 is expected to interact directly with a Mg2+ ion in the native structure6. Indeed, A16 shows the greatest change in SHAPE reactivity of any site in domain AISO upon addition of Mg2+.
Helix 28 is an essential component of domain A
The CPK17 contains helices 1 and 2 (Fig. 3). We anticipated that the structure and stability of the CPK, and of domain AISO, should be dependent on helix 28, because it forms a continuous stack with helix 2 in the intact SSU (Supplementary Fig. 4) and in our model of domain AISO. If our model is correct, then helix 28 contributes globally to the stability of domain AISO. Therefore, we have determined the effect of excision of helix 28 from domain AISO.
Global changes in structure are caused by excision of helix 28. Changes in SHAPE reactivity are distributed throughout domain AISO (Fig. 4d and Supplementary Fig. 2). Reactivity increases near the 5′ terminus. Within helix 1, increases in SHAPE reactivity suggest disruption of base pairs G9-C25, A10-U24, G11-C23, U12-G22, and U13-U20 (Fig. 4d).
Furthermore, it appears that base pairing is precluded between U14 and A16 in both the intact SSU6,17 and in domain AISO. These nucleotides are in a loop region in the native structure, and show a higher SHAPE reactivity than other sites in the CPK (Fig. 4a). However, when helix 28 is omitted from domain AISO, U14 and A16 decrease in reactivity (Fig. 4d), suggesting non-native pairing interactions.
Domain-wide effects from the omission of helix 28 from domain AISO are revealed by CD spectroscopy. Changes in the CD spectrum of domain AISO upon addition of Mg2+ are diminished by excision of helix 28. Figure 5a demonstrates that changes in CD spectra after addition of Mg2+ are lessened by approximately 50% for domain AISO lacking helix 28 compared to intact domain AISO. The diagnostic 265 nm peak does not reach full intensity in the absence of helix 28 (Fig. 5, Supplementary Figs. 5 and 9). The combined SHAPE and CD data suggest that formation of the native folded state of domain AISO is dependent on helix 28, supporting our domain model.
A single mutation of the central pseudoknot impacts the entire domain
Pleij22 and Brink23 demonstrated that a C18A mutation within the CPK inhibits translation by affecting subunit assembly. This mutation is expected to disrupt the C18-G102 base pair. We mutated C18 to A in domain AISO. This mutation is seen to cause domain-wide effects on the structure. The C18A mutation lowers the general SHAPE reactivity of the domain AISO and causes specific changes in helix 2 (U20), helix 19 (C65), helix 27 (U90) and helix 28 (G107, A108) (Fig. 4c and Supplementary Fig. 3). In addition, the unusually high SHAPE reactivities of helix 27 in domain AISO (here) are consistent with those in the assembled SSU (McGinnis and Weeks)24 (Supplementary Fig. 8).
The C18A mutation affects the CD spectra of domain AISO. The C18A mutation, like helix 28 excision, lessens the effect on Mg2+ on the intensity of the 265 nm band by 50% (Fig. 5). These results indicate that domain-wide effects can be incurred by changes in sequence even if the number of nucleotides mutated is small (the C18A mutation changes 1 nucleotide while helix 28 truncation changes ~30 nucleotides compared to intact domain AISO). In sum, the data appear to support our domain model of the SSU rRNA.
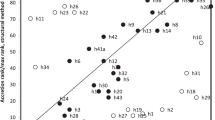
The structure of domain A is conserved in all ribosomes. We superimposed SSU rRNAs from bacterial and eukaryotic domains of life, including T. thermophilus, E. coli, S. cerevisiae, D. melanogaster, and H. sapiens (Fig. 6)6,7,37,38. The root-mean square deviation (RMSD) of backbone atoms of domain A in this superimposition is only 0.78 Å for ribosomes in different domains of life in the same translational state (Supplementary Table 1), consistent with a high degree of conservation of conformation. The greatest deviations are seen in the 5′ terminal region, which is single-stranded (Fig. 6). In addition, we have aligned sequences from 134 species from all three domains of life, and have calculated mutational Shannon entropies. For most of domain A, the sequences are universally conserved, with very low Shannon entropies. The sequences are most divergent in helix 3 and in the 5′ single stranded end (Supplementary Fig. 6).
Discussion
The SSU is a central assembly of all cellular life. The architecture of the SSU has implications for ribosomal function and evolution. Here, we use high-resolution structural information to propose a domain architecture of the SSU rRNA, and have constructed an experimental system to test predictions of the domain model.
We propose a SSU architecture in which four peripheral rRNA domains radiate from a central core, here called domain A (Fig. 1). The SSU is dendritic in structure, in contrast to the monolithic LSU. Domain A is an autonomous core at the structural and functional center of the SSU. Domain A, which includes the CPK, is a hub that connects to the peripheral SSU domains by helical spokes.
To help determine if domain A meets the formal criteria of a domain, we evaluated domain AISO, an experimental model of domain A. We investigated the Mg2+-dependence of SHAPE reactivity and CD spectra of domain AISO and several informative sequence variants. SHAPE and CD experiments suggest compact tertiary folding of domain AISO rRNA to a near-native state in the presence of Mg2+ ions. A C18A mutation or excision of helix 28 causes domain-wide effects. The results of experiments described here support the integrity of domain A, and our domain architecture of the SSU rRNA. The CPK is crucial for biogenesis of the SSU, for stability of the assembled subunits, and for initiation of translation20,21,22,23.
Domain A exhibits certain similarities in structure with tRNA (Fig. 7 and Supplementary Video 1). Similarities in structure to tRNA have previously been observed in select elongation factors, viral RNAs and bacterial non-coding RNA39,40,41,42,43. The similarity of domain A with tRNA lays in the arrangement and local conformations of helices 1, 2, and 27. Helices 1 and 2 are coaxial, and are at right angles to helix 27, giving a L-shape structure. Helix 27 of domain A is a close approximation of the anti-codon stem loop. In this region domain A is very similar to valine tRNA, with correct positioning of the CAA anticodon. However, a significant difference between tRNA and domain A is seen when helix 27 is superimposed on the anticodon stem loop; helices 1 and 2 are offset relative to the acceptor and T-stems of tRNA. Ramakrishnan previously noted a similar structural similarity in the anticodon loop of tRNA and helix 6 of SSU rRNA44. The 5′ end of the SSU rRNA is a rough approximation of the tRNA amino acid acceptor stem, which is formed by the 3′ end of the tRNA. The relevant nucleotides of the SSU rRNA are universally conserved (Supplementary Fig. 6) and are involved in intersubunit bridge B2c via A-minor interactions45,46. Where the CCA amino acid acceptor end of the tRNA comprises a 3′ terminus, the corresponding region of domain A core rRNA contains a 5′ terminus.
tRNA is red and domain A is blue. (a) Superimposition of helices 1, 2, and 27 of domain A with tRNA (tRNA is from PDB ID: 4V51)6. Both domain A and tRNA form L-shaped structures. The anticodon stem-loop superimposes on the helix 27 stem-loop. View down the CCA stem of tRNA shows that it is offset from helices 1 and 2 of domain A. (b–d) These views show the 90° rotations on y-axis. (e,f) These views show 180° rotations on x-axis.
In sum, we propose a predictive model of SSU architecture by defining domain A as a hub connecting to the peripheral domains. We show that the domain concept is applicable and useful for understanding the SSU. Domain A plays a crucial role in SSU structure and function, forming a scaffold that links to each of the other SSU domains and is an evolutionary ancestor to the SSU rRNA (9). Our results support and explain previous in vivo and in vitro observations on inhibition of the protein synthesis by mutations in the CPK22,23. It has been shown that the CPK helps direct biogenesis, folding and function of the SSU. Time-resolved hydroxyl radical footprinting shows that the folding of the CPK occurs very early in subunit sythesis (10−4 to 10−2 s−1)47. Our scheme explains these results in the context of domain A, which includes the CPK. Defects in domain A impact subunit association and ultimately inhibit translation. Our results explain, on a molecular level, the effects of these mutations, which cause domain-wide changes in domain A folding as revealed by CD and SHAPE. Slight orientational alterations in helices 27-28 and 3–19 (which form intersecting orthogonal arcs) affect the overall structure, stability and dynamics of the SSU. Therefore, domain A is central player in protein synthesis machinery in all kingdoms of life.
Methods
Chemical reagents and synthetic oligonucleotides
The chemical reagents used here are molecular biology grade or higher. DNA primers and oligonucleotides were purchased from Operon MWG. All aqueous solutions were prepared with deionized, distilled, nuclease free water (HyClone, Thermo Scientific). For the experiments in the absence of divalent cations, nuclease free water was treated with the Chelex 100 Resin (Biorad) chelating resin and recovered with 0.2 μm Ultrafree–MC–GV Centrifugal Filters (Milipore). All the experiments are reproducible and repeated at least 2 times unless otherwise stated.
Construction of the transcription vector for domain AISO rRNA
The Thermus thermophilus HB8 strain SSU rRNA sequence was obtained from NCBI database. The domain AISO gene minus helix 28 was created by recursive PCR using the four oligonucleotides (5′ to 3′):
Forward 1: GGTGTGGGAATTCTAATACGACTCACTATAGGGTTGTTGGAGAGTTTGATCCTGGCT
Reverse 2: CAGTGAATCCGGGGCCTTACGGCCCCTGAGCCAGGATCAAACTCTCCAAC
Forward 3: GTAAGGCCCCGGATTCACTGGGCGCCGTAAGGCGCCTGGGGAGTACGGCC
Reverse 4: CACCAAGCTTATTCCTTTGAGTTTCAGCCTTGCGGCCGTACTCCCCAGGC
The flanking primers were;
Forward: TGAGTCGTATTAGAATTCCCACACC
Reverse: GAAACTCAAAGGAATAAGCTTGGTG.
The domain AISO gene was cloned into the pUC19 vector using the EcoRI and HindIII restriction sites. The transformation used 5 μL of the ligation mix, which was added to 50 μL DH5α cells using the heat-shock method. Plasmids obtained by minipreps were sequenced bidirectionally by Operon MWG.
Helix 28 was added with Q5 site-directed mutagenesis (NEB) using forward AAGCTTGGCGTAATCATGG and reverse TGTACAAGGGCCTTACGG primers. The C18A mutant was also made by Q5 site-directed mutagenesis, using forward AGAGTTTGATACTGGCTCAGG and reverse CCAACAACCCTATAGTGAG primers. For SHAPE experiments, a primer binding tail was added to the 3′ end by PCR using the reverse primer CACCAAGCTTGAACCGGACCGAAGCCCGATTTGTGTACAAGGGCCTTACGGCCCCCCGTCAATTCCTTTGAGTTTCAGCCTTGC (5′ to 3′). The secondary structure of domain AISO with the SHAPE tail is shown in the Supplementary Fig. 7.
Transcription and purification of the domain AISO rRNA
The pUC19 plasmid containing the domain AISO gene was digested with HindIII-HF (NEB) for 2 hours at 37 °C as described by manufacturer. The reaction mixture was incubated at 80 °C for 20 minutes to deactivate the enzyme. The reaction was purified with SmartSpin nucleic acid & purification columns (Denville Scientific Inc.) using DNA Clean & Concentrator Kit buffers (Zymo Research Corp.) Digested plasmid (400–1,000 ng) was used as a template for T7 RNA polymerase (NEB) transcription. Run-off transcription reaction was prepared according to manufacturer’s description (NEB T7 High Yield RNA Synthesis Kit). The reaction mixture was incubated for 16 hours at 37 °C. After incubation, 1 μL Turbo DNAse (Ambion) was added to the reaction mixture, which was then incubated for 30 minutes at 37 °C. RNA was purified by ammonium acetate precipitation. Ultimately, 40 μL nuclease-free H2O was added to the dried pellet and the OD was measured with a Nanodrop (Thermo Scientific). RNA was further purified by G25 size exclusion chromatography (illustraTMNAPTM-10, GE Healthcare).
SHAPE reactions and di-deoxysequencing
Dideoxy sequencing reactions were carried out by heating a 20 μL solution of 50 ng/μL domain AISO rRNA mixed with 10 μL (0.8 μM) 5′ 6-FAM labeled GAACCGGACCGAAGCCCG primer (Operon MWG). To anneal the primer to the RNA, the reaction was heated to 85 °C and slowly cooled to 30 °C at a rate of 1.5 °C per minute. For domain AISO rRNA lacking helix 28, 5′ 6-FAM labeled TATTCCTTTGAGTTTCAGCC primer was used. After primer annealing, 20 μL mixture of SuperScript® III Reverse Transcriptase (Invitrogen) reaction mixture prepared and added to domain AISO rRNA and primer mixture to give the final concentrations of 1X RT buffer, 2 mM DTT, 0.625 mM dNTPs, and 2.5 mM ddNTPs (TriLink BioTechnologies). The reverse transcription reaction was carried out by incubating 50 μL reaction mixture at 55 °C for 2 hours and quenched for 15 min by heating to 70 °C.
For the SHAPE reactions, a 70 μL solution of 150 ng/μL domain AISO rRNA was incubated for 4 min at 85 °C in the presence of 5 μM 1,2-diaminocyclohexanetetraacetic acid (Sigma) chelating agent and allowed to cool for 10 min into room temperature. This procedure depletes divalent cations from RNA. Divalent-free RNA was divided to two 32 μL samples and 4 μL of 10X folding buffer was added (500 mM HEPES pH 8.0, 2.5 M NaCl) for RNA folding with sodium. Four μL 10X folding buffer was added (500 mM HEPES pH 8.0, 2.5 M NaCl, 20 mM MgCl2) for RNA folding with sodium and magnesium. The sample was folded by incubated at 20 min at 37 °C and was divided two 18 μL solutions. One of the solutions was added to 2 μL of 800 mM benzoyl cyanide in anhydrous DMSO. The other solution was added 2 μL of pure DMSO for a negative background control. The reaction mixture was incubated 2 min at room temperature. The modified RNA was purified using Zymo RNA Clean and Concentrator Kit and eluted in 25 μL modified TE buffer (10 mM Tris, 0.1 mM EDTA). Primer annealing and extension reactions were as described above.
For the capillary electrophoresis, 1.5 μL of reverse transcription reaction mixture was mixed with 0.5 μL ROX-labeled DNA sizing ladder and 9 μL of HiDi Formamide (Applied Biosystems) in a 96-well plate. To denature the cDNA, the plate was incubated for 5 min at 95 °C. The mixture was resolved on a 3130 Genetic Analyzer (Applied Biosystems). Capillary electrophoresis data were processed using in-house MatLab scripts as described28. First, data were aligned via standard peaks and the baseline was subtracted. Sequencing peaks were matched with SHAPE data peaks. The traces were integrated and processed with a signal decay correction, and were scaled and normalized.
Circular dichroism spectroscopy
A solution of 25 ng/mL RNA, 5 mM sodium cacodylate, pH 6.8 was titrated with either a EDTA or Mg2+. The RNA was titrated first with the chelator, followed by back-titration with Mg2+, taking CD scans on a Jasco J-810 spectropolarimeter after each addition. Four CD spectra collected and averaged, from 350 to 220 nm with an integration time of 4 seconds, bandwidth of 4 nm, a scan speed of 50 nm/min. The temperature was kept at 20 °C. RNA concentrations were kept constant for mutant and intact RNAs. Since titration of 0.5–1 mM Mg2+ gives a plateau for all of the RNAs in the CD scans, titration was stopped after 1 mM divalent cations added. All RNAs are present in 74 μM of nucleotides, which corresponds to strand concentrations of 0.55 μM domain AISO, 0.55 μM domain AISO C18A mutant and 0.70 μM domain AISO w/o helix 28. CD experiments were repeated 3 times and results are reproducible.
3D Modeling and Minimization
The domain A three-dimensional structure was modeled the ribosome structure of Ramakrishnan (PDB ID:4V51)6. Nucleotides 1–29, 554–569, 881–929, and 1388–1396 were extracted from the crystal structure and capped by a stem-loop containing the three base pairs and a tetra loop with a sequence GCCGUAAGGC. The 3D coordinates of the stem loop were obtained from Hsiao et al.29. The stem loops were positioned as extensions of the domain A helices and connected to it by adding O3′-P bonds. The stem loops along with their two adjacent base-pairs from the domain A were subjected by the partial energy minimization, while the rest of the structure was held fixed.
Energy Minimization
Partial minimization of the re-ligated rRNAs was performed with Sybyl-X 1.2 software (Tripos International, St. Louis, MO, USA) with the AMBER FF99 force field using an implicit solvent model with the distance dependent dielectric function D(r) = 20r. The non-bonded cut-off distance was set to 12 Å. Each system was minimized by 1,000 steps of steepest decent followed by 5,000 steps of conjugate gradient minimization.
Superimposition
PDB IDs: 4V51, 4V9D, 4V88, 4V6W, 4V6X for Thermus thermophilus, Escherichia coli, Saccharomyces cerevisiae, Drosophila melanogaster, Homo sapiens were obtained from Protein Data Bank6,7,37,38.Structures were superimposed pairwise using PyMol “super” command with default settings.
Data Mapping
SHAPE data are normalized and mapped on in-house RiboVision48 server using the custom data function.
Shannon Entropy
Shannon Entropies were calculated as previously described49.
Figures and Images
Figures of three-dimensional structures are prepared with PyMol or Maxon Cinema 4D with the ePMV plugin50. Secondary structures are obtained from in-house RiboVision server48. Labels are added in Adobe Illustrator or Adobe Photoshop.
Additional Information
How to cite this article: Gulen, B. et al. Ribosomal small subunit domains radiate from a central core. Sci. Rep. 6, 20885; doi: 10.1038/srep20885 (2016).
References
Noller, H. F., Hoffarth, V. & Zimniak, L. Unusual resistance of peptidyl transferase to protein extraction procedures. Science 256, 1416–1419 (1992).
Ban, N., Nissen, P., Hansen, J., Moore, P. B. & Steitz, T. A. The complete atomic structure of the large ribosomal subunit at 2.4 Å resolution. Science 905, 10.1126/science.289.5481.905 (2000).
Ramakrishnan, V. & White, S. W. Ribosomal protein structures: insights into the architecture, machinery and evolution of the ribosome. Trends in Biochemical Sciences 23, 208–212 (1998).
Cate, J. H., Yusupov, M. M., Yusupova, G. Z., Earnest, T. N. & Noller, H. F. X-ray crystal structures of 70S ribosome functional complexes. Science 285, 2095–2104, 10.1126/science.285.5436.2095 (1999).
Harms, J. et al. High resolution structure of the large ribosomal subunit from a mesophilic eubacterium. Cell 107, 679–688 (2001).
Selmer, M. et al. Structure of the 70S ribosome complexed with mRNA and tRNA. Science (New York, N.Y.) 313, 1935–1942, 10.1126/science.1131127 (2006).
Anger, A. M. et al. Structures of the human and Drosophila 80S ribosome. Nature 497, 80–85, 10.1038/nature12104 (2013).
Frank, J. et al. A model of protein synthesis based on cryo-electron microscopy of the E. coli ribosome. Nature 376, 441–444, 10.1038/376441a0 (1995).
Petrov, A. S. et al. History of the ribosome and the origin of translation. Proc. Natl. Acad. Sci. USA, 112, 15396–15401 10.1073/pnas.1509761112 (2015).
Lescoute, A., Leontis, N. B., Massire, C. & Westhof, E. Recurrent structural RNA motifs, Isostericity Matrices and sequence alignments. Nucleic Acids Res. 33, 2395–2409 (2005).
Porter, L. L. & Rose, G. D. A thermodynamic definition of protein domains. Proc. Natl. Acad. Sci. USA 109, 9420–9425, 10.1073/pnas.1202604109 (2012).
Richardson, J. S. The anatomy and taxonomy of protein structure. Adv. Protein Chem. 34, 167–339 (1981).
Orengo, C. A. et al. CATH–a hierarchic classification of protein domain structures. Structure 5, 1093–1109 (1997).
Gutell, R. R., Larsen, N. & Woese, C. R. Lessons from an evolving rRNA: 16S and 23S rRNA structures from a comparative perspective. Microbiol Rev 58, 10–26 (1994).
Woese, C. R. et al. Secondary structure model for bacterial 16S ribosomal RNA: phylogenetic, enzymatic and chemical evidence. Nucleic Acids Res 8, 2275–2293 (1980).
Wimberly, B. T. et al. Structure of the 30S ribosomal subunit. Nature 407, 327–339, 10.1038/35030006 (2000).
Petrov, A. S. et al. Secondary Structures of rRNAs from all three Domains of Life. PLoS One 9, e88222, 10.1371/journal.pone.0088222 (2014).
Valle, M. et al. Locking and Unlocking of Ribosomal Motions. Cell 114, 123–134, http://dx.doi.org/10.1016/S0092-8674(03)00476-8 (2003).
Ogle, J. M. et al. Recognition of Cognate Transfer RNA by the 30S Ribosomal Subunit. Science 292, 897–902, 10.1126/science.1060612 (2001).
Pleij, C. W. A., Rietveld, K. & Bosch, L. A new principle of RNA folding based on pseudoknotting. Nucleic Acids Research V, 1717–1731 (1985).
Poot, R. a., Pleij, C. W. & van Duin, J. The central pseudoknot in 16S ribosomal RNA is needed for ribosome stability but is not essential for 30S initiation complex formation. Nucleic Acids Research 24, 3670–3676 (1996).
Poot, R. A., van den Worm, S. H. E., Pleij, C. W. A. & van Duin, J. Base complementarity in helix 2 of the central pseudoknot in 16S rRNA is essential for ribosome functioning. Nucleic Acids Research 26, 549–553, 10.1093/Nar/26.2.549 (1998).
Brink, M. F., Verbeet, M. P. & deBoer, H. A. Formation of the central pseudoknot in 16S rRNA is essential for initiation of translation. The EMBO journal 12, 3987–3996 (1993).
McGinnis, J. L. & Weeks, K. M. Ribosome RNA assembly intermediates visualized in living cells. Biochemistry 53, 3237–3247, 10.1021/bi500198b (2014).
Wilkinson, K. a., Merino, E. J. & Weeks, K. M. Selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE): quantitative RNA structure analysis at single nucleotide resolution. Nature Protocols 1, 1610–1616, 10.1038/nprot.2006.249 (2006).
Mortimer, S. a. & Weeks, K. M. A fast-acting reagent for accurate analysis of RNA secondary and tertiary structure by SHAPE chemistry. Journal of the American Chemical Society 129, 4144–4145, 10.1021/ja0704028 (2007).
Athavale, S. S. et al. Domain III of the T. thermophilus 23S rRNA folds independently to a near-native state. RNA (New York, N.Y.) 18, 752–758, 10.1261/rna.030692.111 (2012).
Athavale, S. S. et al. RNA folding and catalysis mediated by iron (II). PloS One 7, e38024–e38024, 10.1371/journal.pone.0038024 (2012).
Hsiao, C. et al. Molecular paleontology: a biochemical model of the ancestral ribosome. Nucleic Acids Research, 1–13, 10.1093/nar/gkt023 (2013).
Deigan, K. E., Li, T. W., Mathews, D. H. & Weeks, K. M. Accurate SHAPE-directed RNA structure determination. Proc Natl Acad Sci USA 106, 97–102, 10.1073/pnas.0806929106 (2009).
Woodson, S. A. Compact intermediates in RNA folding. Annual Review of Biophysics 39, 61–77 (2010).
Bowman, J. C., Lenz, T. K., Hud, N. V. & Williams, L. D. Cations in charge: magnesium ions in RNA folding and catalysis. Curr Opin Struct Biol 22, 262–272, 10.1016/j.sbi.2012.04.006 (2012).
Romer, R. & Hach, R. tRNA conformation and magnesium binding. A study of a yeast phenylalanine-specific tRNA by a fluorescent indicator and differential melting curves. Eur J Biochem 55, 271–284 (1975).
Frederiksen, J. K., Li, N.-S., Das, R., Herschlag, D. & Piccirilli, J. A. Metal-ion rescue revisited: Biochemical detection of site-bound metal ions important for RNA folding. RNA 18, 1123–1141 (2012).
Merino, E. J., Wilkinson, K. A., Coughlan, J. L. & Weeks, K. M. RNA Structure Analysis at Single Nucleotide Resolution by Selective 2′-Hydroxyl Acylation and Primer Extension (SHAPE). J. Am. Chem. Soc. 127, 4223–4231 (2005).
Mortimer, S. A. & Weeks, K. M. Time-Resolved RNA SHAPE Chemistry. J. Am. Chem. Soc. 130, 16178–16180 (2008).
Dunkle, J. A. et al. Structures of the bacterial ribosome in classical and hybrid states of tRNA binding. Science 332, 981–984, 10.1126/science.1202692 (2011).
Ben-Shem, A. et al. The structure of the eukaryotic ribosome at 3.0 A resolution. Science 334, 1524–1529, 10.1126/science.1212642 (2011).
Giegé, R., Frugier, M. & Rudinger, J. tRNA mimics. Current Opinion in Structural Biology 8, 286–293, http://dx.doi.org/10.1016/S0959-440X(98)80060-2 (1998).
Chen, X., Sim, S., Wurtmann, E. J., Feke, A. & Wolin, S. L. Bacterial noncoding Y RNAs are widespread and mimic tRNAs. RNA 20, 1715–1724, 10.1261/rna.047241.114 (2014).
Felden, B., Florentz, C., Giegé, R. & Westhof, E. A central pseudoknotted three-way junction imposes tRNA-like mimicry and the orientation of three 5′upstream pseudoknots in the 3′terminus of tobacco mosaic virus RNA. RNA 2, 201–212 (1996).
Fujiwara, T., Ito, K. & Nakamura, Y. Functional mapping of ribosome-contact sites in the ribosome recycling factor: a structural view from a tRNA mimic. RNA 7, 64–70 (2001).
Hirokawa, G. et al. Post-termination complex disassembly by ribosome recycling factor, a functional tRNA mimic. EMBO J 21, 2272–2281, 10.1093/emboj/21.9.2272 (2002).
Carter, A. P. et al. Functional insights from the structure of the 30S ribosomal subunit and its interactions with antibiotics. Nature 407, 340–348 (2000).
Gao, N. & Frank, J. A library of RNA bridges. Nature Chemical Biology 2, 231–232 (2006).
Schuwirth, B. S. et al. Structures of the Bacterial Ribosome at 3.5 Å Resolution. Science 310, 827–834, 10.1126/science.1117230 (2005).
Sykes, M. T. & Williamson, J. R. A complex assembly landscape for the 30S ribosomal subunit. Annual Review of Biophysics 38, 197–215 (2009).
Bernier, C. R. et al. RiboVision suite for visualization and analysis of ribosomes. Faraday Discuss 169, 195–207, 10.1039/c3fd00126a (2014).
Petrov, A. S. et al. Secondary structure and domain architecture of the 23S and 5S rRNAs. Nucleic Acids Res 41, 7522–7535, 10.1093/nar/gkt513 (2013).
Johnson, G. T., Autin, L., Goodsell, D. S., Sanner, M. F. & Olson, A. J. ePMV Embeds Molecular Modeling into Professional Animation Software Environments. Structure 19, 293–303, 10.1016/J.Str.2010.12.023 (2011).
Acknowledgements
This work was supported in part by the NASA Astrobiology Institute (NNA09DA78A). The authors thank Dr. Kevin Weeks for providing processed SHAPE data from the intact SSU rRNA and Dr. Roger Wartell for valuable discussions and comments on the manuscript and Mrs. Susann Orth for valuable assistance with figure preparation.
Author information
Authors and Affiliations
Contributions
B.G., N.V.H. and L.D.W. conceived of the approach and designed the experiments. B.G. performed the experiments and processed the data except that C.D.O. and D.V.W. conducted the CD experiments. E.B.O. designed the constructs, produced and purified the RNA. A.S.P. designed, constructed and minimization of the three-dimensional model of domain AISO. B.G. prepared the figures. B.G. and L.D.W. wrote the manuscript. L.D.W. provided overall supervision of the project. All authors contributed to writing manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Gulen, B., Petrov, A., Okafor, C. et al. Ribosomal small subunit domains radiate from a central core. Sci Rep 6, 20885 (2016). https://doi.org/10.1038/srep20885
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep20885
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.