Abstract
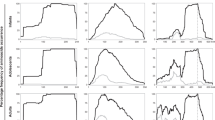
Current serological diagnostic assays for typhoid fever are based on detecting antibodies against Salmonella LPS or flagellum, resulting in a high false-positive rate. Here we used a protein microarray containing 2,724 Salmonella enterica serovar Typhi antigens (>63% of proteome) and identified antibodies against 16 IgG antigens and 77 IgM antigens that were differentially reactive among acute typhoid patients and healthy controls. The IgG target antigens produced a sensitivity of 97% and specificity of 80%, whereas the IgM target antigens produced 97% and 91% sensitivity and specificity, respectively. Our analyses indicated certain features such as membrane association, secretion and protein expression were significant enriching features of the reactive antigens. About 72% of the serodiagnostic antigens were within the top 25% of the ranked antigen list using a Naïve bayes classifier. These data provide an important resource for improved diagnostics, therapeutics and vaccine development against an important human pathogen.
Similar content being viewed by others
Introduction
Typhoid fever is a life threatening bacterial infection caused by typhoidal members of the genus Salmonella, including Salmonella enterica serovar Typhi and Salmonella enterica Paratyphi A. The disease is common in South and Southeast Asia, parts of Africa and other developing regions that have poor sanitation and limited access to clean water1. S. Typhi is the dominant agent globally, estimated to cause over 21 million cases of typhoid and 200,000 deaths. S. Paratyphi A is less common than S. Typhi, responsible for approximately 5 million illnesses in the year 20002,3, yet is becoming increasingly prevalent across Asia2,4,5,6.
Typhoidal organisms are stealth pathogens, with the ability to regulate the host immune response during an infection3. Yet, the majority of patients do have detectable antibody responses to the Vi capsular polysaccharide, the O (lipopolysaccharide) and the H (flagellar) antigens during active infection and for prolonged periods afterwards7,8. Seroprevalence is highest in endemic countries, although individual titers vary according to geographic location, age and exposure rates9. This is highlighted by individuals in endemic areas frequently having elevated Vi antibody and O antibody titers with no clinical history of typhoid and by a frequent inability to detect elevated antibody responses in culture confirmed patients4,5,6,7. The Widal test, first introduced in 1896, detects agglutinating antibodies against the O and H antigens of S. Typhi10. These antigens are cross-reactive with antibodies from other Salmonella serovars and related Gram-negative bacteria, resulting in a high false-positive rate. Despite this, the Widal test is still widely used for typhoid diagnosis in developing countries. Commercial serological tests, such as Tubex and Typhidot11,12, which typically titrate and differentiate IgM and IgG against the O and H antigens, are fraught with the same limitations of the Widal test, having sensitivities and specificities of around 70% and 80%, respectively13,14.
Current knowledge of the antigen repertoire recognized by patients during an acute typhoid infection is sparse15,16, limiting a detailed interrogation of immunity, exposure and hindering preclinical vaccine development. Protein microarrays have been used previously to probe the natural immune response to a multitude of infections, adding insight in the pathogenicity and natural history of bacterial, viral and parasitic infections17,18,19,20,21,22,23,24,25. We aimed to detail a comprehensive overview of antigenic targeting using sera from patients infected with S. Typhi, with acute, active typhoid fever. Here we describe the identification of Salmonella-specific IgG and IgM target antigens following natural human infection after probing protein microarrays representing >2,700 S. Typhi antigens. Our unique study provides an antibody signature during acute S. Typhi infections and identifies novel antigens suitable for both vaccine development and diagnostics.
Results
Gene amplification, cloning and protein expression
A set of 2,724 ORFs from S. Typhi TY2, representing approximately 63% of the S. Typhi genome, were selected for screening26. These ORFs were prioritized using markers we found to be enriched in seroreactive antigens in other bacterial species22. These features included: signal peptides motifs, proteins with characteristics of the ‘outer membrane’, the ‘periplasm’, ‘heat shock’, ‘chaperone’, ‘transport protein’, ‘lipoprotein’, or ’virulence’. To minimize the potential for background cross-reactivity, proteins that were non-homologous (<50% identify at the amino acid level) to E. coli were also selected. S. Typhi TY2 ORFs cloned in pXT7 vector were expressed under T7 promoter in the E. coli in vitro transcription/translation system, printed on microarrays. By detecting HA- or HIS-tag expression, we were able to confirm expression of 96% of the proteins ( Supplemental Figure 1 ).
Human IgG profile and identification of serodiagnostic IgG antigens
The constructed protein array was probed with sera from 34 acute typhoid patients. The IgG and IgM immunoproteome, defined as the total number of antigens reacting with serum from at least 1 acute typhoid patient, consisted of 2,442 (89%) and 809 (30%) antigens respectively ( Figure 1 ). A subset of the IgG proteome, consisting of 127 antigens was identified as serodominant (Materials and Methods, Data Analysis) ( Figure 2A, 2B, 2C and Table S1, S2 ), with 16 antigens found to be significantly serodiagnostic (the ability to distinguish between serum from acute typhoid patients and control serum) (Benjamini and Hochberg adjusted Cyber-T p value <0.05). As there is extensive DNA homology within the Salmonella genus, we compared seroreactivity of the typhoid patients against that of African patients with nontyphoidal Salmonella infections from our previously published work (Figure 2A26. We found 1 of the 16 serodiagnostic typhoid antigens (t1459), was also serodiagnostic for nontyphoidal Salmonella patients from Africa26 (Table S1). Six of the 16 serodiagnostic antigens were also able to distinguish between the Vietnamese typhoid patients and the nontyphoidal Salmonella patients from Africa. The remaining 10 antigens reacted similarly between both groups (Figure S2).
Composition of the IgG and IgM Immunoproteomes.
(A) IgG immunoproteome consists of 89% of the antigens cloned. There are 2442 antigens with detected IgG response in at least 1 acute sample (3% of all acute samples), 1608 antigens reactive in at least 9% of all acute samples, 38 antigens reactive in at least 50% of all acute samples and 1 antigen reactive in 94% acute samples. (B) IgM immunoproteome consists of 30% of the antigens cloned. There are 809 antigens reactive in at least 1 (3%) acute sample, 410 antigens in at least 9% of all acute samples, 36 antigens in at least 50% of acute samples and 118 antigens are reactive in 10–34 samples.
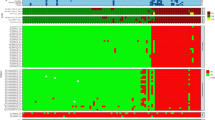
Human IgG profile.
Arrays containing 2724 S.enterica proteins were probed with sera from human acute typhoid patients and control subjects in Vietnam and USA. (A) Heatmap showing signal intensity with red strongest, bright green weakest and black in between. Only the serodiagnostic antigens (Benjamini Hochberg corrected p value <0.05) are shown. The human samples are in columns and sorted left to right by increasing average intensity to serodiagnostic antigens. One antigen marked in red, t1459, is also found serodiagnostic for nontyphoidal salmonellosis in Africa. (B) Reactivity of cross-reactive (Benjamini Hochberg corrected p value >0.05) antigens are shown in heatmap. (C) The mean IgG reactivity of the antigens was compared between the acute typhoid patients and endemic controls. Antigens with Benjamini Hochberg corrected p-value less than 0.05 are organized to the left and cross-reactive antigens to the right. The 16 serodiagnostic and 22 of the most reactive cross-reactive antigens are shown. (D) The LOOCV ROC graphs show classifiers with increasing number of human serodiagnostic IgG antigens.
In addition to LPS from Salmonella enterica and E. coli, we identified a further 111 antigens (including the H antigen; ORF t0918) that reacted comparably among all serum samples, irrespective of whether from typhoid cases or from non-infected controls ( Figure 2 , Table S2 ). These cross-reactive antigens may be indicative of past exposure to Salmonella and use of these antigens should be selectively avoided as serodiagnostic markers for acute typhoid in the future. Six of these cross-reactive antigens shared homology with S. Typhimurium antigens and were differentially reactive between the nontyphoidal Salmonella patients and controls in Africa. Furthermore, when we compared seroreactivity between differing human populations, we observed significantly higher overall antibody response in control group in Vietnam than in serum from a non-endemic area in USA, indicative of an elevated typhoid exposure in Vietnam compared to the USA.
To assess the ability of the serodiagnostic antigens to accurately distinguish between controls and typhoid cases, we generated “leave one out cross-validation (LOOCV) receiver operating characteristic (ROC) curves” ( Figure 2D ). The serodiagnostic antigens were ordered by decreasing BH corrected p value. We used kernel methods and support vector machines to build linear and nonlinear classifiers. As input to the classifier, we used the highest-ranking 1, 3, 5, 10 and 16 antigens. The results show that increasing the antigen number from 1 to 10 produced significant improvement in sensitivity and specificity ( Figure 2D ). This was optimum for 10 antigens, with this classifier yielding a sensitivity of 98% and a specificity of 80%.
Human IgM profile
Measuring the IgM response is the most suitable approach for distinguishing between the early stages of an acute infection and healthy controls. A summary of the IgM responses is shown in Figure 3A . We could identify 77 antigens that discriminated between the control population and those with acute typhoid (BH-corrected p value <0.05; Figure 3B , Table S3 ). The overall reactivity against these 77 antigens in the control group was low, with an elevated response against a limited number of antigens detected in only a small number of serum samples, predicting a greater degree of serodiagnostic sensitivity and specificity than the IgG antigen response. Additionally, the reactivity in typhoid patients was significantly greater in comparison to the control group, indicating a strong IgM response during an acute typhoid infection. Twenty-two antigens that reacted similarly among all human samples were also identified, signifying non-specific antigenic cross-reactivity ( Figure 3 , Table S4 ).
Human IgM profile.
S.enterica arrays were probed for IgM response with sera from human acute typhoid patients and control subjects in Vietnam. (A) Antigens are listed on the vertical axis and the individual donors along the top. Only the serodominant IgM antigens are shown. These antigens were sub-classified by Benjamini Hochberg corrected cyber T test p value into discriminatory (p<0.05; n = 77) and non- discriminatory (p>0.05; n = 22) by comparing the endemic control group with human typhoid patients in Vietnam. The antigens are ranked by the average response of the typhoid patients group and donors sorted from left to right by increasing average signal. (B) The mean IgM reactivity of the antigens was compared between the typhoid patients and endemic controls. Antigens with Benjamini Hochberg corrected p-value less than 0.05 are organized to the left and cross-reactive antigens to the right. (C) The LOOCV ROC graphs show classifiers with increasing number of human serodiagnostic IgM antigens. Overall, all 77 antigens produced sensitivity and specificity of 90% and 91%, respectively.
Cross-validation receiver operating characteristic (ROC) curves were generated to assess the accuracy of IgM response to the 77 antigens to distinguish between acute typhoid patients and controls ( Figure 3 ). The candidate serodiagnostic antigens to IgM were ranked by decreasing BH corrected p values, showing an elevated sensitivity and specificity from 1 to 30 antigens. This classifier yielded sensitivity and specificity rates of 97% and 91%, respectively for the top 30 IgM antigens, yet combining all 77 antigens yielded a drop in sensitivity to 90%, with specificity remaining at 91%.
Class switching from IgM to IgG
Immunoglobulin class-switching from IgM to other isotypes is an indicative component of immune response maturation. The extent of overlap of the IgG and IgM profiles from uninfected control and typhoid patients is illustrated in Figure 4. There were a total of 199 IgM and IgG seroreactive antigens, of which 30 were consistently detected between IgM and IgG, indicative of class switching. Furthermore, 69 antigens were observed in the IgM profile but not the IgG profile and an additional 100 antigens were observed in the IgG profile but not the IgM profile.
Overlap between IgM and IgG profiles.
Scatter plots of corresponding mean IgG and IgM responses against the 199 combined IgM and IgG target antigens that discriminate between negative control and typhoid individuals. The horizontal and vertical hashed lines are seropositive cutoffs for IgM and IgG, respectively, defined as the mean+3SD of the arrays ‘no-DNA’ control spots.
Validation of serodiagnostic accuracy with immunostrips
To investigate the diagnostic potential of the serodiagnostic antigens, six serodiagnostic proteins were extracted from inclusion bodies of E. coli BL21 in vitro ( Figure 5A ) and were printed onto Nitrocellulose membranes, representing a simple analytical assay. The immunostrips were probed with serum from 16 acute typhoid patients and 14 controls. Serum from the 16 typhoid patients demonstrated greater reactivity than serum from the controls (Figure 5B). To assess the ability of these six antigens to distinguish between typhoid and controls, a LOOCV ROC curve was generated ( Figure 5C ) and demonstrated that the six selected antigens yielded a sensitivity of 94% and a specificity of 100%.
Immunostrips.
(A) Six serodiagnostic antigens were expressed in E.coli and extracted from the inclusion body. Proteins were loaded in 4–12% SDS PAGE, stained with blue stain or transferred to nitrocellulose membrane and probed for HIS tag. All proteins were confirmed positive with HIS tag. (B) 0.1 mg/mL of each protein was printed onto nitrocellulose paper in adjacent stripes using a BioDot jet dispenser. Membrane was cut into 3 mm strips. Strips were probed with human acute typhoid patient or endemic naive sera. Weak reactivity in the naïve healthy controls can be distinguished from the strong reactivity in infected group. (C) The LOOCV ROC curve was generated and sensitivity and specificity of all 6 antigens on immunostrips probing test is 94% and 100%, respectively.
Enrichment analysis
To understand the nature of the immune response to S. Typhi we performed a detailed analysis of the serodominant and serodiagnostic IgG antigens. Proteins were annotated using the NCBI Clustered Orthologous Group (COG) functional categories and in silico predictions were made for transmembrane domains, signal peptides, subcellular localizations and isoelectric points (pI) and compared to published mass-spectrometry data27.
Proteins with predicted COG-M function, involved in cell envelope biogenesis and outer membrane, were 1.8- fold enriched in serodominant antigens (p = 3.42E-02) and proteins with predicted COG-N function, involved in cell motility and secretion, were 2.0-fold enriched in serodominant antigens. Proteins with predicted COG-O function, posttranslational modification, protein turnover, chaperones, were also significantly enriched at 2.6- and 5.5- fold in the serodominant (p = 2.15E-03) and serodiagnostic (p = 1.53E-02) antigens, respectively. Conversely, proteins with predicted COG-G (carbohydrate transport and metabolism) or COG-K (transcription) functions were significantly underrepresented in serodominant antigens ( Table S5–S7 ).
Proteins with one transmembrane domain were also significantly enriched in serodominant and serodiagnostic antigen groups, with proteins lacking transmembrane domains or more than one transmembrane domain underrepresented. Proteins with predicted signal peptides (SignalP score > 0.7) were significantly enriched at 2.0- fold and proteins without signal peptides were significantly underrepresented. PSortb predicted 19 serodominant and two serodiagnostic periplasmic antigens, resulting in 3.4- and 2.9- fold enrichment, respectively (p = 1.28E-06 and 1.53E-01, respectively). However, pSortb cytoplasmic and cytoplasmic membrane proteins were 0.5- and 0.4- fold underrepresented in serodominant antigen group. We also found that proteins with pI 9–14 were 0.7-fold underrepresented in serodominant antigen group.
As supported by our previous study19, we found that the level of protein expression is an important factor contributor to antigenicity. A recent mass spectrometry study of S. Typhi TY2 proteins identified 2,062 expressed proteins, of which, 923 proteins were represented in this study27. Our results indicate that expression was a significant feature of enrichment, with 1.6- fold increase in serodominant antigens (p = 8.11E-06). Furthermore, as the number of detected peptides increased, the fold enrichment also increased. There was 1.9- fold enrichment of serodiagnostic antigens in proteins detected with at least 10 peptides (p = 4.30E-02), 3.3- fold in proteins detected with at least 50 peptides (p = 2.81E-02). However, proteins that were not identified by MS were underrepresented at 0.7- fold among both the serodominant and the serodiagnostic antigens.
Ranking antigenicity of antigens using Naïve Bayes classifier
To investigate the relationship between proteomic features and the seroreactivity, we used a naïve Bayes classification scheme to rank the probability of antigenicity of all S. Typhi proteins on chip28,29. The likelihood ratio was calculated using all the proteomic features for all represented proteins on chip, proteins were ranked according to this calculation for IgG and IgM combined serodominant or serodiagnostic antigens.
Our analysis suggested that we would need to screen only 5% of the genome to identify 9% of all seroreactive IgG and IgM antigens and 25% of all serodiagnostic antigens ( Figure 6 , Table 1 and Table 2 ), resulting in 1.84- and 5.01-fold enrichment, respectively. Therefore, as the number of proteins increases, more seroreactive and serodiagnostic antigens are identified. Precisely 72% of the serodiagnostic antigens are within the top 25% of the ranked antigen list, yet 50% of the proteome would require to be tested to identify 72% of seroreactive hits. These data suggest that the antibody responses against the cross-reactive antigens are derived from previous exposure to unrelated infections and not associated with the active infection in the infected group.
Combined Naïve bayes classifier on ranking of seroreactive and serodiagnostic antigens.
The cloned genome (2724 genes) was ranked by naïve bayes classifier based on 196 IgG/IgM seroreactive and 92 IgG/IgM serodiagnostic antigens features. As the percentage of genome increases, prediction rate also increases.
Discussion
Here we report the first proteome microarray analysis of S. Typhi proteins that are antigenic in the context of naturally acquired human typhoid infections. These data have profound implications for understanding bacterial pathogenesis and the practical development of diagnostics, therapeutics and vaccines for a disease that still kills over 200,000 people every year.
Current serological diagnosis of human typhoid suffers from the inability to distinguish between an active infection and historical exposure. These traditional serological assays are based primarily on identification of antibody to the O antigen, the H antigen or the Vi capsule and have not changed substantially for over a century. However, LPS is notoriously cross-reactive between bacterial species30 and exposure to the S. Typhi O, H and Vi antigen may be highly prevalent in endemic areas7,8,31. High background reactivity was confirmed by our microarray study, where O and H antigens reacted similarly in endemic controls and acute typhoid patients, explaining the limited utility of these antigens for accurate diagnosis. In comparison, the top 10 serodiagnostic IgM antigens discriminated between typhoid cases and healthy controls in Vietnam with a sensitivity and specificity of 94% and 91% respectively and the top 10 IgG antigens demonstrated 98% sensitivity and 80% specificity with immunostrips. Thus, our findings should allow the development of a new generation of diagnostic tests for this important bacterial infection.
Designing a field-deployable diagnostic for typhoid requires an understanding of the background response to Salmonella in endemic areas. During our analyses we noted a significant difference in background reactivity to S. Typhi proteins in uninfected individuals from endemic against uninfected individuals from non-endemic areas. In Vietnam, control subjects had much higher background reactivity to Salmonella antigens, compared to control subjects from the USA ( Figure 2 ). The higher background in endemic countries, like Vietnam, makes it more challenging to identify IgG antigens recognized specifically by active infection, whereas this is less challenging in non-endemic areas that have lower background. Consideration of these differences is important for the development of diagnostic assays intended for use in endemic and non-endemic regions. Moreover, we did not observe high background of IgM antibodies even in endemic countries (Vietnam), indicating that IgM may be a more suitable approach for diagnosing acute typhoid in endemic regions.
Two antigens that we found to discriminate between typhoid patients and control subjects in Vietnam, hemolysin E (hlyE, t1477) and putative toxin-like protein (cdtB, t1111) have been identified in Bangladeshi patients by an immune-affinity technique15. Furthermore, a nonspecific acid phosphatase precursor (phoN, t4225), also identified in Bangladeshi patients, was identified to be reactive in Vietnamese patients and controls. We further identified 14 novel serodiagnostic IgG antigens and 77 novel serodiagnostic IgM antigens that may be specific for human typhoid patients in Vietnam( Table S1 and Table S3 ). It is noteworthy, that despite high genome conservation within the genus Salmonella, only one antigen was also found to be serodiagnostic for nontyphoidal Salmonella infections in African individuals. Furthermore, six antigens could distinguish between typhoid patients in Vietnam and nontyphoidal Salmonella patients in Africa. Further studies on diagnostic antigens that are able to distinguish between typhoid, nontyphoidal Salmonella and other febrile diseases from the same geographic areas are of high practical importance.
We were interested in expanding the utility of work by understanding the mechanisms of antigenicity of S. Typhi proteins; we addressed this by identifying proteomic features that were enriched in the serodominant antigen set. Enrichment analysis identified nine proteomic features that were over represented in the serodominant antigens. The proteomic features that were overrepresented included; COGs M, N, O and W, the predicted features TMHMM = 1, SignalP>0.7, pSort periplasmic, pSort unknown and evidence of expression in viable organisms. No single proteomic feature or category of features was sufficient to identify all the signature antigens. However, our analyses may exhibit bias as consequence of the proteins selected to go on the array. Yet, we anticipate similar results from a full proteome chip and have previous catalogues of serodominant antigens for numerous bacteria and have consistently found these features predict antigenicity19,21,22,32.
To increase the accuracy of predicting serodominant and serodiagnostic antigens over the entire proteome, we used all proteomic features information in a naïve Bayes classification approach to rank the cloned proteome for increased likelihood of seroreactivity. By doing so we were able to segregate 19% of the serodominant proteins and 39% of the serodiagnostic proteins within 10% of the cloned proteome. Within the top 25% of ranking of the cloned genome, we were able to predict 45% of the serodominant antigens and 72% of the serodiagnostic antigens. We anticipate a higher prediction of serodominant and serodiagnostic antigens in using the full proteome. This approach is relevant and necessary for studies focusing on small portions of the proteome that are the most likely to contain serodiagnostic antigens.
In conclusion, our unique approach to studying typhoid infections provides an empirical basis for understanding the breadth and specificity of the human immune response to S. Typhi. The results presented here represent an analysis of 63% of predicted proteins in the S. Typhi genome. Furthermore, the subset of diagnostic antigens identified here provide an initial estimated accuracy rate of 90% for the diagnosis of typhoid fever and will open the door to long-needed rapid diagnostics for this important and under-studied human restricted infection.
Methods
Ethics statement
This study was conducted according to the principles expressed in the Declaration of Helsinki and the studies contributing samples were approved by the institutional ethical review boards of the Hospital for Tropical Diseases, the Dong Thap Hospital and the Health services of Dong Thap Province in Vietnam and The Oxford University Tropical Research Ethics Committee (OxTREC) in the UK. All enrollees were required to provide written informed consent for the collection of samples, in the case of children this was provided by the parent or guardian.
Human serum samples
For the purposes of this work, serum was collected from Vietnamese individuals with acute typhoid fever and from a control group without disease originating from previously published epidemiologic or clinical studies33,34,35,36. The typhoid studies were conducted at the Hospital for Tropical Diseases in Ho Chi Minh City or and Dong Thap Provincial Hospital in Dong Thap Province, southern Vietnam in 2002 and 2003. For all acute febrile patients with confirmed etiology, serum was typically separated from 2 ml of venous blood within three days of fever onset and hospital admission. Children were enrolled with the same criteria as the adults and none of the children were < 1 year old due to ethical considerations of taking required volumes of blood for serology. The control serum samples were collected from healthy adults working at the Oxford University Clinical Research Unit in Ho Chi Minh City described previously in Thomson, et al35 and from umbilical cord blood samples from babies born at Dong Thap Provincial Hospital between 2002 and 200334,35. We also tested a group of 27 control individuals from the US with no prior symptoms of typhoid fever.
Microarray fabrication and probing
Genes were amplified and cloned using a high-throughput PCR and recombination method19. ORFs from S. Typhi TY2 genomic DNA were identified using GenBank NC_004613 and amplified using gene specific primers containing 20 bp nucleotide extension complementary to ends of linearized pXT7 vector, which allows homologous recombination between the PCR product and pXT7 vector in competent DH5a cells. The resulting fusion proteins also harbored a hemagglutinin epitope at 3′ end and polyhistidine at the 5′ end. ORFs longer than 3 kb in length were split into multiple in-frame overlapping segments shorter than 3 kb for cloning. Plasmids were expressed at 24°C in a 16 hour- in vitro transcription/translation E. coli system (expressway kits from Invitrogen). For no DNA controls, no plasmid DNA was added to the same amount of reagent from in vitro transcription/translation E.coli system to test E.coli background reactivity. For microarrays, 10 μl of reaction was mixed with 3.3 μl 0.2% Tween 20 to give a final concentration of 0.05% Tween 20 and printed onto nitrocellulose coated glass FAST slides (Whatman) using an Omni Grid 100 microarray printer (Genomic Solutions). Protein expression and printing was monitored by immunoprobing with anti-polyhistidine (clone His-1, Sigma) and anti-hemagglutinin (clone 3F10, Roche). Salmonella enterica lipopolysaccharide and E.coli lipopolysaccharide were purchased from Sigma and printed at 0.1 mg/mL and 0.01 mg/mL (L6511, L2880, Sigma). Human sera samples were diluted to 1:200 with 10 mg/ml E. coli lysate (Mclab). Microarray slides were incubated in biotin-conjugated secondary antibody (Jackson ImmunoResearch) diluted 1/200 in blocking buffer and detected by incubation with streptavidin-conjugated SureLight® P-3 (Columbia Biosciences). The slides were washed and air-dried by brief centrifugation. Microarray slides were scanned and analyzed using a Perkin Elmer ScanArray Express HT microarray scanner. Intensities were quantified using QuantArray software. All signal intensities were corrected for spot-specific background. All signal intensities are corrected for spot-specific background.
Protein expression and Immunostrips probing
Six plasmids of interest were single colony purified and transformed to BL21 cells. Single colonies were picked and inoculated into 3 ml LB with 50 ug/ml Kanamycin and incubated overnight at 37°C at 300 rpm. The overnight cultures were transferred to TB until OD600 nm reached 0.6. Isopropyl β-D-1-thiogalactopyranoside (IPTG) was then added at 0.4 mM and the proteins were induced for another 3 hrs at 30°C at 200 rpm. Cells were harvested and lysed in 1% Bugbuster solution (Novagen). The remaining pellets were solubilized in 7 M urea and filtered through a 0.2 μm filter. The proteins were then quantified and printed at approximately 0.1 mg/ml on Optitran BA-S 85 0.45 μm Nitrocellulose membrane (Whatman) using BioJet dispenser (BioDot). The membrane was then cut into 3 mm strips. For immunostrips probing, human sera were diluted to 1/200 in 5% non-fat dry milk solution dissolved in 10 mM Tris (pH 8.0) and 150 mM NaCl containing 0.05% (v/v) Tween 20 (T-TBS) containing 10 mg/ml E. coli lysate (McLab). Each strip was then incubated with pretreated sera for 2 hrs at room temperature with gentle mixing. Strips were extensively washed and then incubated in alkaline phosphatase conjugated donkey anti-human immunoglobulin (anti-IgG, Fcγ fragment-specific, Jackson ImmunoResearch) secondary antibody, diluted to 1/2000 and after extensive wash, reactive bands were visualized by incubating with 1-Step™ Nitro-Blue Tetrazolium Chloride/5-Bromo-4-Chloro-3′-Indolyphosphate p-Toluidine Salt (NBT/BCIP) developing buffer (Thermo Fisher Scientific). Immunostrips were scanned with Hewlett-Packard document scanner and were quantified using Image J software (NIH).
Data analysis
All analysis was performed using the R statistical environment (http://www.r-project.org) and SAS (http://www.sas.com/) statistical software. The vsn method implemented as part of the Bioconductor suite (www.bioconductor.org) is applied to the quantified array intensities. In addition to removing heteroskedacity, this procedure corrects for non-specific noise effects by finding maximum likelihood shifting and scaling parameters for each array such that control probe variance is minimized37,38. Antigens were considered “serodominant” with mean reactivity among typhoid patients greater than 3 times of standard deviations above the mean of the no DNA controls. Protein antigens with multiple segments were considered “serodominant” if one or more segments of this protein were “serodominant”.
Diagnostic biomarkers of typhoid were determined by comparing typhoid and control groups using a Bayes regularized t-test adapted from Cyber-T for protein arrays39,40. To account for multiple testing conditions, the Benjamini and Hochberg (BH) method was used to control the false discovery rate41. After Benjamini and Hochberg correction, a p-value smaller than 0.05 was considered significant and the corresponding proteins were considered differentially recognized or “serodiagnostic”, whereas non-differentially recognized or “crossreactive” proteins had p-value larger than 0.05. Multiplex classifiers were constructed using linear and non-linear Support Vector Machines (SVMs) using the “e1071” R package. SVM is a supervised learning method that has been successfully applied to microarray data characterized by small samples sizes and a large number of attributes42,43. The SVM approach, as any other supervised classification approach, uses a training dataset to build a classification model and a testing set to validate the model. To generate unbiased training and testing sets, leave one out cross-validation (LOOCV) was used. With this methodology, each data point is tested with a classifier trained using all of the remaining data points. Plots of receiver operating characteristic (ROC) curves were made with the ‘ROCR” R package.
The following programs were utilized for computational prediction of structural features. TMHMM v2.0 was utilized for transmembrane domains prediction44 (http://www.cbs.dtu.dk/services/TMHMM/), SignalP v3.0 for signal peptide prediction45 (http://www.cbs.dtu.dk/services/SignalP/), PSORTb v3.0. software for cellular location prediction46 (http://www.psort.org/psortb/). PI/MW tool from Swiss Institute of Bioinformatics was used to determine isolectric point (http://ca.expasy.org/tools/pi_tool.html). P value for enrichment statistical analysis was calculated using Fisher's exact test in the R environment. A combined naïve Bayes classifier approach29 originally applied by us to classify antigens from the Francisella tularensis proteome28, was used here to rank all Salmonella proteins.
References
Baker, S. et al. Combined high-resolution genotyping and geospatial analysis reveals modes of endemic urban typhoid fever transmission. Open biology 1, 110008 (2011).
Woods, C. W. et al. Emergence of Salmonella enterica serotype Paratyphi A as a major cause of enteric fever in Kathmandu, Nepal. Trans R Soc Trop Med Hyg 100, 1063–1067 (2006).
Baker, S., Favorov, M. & Dougan, G. Searching for the elusive typhoid diagnostic. BMC Infect Dis 10, 45 (2010).
Connor, B. A. & Schwartz, E. Typhoid and paratyphoid fever in travellers. Lancet Infect Dis 5, 623–628 (2005).
Ochiai, R. L. et al. Salmonella paratyphi A rates, Asia. Emerg Infect Dis 11, 1764–1766 (2005).
Sood, S. et al. Paratyphoid fever in India: An emerging problem. Emerg Infect Dis 5, 483–484 (1999).
House, D. et al. Serology of typhoid fever in an area of endemicity and its relevance to diagnosis. J Clin Microbiol 39, 1002–1007 (2001).
House, D. et al. Use of paired serum samples for serodiagnosis of typhoid fever. J Clin Microbiol 43, 4889–4890 (2005).
Pulickal, A. S. et al. Kinetics of the natural, humoral immune response to Salmonella enterica serovar Typhi in Kathmandu, Nepal. Clin Vaccine Immunol 16, 1413–1419 (2009).
Parry, C. M. et al. Value of a single-tube widal test in diagnosis of typhoid fever in Vietnam. J Clin Microbiol 37, 2882–2886 (1999).
Lim, P. L., Tam, F. C., Cheong, Y. M. & Jegathesan, M. One-step 2-minute test to detect typhoid-specific antibodies based on particle separation in tubes. J Clin Microbiol 36, 2271–2278 (1998).
Prakash, P. et al. Dot enzyme immunoassay (Typhidot) in diagnosis of typhoid fever in children. J Trop Pediatr 53, 216–217 (2007).
Dutta, S. et al. Evaluation of new-generation serologic tests for the diagnosis of typhoid fever: data from a community-based surveillance in Calcutta, India. Diagn Microbiol Infect Dis 56, 359–365 (2006).
Ochiai, R. L. et al. A study of typhoid fever in five Asian countries: disease burden and implications for controls. Bull World Health Organ 86, 260–268 (2008).
Charles, R. C. et al. Characterization of anti-Salmonella enterica serotype Typhi antibody responses in bacteremic Bangladeshi patients by an immunoaffinity proteomics-based technology. Clin Vaccine Immunol 17, 1188–1195 (2010).
Srinivasan, A. & McSorley, S. J. Visualizing the immune response to pathogens. Curr Opin Immunol 16, 494–498 (2004).
Kunnath-Velayudhan, S. et al. Dynamic antibody responses to the Mycobacterium tuberculosis proteome. Proc Natl Acad Sci U S A 107, 14703–14708 (2010).
Crompton, P. D. et al. A prospective analysis of the Ab response to Plasmodium falciparum before and after a malaria season by protein microarray. Proc Natl Acad Sci U S A 107, 6958–6963 (2010).
Liang, L. et al. Systems Biology Approach Predicts Antibody Signature Associated with Brucella melitensis Infection in Humans. J Proteome Res 10, 4813–4824 (2011).
Liang, L. et al. Identification of potential serodiagnostic and subunit vaccine antigens by antibody profiling of toxoplasmosis cases in Turkey. Mol Cell Proteomics 10, M110.006916 (2011).
Eyles, J. E. et al. Immunodominant Francisella tularensis antigens identified using proteome microarray. Proteomics 7, 2172–2183 (2007).
Felgner, P. L. et al. A Burkholderia pseudomallei protein microarray reveals serodiagnostic and cross-reactive antigens. Proc Natl Acad Sci U S A 106, 13499–13504 (2009).
Davies, D. H. et al. Proteome-wide analysis of the serological response to vaccinia and smallpox. Proteomics 7, 1678–1686 (2007).
Kalantari-Dehaghi, M. et al. Discovery of potential diagnostic and vaccine antigens in herpes simplex virus 1 and 2 by proteome-wide antibody profiling. J Virol 86, 4328–4339 (2012).
Barbour, A. G. et al. A genome-wide proteome array reveals a limited set of immunogens in natural infections of humans and white-footed mice with Borrelia burgdorferi. Infect Immun 76, 3374–3389 (2008).
Lee, S. J. et al. Identification of a common immune signature in murine and human systemic Salmonellosis. Proc Natl Acad Sci U S A 109, 4998–5003 (2012).
Ansong, C. et al. Proteomics analysis of the causative agent of typhoid fever. J Proteome Res 7, 546–557 (2008).
Eyles, J. E. et al. Immunodominant Francisella tularensis antigens identified using proteome microarray. Proteomics 7, 2172–2183. PMID 17533643 (2007).
Witten, I. H. & Frank, E. Data mining : practical machine learning tools and techniques. 2nd edn, (Morgan Kaufman, Amsterdam, 2005).
Barclay, G. R. & Scott, B. B. Serological relationships between Escherichia coli and Salmonella smooth- and rough-mutant lipopolysaccharides as revealed by enzyme-linked immunosorbent assay for human immunoglobulin G antiendotoxin antibodies. Infect Immun 55, 2706–2714 (1987).
House, D. et al. Antibodies to the Vi capsule of Salmonella Typhi in the serum of typhoid patients and healthy control subjects from a typhoid endemic region. J Infect Dev Ctries 2, 308–312 (2008).
Magnan, C. N. et al. High-throughput prediction of protein antigenicity using protein microarray data. Bioinformatics 26, 2936–2943 (2010).
Dolecek, C. et al. A multi-center randomised controlled trial of gatifloxacin versus azithromycin for the treatment of uncomplicated typhoid fever in children and adults in Vietnam. PLoS One 3, e2188 (2008).
Dunstan, S. J. et al. A TNF region haplotype offers protection from typhoid fever in Vietnamese patients. Human genetics 122, 51–61 (2007).
Thompson, L. J. et al. Transcriptional response in the peripheral blood of patients infected with Salmonella enterica serovar Typhi. Proc Natl Acad Sci U S A 106, 22433–22438 (2009).
Alexander, N. et al. Multicentre prospective study on dengue classification in four South-east Asian and three Latin American countries. Tropical medicine & international health : TM & IH (2011).
Durbin, B. P., Hardin, J. S., Hawkins, D. M. & Rocke, D. M. A variance-stabilizing transformation for gene-expression microarray data. Bioinformatics 18 Suppl 1, S105–110 (2002).
Huber, W., von Heydebreck, A., Sultmann, H., Poustka, A. & Vingron, M. Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics 18 Suppl 1, S96–104 (2002).
Baldi, P. & Hatfield, G. W. DNA Microarrays and Gene Expression: From Experiments to Data Analysis and Modeling (2002).
Baldi, P. & Brunak, S. R. Bioinformatics: the machine learning approach (MIT Press, 2001).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Roy. Stat. Soc., B 57, 289–300 (1995).
Sundaresh, S. et al. From protein microarrays to diagnostic antigen discovery: a study of the pathogen Francisella tularensis. Bioinformatics 23, i508–i518 (2007).
Doran, M. et al. Oligonucleotide microarray identification of Bacillus anthracis strains using support vector machines. Bioinformatics 23, 487–492 (2007).
Moller, S., Croning, M. D. & Apweiler, R. Evaluation of methods for the prediction of membrane spanning regions. Bioinformatics 17, 646–653 (2001).
Bendtsen, J. D., Nielsen, H., von Heijne, G. & Brunak, S. Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 340, 783–795 (2004).
Gardy, J. L. et al. PSORTb v.2.0: expanded prediction of bacterial protein subcellular localization and insights gained from comparative proteome analysis. Bioinformatics 21, 617–623 (2005).
Acknowledgements
We thank Renee Tsolis for providing Salmonella enterica ty2 genomic DNA. This work was supported by U01AI078213 (PLF) and a subcontract to DHD from R01AI073672 (SJMcS). SB is funded by the OAK foundation through Oxford University, SD and TVTN are funded by the Wellcome Trust of the United Kingdom.
Author information
Authors and Affiliations
Contributions
L.L., S.B., P.L.F. conceived and designed the experiments; L.L., S.J., R.N. performed experiments; L.L., P.L.F. analyzed data; T.N., S.D., S.B. provided reagents; L.L., D.H.D., S.M., S.B. and P.L.F. wrote the paper.
Ethics declarations
Competing interests
L.L., P.L.F. and D.H.D. have patent applications pertaining to this work. P.L.F. and D.H.D. own stock in Antigen Discovery Inc. that has licensed the technology in this work.
Electronic supplementary material
Supplementary Information
Supplementary figures and tables
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Liang, L., Juarez, S., Nga, T. et al. Immune profiling with a Salmonella Typhi antigen microarray identifies new diagnostic biomarkers of human typhoid. Sci Rep 3, 1043 (2013). https://doi.org/10.1038/srep01043
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01043
This article is cited by
-
Typhoid fever
Nature Reviews Disease Primers (2023)
-
Detection of Salmonella Typhi nucleic acid by RT-PCR and anti-HlyE, -CdtB, -PilL, and -Vi IgM by ELISA at sites in Ghana, Madagascar and Ethiopia
BMC Infectious Diseases (2022)
-
The identification of novel immunogenic antigens as potential Shigella vaccine components
Genome Medicine (2021)
-
Evaluation of Salmonella Typhi antigen YncE alongside HlyE for the detection of typhoid fever and its carriers
Medical Microbiology and Immunology (2020)
-
Immunogenicity profiling of protein antigens from capsular group B Neisseria meningitidis
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.