Abstract
Predicting the synthesizability of hypothetical crystals is challenging because of the wide range of parameters that govern materials synthesis. Yet, exploring the exponentially large space of novel crystals for any future application demands an accurate predictive capability for synthesis likelihood to avoid a haphazard trial-and-error. Typically, benchmarks of synthesizability are defined based on the energy of crystal structures. Here, we take an alternative approach to select features of synthesizability from the latent information embedded in crystalline materials. We represent the atomic structure of crystalline materials by three-dimensional pixel-wise images that are color-coded by their chemical attributes. The image representation of crystals enables the use of a convolutional encoder to learn the features of synthesizability hidden in structural and chemical arrangements of crystalline materials. Based on the presented model, we can accurately classify materials into synthesizable crystals versus crystal anomalies across a broad range of crystal structure types and chemical compositions. We illustrate the usefulness of the model by predicting the synthesizability of hypothetical crystals for battery electrode and thermoelectric applications.
Similar content being viewed by others
Introduction
The structure–property relationship often necessitates the synthesis of a specific crystal structure for a chemical composition. An essential aspect to guide and accelerate the discovery of future materials is to predict whether a novel crystalline material is synthesizable or not. However, providing a general metric to identify the probability of successful synthesis of hypothetical crystals is a challenging task because of the broad range of parameters controlling the synthesis process, including processing rates and routes, thermodynamic handles, synthesis techniques, and synthesis scales. While materials synthesis is traditionally guided by the expert-interpreted knowledge of various synthesis conditions1,2, recent computational methods and machine learning approaches provide prospects for predictive capability and guidelines for the synthesis of future materials3,4,5,6,7,8,9,10,11,12. However, there remains a lack of general and accurate predictive models for synthesizability across various crystal structure types and chemical compositions.
A group of pioneering studies has provided metrics of synthesizability based on the relevant thermodynamic free energies of crystalline materials7,8,13. Sun et al.13 performed a high-throughput screening of the energy distribution for a large number of crystal structures by means of density functional theory (DFT) calculations, indicating that a considerable number of low-energy hypothetical crystals are not observed for well-explored chemical compositions and are most likely not easily synthesizable. Based on the presence of these low-energy crystals, they concluded that the energy above the ground state cannot act as a reliable metric for synthesizability. In a later study7, the energy of the amorphous solid (or super-cooled liquid state) with a given chemical composition was used as the limit on the energy scale useful for establishing a necessary condition for synthesis. The basis of this framework is that the zero-temperature enthalpy of the amorphous phase provides an accurate upper bound for the Gibbs energy of synthesizable crystals at any temperature due to the inevitably larger entropy of the amorphous solid compared to ordered crystals. Therefore, crystal structures with enthalpies higher than the amorphous state are predicted as unsynthesizable. However, this benchmark of synthesizability is limited to a specific chemical composition. In other words, while the amorphous solid energy can evaluate the synthesizability of any crystal structure for a given chemical composition, a new energy benchmark must be obtained for a different composition. Additionally, this approach cannot predict low-energy unsynthesizable crystals or high-energy synthesizable crystals (e.g., high-pressure crystalline materials). A detailed discussion is provided in the “Discussion” section.
Only a few studies have employed machine learning to address the issue of synthesizability of crystalline materials8,9,10,11,12,14,15,16,17,18,19. In one of the earliest studies by Hautier et al.15,16 developed a probabilistic model built on an experimental crystal structure database to quantify the likelihood of substitution of certain ions in a compound leading to another compound with the same crystal structure. Later Ryan et al.14 used an atomic fingerprint (which captures the local topology around each crystallographically unique site) alongside other descriptors in a neural network to anticipate the likelihood of substituting a lattice site with other components of a given crystalline compound. Aykol et al.8 modeled the free energy convex hull in the composition space that encompasses the chronological discovery timeline of each composition by an evolving network, where the nodes encode the convex hull and the edges encode the circumstantial factors. They utilized their model to predict the likelihood of successful experimental synthesis of hypothetical materials. Some recent studies utilize expert-knowledge-based parameters instead of structural or chemical features to predict synthesis success9,10,11,12. For example, Kim et al.10 analyzed the literature text via natural language processing methods to determine the significant parameters involved in the synthesis of titania nanotubes by hydrothermal methods, and Raccuglia et al.9 used failed hydrothermal synthesis experimental data to predict the crystallization of vanadium selenites. Tang et al.12 utilized machine learning to optimize synthesis conditions in order to enhance process-related properties in inorganic crystalline solids.
In this work, we present a deep-learning model that can predict the synthesizability of hypothetical crystalline materials. The predictive capability of most existing models for synthesizability is confined to either a specific crystal structure type or a given chemical composition. Our model, however, can predict the synthesizability of any given chemical composition in any given crystal form (see the “Discussion” section for more details). This is achieved by our model’s ability to simultaneously capture the structural and chemical features of synthesizability. Our model represents crystalline materials with color-coded three-dimensional images, from which a low-dimension set of latent structural and chemical features are encoded by a convolutional neural network (CNN). Moreover, we provide instances of the crystal anomaly class. We define crystal anomalies as the hypothetical crystalline materials that are highly unlikely to be synthesized. We select crystal anomalies from the unobserved crystal structures for the most-studied chemical compositions in the published literature. This approach ensures the selection of the most pertinent crystalline materials as anomalies, which would otherwise be very difficult to identify. The machine learning framework used in this study can be extended to serve as a predictive tool for the synthesizability likelihood across a wide range of crystalline materials, from elemental, ionic, and covalent crystals to complex molecular crystals.
Results
The overall framework of the crystal synthesizability model
Our crystal synthesizability model consists of two main components: feature learning and classification. Feature learning consists of encoding the hidden structural and chemical patterns from crystalline materials data, here in the form of three-dimensional images, into a latent space representation. Here, we examine two different approaches of supervised and unsupervised feature learning. In the supervised learning, the two tasks of feature learning and classification are intertwined, i.e., the feature learning is performed using a convolutional encoder that is connected to a neural network classifier, where they share the same parameterization. Consequently, the latent space is learned throughout the classification of the labeled crystal images. In the unsupervised learning, the latent space is learned using a convolutional auto-encoder (CAE) on unlabeled crystal images. The learned latent space is then used as the input layer of a neural network classifier, which is trained on labeled crystal images.
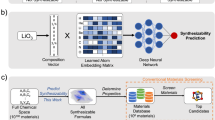
Classification is performed based on the extracted features (or the latent space representation) of two classes of crystalline materials: synthesizable crystals and crystal anomalies, the latter being the hypothetical crystalline materials that are unlikely to be synthesized. Figure 1 summarizes the overall framework of the crystal synthesizability model. In the following sections, we elaborate on data collection and input preparation, model training and validation, and applications of our model to predict the synthesizability of candidate crystalline materials for battery electrodes and thermoelectric applications. Moreover, we apply our model to predict the synthesizability of different crystal structures of molybdenum disulfide.
a Crystal samples for the synthesizable class are obtained from the Crystallographic Open Database (COD). We prepare crystal anomaly samples by using the crystal structure prototype database (CSPD) to generate crystal structures for the most-studied compositions in the published literature that are absent in the COD. b The crystal information files (CIFs) (extracted from the COD or generated by the CSPD) are converted into digitized three-dimensional images, which are used as the inputs for the convolutional encoder or convolutional auto-encoder. c Supervised and unsupervised feature encoding of three-dimensional images are performed using a convolution encoder followed by a multi-layer perceptron (MLP) classifier, referred to as the CNN classifier, and a convolutional auto-encoder (CAE), respectively. The unsupervised latent space representation of crystals from the auto-encoder is used as the input in an MLP classifier, referred to as the CAE+MLP classifier.
Crystal data collection
Crystal anomalies
Generating crystal anomalies is challenging because the unobserved crystals in experimentally synthesized crystal databases can be either crystal anomalies or synthesizable crystals that have not been explored yet. In this study, we identify crystal anomalies to be the unobserved crystal structures for those chemical compositions that are highly repeated in the literature (see Fig. 2a). The underlying assumption is that these compositions are explored enough so that all possible synthesizable crystal structures have already been observed. The hypothetical crystal structures that are not observed are most likely unsynthesizable, i.e., crystal anomalies. We utilize a natural language processing model developed by Tshitoyan et al.20 that encompasses material science literature knowledge from 1922 to 2018. We rank the available chemical compositions in the literature according to their repetition frequency. For example, Fig. 2b illustrates the 15 most-frequently-repeated chemical compositions in the materials science literature since 1922. The top 0.1% constitutes the first 108 unique compositions that are repeated at least 3306 times. We select the unobserved crystal structures for the top 108 compositions as crystal anomaly samples. For each selected composition, we balance the number of structures between the two classes of synthesizable and anomaly by restricting the number of generated anomaly structures to at most the same number of distinct structures that have been already synthesized and are available in the Crystallographic Open Database (COD, 2019)21,22,23,24,25,26. Additionally, we ensure that at least five unobserved structures are generated for each composition. Figure 2c shows the number of anomaly crystals and synthesized crystals for the 15 most-studied compositions. A total number of 600 crystal anomaly samples are generated according to the aforementioned approach. More details about crystal anomaly generation are provided in the “Methods” section, Supplementary Note 1, Supplementary Table 1, and Supplementary Fig. 1.
a We select the unobserved hypothetical crystal structures of well-studied compositions in the literature and label them as crystal anomalies. b The total repetitions of the 15 most-studied compositions in materials science literature are shown as an example. We use the top 108 compositions to generate crystal anomalies in this study. c The number of distinct crystal anomalies and observed/synthesized crystal structures for the 15 most-studied compositions are shown by blue and orange bars, respectively. The 15 most-studied compositions are shown as an example while we use the 108 most-studied compositions to generate the anomaly and synthesizable crystal samples.
Synthesizable crystals
We collect synthesizable crystal samples from the COD (2019), an open-access crystallography database of experimentally synthesized crystalline materials. We select 3000 crystal samples from the COD, five times more than the 600 samples generated for the crystal anomaly class. Within the 3000 samples, we include all the distinct crystalline polymorphs available in the COD for the 108 chemical compositions that we have used to generate the crystal anomaly samples, which amount to 367 crystal samples. This strategy ensures that the classifier is not overfitted to non-generalizable patterns in the small set of chemical compositions in the anomaly class. Instead, by including all structurally distinct polymorphs of the synthesizable class that share the same chemical compositions with the anomaly class samples, we provide the necessary structural information that the classifier needs to learn the distinction between the synthesizable and anomaly classes. In fact, our studies on a variety of data sets indicate that including distinct structural polymorphs of the same composition that belong to the two distinct classes significantly enhances the predictive performance of the classifier. The remaining 2633 synthesizable crystal samples are randomly selected from other chemical compositions. The synthesizable crystal samples in our data set span across 156 distinct space groups (see Supplementary Note 2 and Supplementary Fig. 2). Details about the preparation of crystal samples for each class are provided in the “Methods” section.
Limited availability of crystal anomalies
Crystal synthesizability prediction, as the focus of this study, is inherently limited by the availability of crystal samples that could be labeled as unsynthesizable or anomalies with high confidence. Our case studies suggest that including more than a couple hundred of the most-studied chemical compositions in the anomaly class will hamper the predictive power of the classifier. This indicates that while including a wider range of chemical compositions can reduce the sample bias, it increases the risk of mislabeling positive samples as negative. The increased risk of mislabeling arises from the lower confidence we have in labeling the unobserved crystal polymorphs of a composition that is not repeated enough in the literature, or explored enough, as anomaly.
Data set
The final data set consists of 3000 synthesizable crystals (i.e., positive samples) and 600 crystal anomalies (i.e., negative samples), which are randomly partitioned into the training (49%), validation (21%), and test (30%) sets. To correct the disproportion in the number of positive and negative samples, we randomly duplicate the negative samples in the training set, resulting in an equal number of samples between the two classes.
Crystal representation by three-dimensional images
We represent the atomic structure of crystals by three-dimensional pixel (or voxel) images that incorporate both the structural and chemical patterns embedded within crystals. Each crystal unit cell is replicated to fill a cube with a side length of 70 Å, which is then digitized into 128 voxels on each side. If a voxel is occupied by an element, i.e., a chemical component lies inside a voxel, then the normalized atomic number, periodic row number, and periodic group number are assigned as its three channels. Otherwise, all the channels are set to zero. The three-dimensional image representation of crystals in this study enables the use of CNNs to encode the latent space representation as features of synthesizability. More details about preparing three-dimensional image representation of crystals are presented in the “Methods” section.
Supervised feature learning and classification
In the supervised model, the features of synthesizability are learned from labeled crystal images. A CNN or convolutional encoder is used to map the high-dimensional information embedded in raw crystal images into the reduced dimension latent space representation. The encoder is connected to a multi-layer perceptron (MLP) classifier with the latent space representation as the input layer. The parameters of the encoder and the classifier are optimized simultaneously over the labeled crystal images using a single loss function. Hereafter, we refer to this model as the CNN classifier. Supplementary Figure 7a illustrates the detailed architecture of the CNN classifier. More details about the CNN classifier optimization are provided in the “Methods” section.
To evaluate the predictive power of the CNN classifier, we use the area under the receiver operating characteristic curve (ROC-AUC). Figure 3 illustrates the ROC curve for the CNN classifier, evaluated on the test set with 1080 crystal images. The ROC-AUC value is 0.981 and the calculated accuracy is 93.7%. The decision threshold is set to 0.5, which means that a synthesizability likelihood prediction equal to or above 0.5 is labeled as synthesizable and below 0.5 is labeled as anomaly.
The receiver operating characteristic (ROC) curves for the (a) CNN and (b) CAE+MLP classifiers shown by the red line. Each classifier’s ROC is compared with a dumb classifier, which randomly assigns class labels, shown by blue dashed lines. The normalized distribution of synthesizability likelihood of crystals in the test set are represented for (c) the CNN classifier and (d) the CAE+MLP classifier. e The main classification metrics based on a decision threshold of 0.5. AUC stands for the area under the ROC curve.
Unsupervised feature learning and classification
In the unsupervised model, we utilize a CAE for learning the latent structural and chemical features in unlabeled crystal images. The CAE maps the voxels of crystal images to the latent representation vector by an encoder and then maps the latent representation vector onto a reconstructed crystal image using a decoder. The learned latent space representation is a set of reduced-dimension patterns underlying the raw crystal images that can be used to reconstruct them. Since the CAE has no explicit knowledge of the labels, the learned features can be interpreted as a more general set of features to be used as the feature space for any learning task that maps structural and chemical patterns at the atomic level to a class label or property. Supplementary Figure 6 illustrates the detailed architecture of the CAE used to encode and decode the three-dimensional crystal images.
The latent space representation is flattened and is then passed as the input to a separate MLP classifier. The MLP architecture is identical to the fully-connected layers of the CNN classifier (see Supplementary Fig. 7). The MLP classifier is optimized using the pre-trained latent space representation from the CAE. The latent space representation provides cues about how to group the crystal image examples in the training set. Hereafter, we refer to this model as the CAE+MLP classifier. More details about the design of the CAE+MLP classifier and its optimization are provided in Supplementary Fig. 7b and in the “Methods” section.
In general, neural networks tend to overfit to the training data due to the large number of fitting parameters in their complex non-linear structure. In the CAE, the number of fitting parameters is large because of the size of three-dimensional images and the design of the hidden layers. Therefore, we apply dropout layers for regularizing the CAE. The regularization helps to avoid overfitting to the training data and renders the CAE+MLP model to generalize better (see more details in the “Methods” section).
Figure 3 illustrates the ROC curve for the CAE+MLP classifier, evaluated on the test set with 1080 crystal images. The ROC-AUC value is 0.968 and the calculated accuracy is 91.9% for a decision threshold of 0.5.
Importance of feature learning
To assess the importance of feature learning in our models, we design an MLP classifier that uses flattened raw images as the input layer. Instead of using a convolutional encoder to learn the latent space representation of three-dimensional crystal images, we flatten raw crystal images and pass them to the input layer of an MLP classifier. Here, the MLP classifier has the same design as in the CNN and the CAE+MLP models. Hereafter, we refer to this model as the raw image classifier. The raw image classifier is overfitted to the training data, as shown by its low accuracy on the validation data during training in Supplementary Fig. 9. This overfitting is inevitable because of the design of the raw image classifier. Supplementary Table 2 shows the details of the raw image classifier design. As shown in Supplementary Table 2, the raw image classifier has more than 80 million trainable parameters, making it a highly over-parameterized model for such limited training data. This result indicates the importance of the feature learning step in our model as an efficient dimension reduction technique, which avoids over-parameterization and the consequent overfitting. As expected, the performance of the raw image classifier is poor compared to the CNN and CAE+MLP classifiers. As shown in Supplementary Fig. 9, for the raw image classifier, the ROC-AUC and the classification accuracy on the test data are 0.685 and 80%, respectively. A classifier that assigns a synthesizable label to all the samples in the test set reaches an accuracy of 83%. The accuracy of the raw image classifier is below such classifier. See Supplementary Note 8 for more detail.
Comparison of supervised and unsupervised feature learning
Figure 3e compares the performance of the CNN and the CAE+MLP classifiers on the test data. The accuracy and sensitivity of the two classifiers are almost the same; however, the specificity of the CNN classifier is higher than the CAE+MLP classifier. The out-performance of the CNN specificity stems from the supervised nature of the feature learning process, in which the latent space has been trained with the explicit knowledge of the class labels. Considering that the unsupervised feature learning in the CAE has no knowledge about the class labels, the CAE+MLP prediction is very accurate in the context of a binary classification task. This accurate classification shows that the auto-encoder can successfully extract the correct latent features.
To compare the generalization of the CNN and the CAE+MLP classifiers, we predict the synthesizability of two sets of crystal samples beyond the test set, namely electrodes and thermoelectric crystalline materials. By going outside the test data, we can evaluate the generality of the model for a set with a different distribution of samples. Because we randomly partition the samples between the training and test sets, the distribution of crystal samples in the test set is more similar to the training set than the distribution of samples in the electrode and the thermoelectric sets (see Supplementary Note 5 and Supplementary Fig. 5).
As detailed in the following subsections, the CAE+MLP shows a stronger generalization compared to the CNN classifier on both the electrode and thermoelectric materials. This indicates the power of unsupervised learning, which does not overfit to a particular data set. In other words, the general nature of the features that are learned by the CAE in an unsupervised manner leads to the stronger generalization of the CAE+MLP classifier. This result is remarkable and further encourages the use of unsupervised learning for different tasks, given that proper unsupervised learning methods are used. Supplementary Note 9 and Supplementary Fig. 10 provide a more detailed comparison of the two classifiers.
Case study: electrode materials
We apply our model to predict the synthesizability likelihood for hypothetical crystals in the Battery Explorer database of the Materials Project27,28,29, which are electrode candidates satisfying critical criteria such as high voltage, high volumetric capacity, and high energy density. We selected 2088 crystal samples of electrode materials, out of which 264 samples exist in the COD database. As shown in Fig. 4, the CNN and the CAE+MLP classifiers predict the synthesizability likelihood for the battery materials that exist in the COD (i.e., samples that are positive, or synthesizable) with an accuracy of 82% and 89%, respectively. The prediction accuracy for COD samples is a measure of the recall of the classifier, which is slightly lower than the recall for the test data (93% versus 82% for the CNN classifier and 94% versus 89% for the CAE+MLP classifier). The lower recall on electrode samples is likely due to the larger statistical error arising from the small sample size (264 samples here versus 1080 samples in the test set). Additionally, the test set samples are more similar to the crystal samples in the training set because they have been partitioned randomly, unlike the electrode samples that belong to a different range of chemical compositions (see Supplementary Fig. 5). This similarity between the training and test set samples lead to a higher recall on the test set. The CNN and the CAE+MLP classify 73% and 85% of the non-COD samples as synthesizable, respectively.
The probability distribution function of the 2088 electrode crystals from the Battery Explorer database of the Materials Project (MP) is predicted by (a) the CNN and (b) the CAE+MLP classifiers, respectively. The green and yellow bars show the samples from the COD and those absent in the COD, respectively. Synthesizability refers to the ratio of predicted synthesizable samples to the crystal samples in the non-COD electrode data. The sensitivity (or recall) is calculated based on the 264 electrode samples that belong to the COD.
To illustrate the usefulness of our model, we show the synthesizability predictions against a second property of interest, inspired by Ashby charts30, where synthesizability likelihood can serve as a design parameter or a parameter for materials selection. For example, Fig. 5 shows the synthesizability of samples, predicted by the CAE+MLP, against their total volumetric capacity and average voltage. These plots can be utilized to identify the best materials, where the ideal candidates lie near the top right on the charts in this case. The top right region in Fig. 5a corresponds to a high volumetric capacity combined with a high likelihood of synthesizability. Similarly, the top right region in Fig. 5b corresponds to a high voltage combined with a high likelihood of synthesizability. For more synthesizability predictions for electrode materials, refer to Supplementary Fig. 11, Supplementary Table 3, and Supplementary Note 10.
Synthesizability likelihood of selected electrode samples against their (a) volumetric capacity and (b) average voltage. Each crystal sample is identified by its chemical formula followed by the space group number in parenthesis. For the sake of visualization, each working-ion is color-coded differently: Li as green, Na as orange, and Ca as blue. These plots can be utilized similarly to Ashby charts for materials selection. In this case, the ideal candidates lie near the top right of each plot to increase the synthesizability and the desired electrode property.
Case study: thermoelectric materials
As a second case study, we employ our synthesizabilty prediction model on hypothetical crystal structures that have been proposed as thermoelectric materials by Tshitoyan et al.20. To discover promising thermoelectric materials, Tshitoyan’s model encodes the knowledge in the published literature into vector representation of words. They proposed a total of 9483 chemical compositions with cosine similarities to the word “thermoelectric”, of which the top 10 thermoelectric composition candidates (not crystal structures) were identified for each year from 2001 to 2018. We selected all the available crystal structures in the Materials Project and the COD databases for the yearly proposed candidate compositions, which resulted in 122 crystal samples. As shown in Fig. 6, we predicted the synthesizability likelihood of these samples based on the CNN and the CAE+MLP classifiers. The classification recall on the COD thermoelectric samples (56 out of 122 crystal samples), which indicates the rate of positive predictions to the total true positive samples, is 64.3% and 78.6% for the CNN and the CAE+MLP classifiers, respectively. On the other hand, the classification recall of the test data set is 93% and 94% for the CNN and the CAE+MLP classifiers, respectively (see Fig. 3e). The lower recall of the classifiers for the thermoelectric samples is a likely result of larger statistical errors due to smaller sample size (56 versus 1080) and a possible out-performance of predictions on the test data due to their similarities with the training set samples (see Supplementary Fig. 5).
The synthesizability likelihoods evaluated by (a) the CNN and (b) the CAE+MLP classifiers. The total number of thermoelectric crystal samples are 122, from which 56 are collected from the COD and the rest are collected from the Materials Project database. The green and yellow bars show the samples from the COD and those absent in the COD, respectively. Synthesizability refers to the ratio of predicted synthesizable samples to the non-COD crystal samples in our thermoelectric data. The sensitivity is calculated based on the 56 samples that belong to the COD.
We ranked the synthesizability likelihood for a subset of the thermoelectric crystals in Table 1. The synthesizability likelihood predictions for all the thermoelectric crystal candidates from 2002 to 2018 are presented in Supplementary Data 1 and Supplementary Note 11.
Case study: molybdenum disulfide
Our model can be used to identify synthesizable crystal structures for a given chemical composition. For example, Fig. 7 illustrates different crystal structures of molybdenum disulfide, or MoS2, sorted from bottom to top according to their synthesizability likelihood (predicted by the CAE+MLP model). According to Pauling’s rules, one can obtain some insight about the most probable cation and anion coordination in ionic compounds. For MoS2, the cation to anion radius ratio is \(\frac{65}{184}=0.35\). According to the radius ratio rule, the cation (or Mo4+) coordination number should be 4. This results in an electrostatic bond strength of 1 to each coordinated anion (i.e., \(\frac{4}{4}=1\)). According to the electrostatic valence rule, the anion (or S2−) coordination number is 2. Therefore, Pauling’s rules suggest a fourfold cation and a twofold anion coordination.
For MoS2, the experimentally reported bulk crystal structures include three different polytypes, namely 1T (1 atomic layer of tetragonal structure), 2H (2 atomic layer of hexagonal structure), and 3R (3 atomic layer of rhombohedral structure)31. Natural and stable MoS2 is composed of 2H (space group P63/mmc) with less than 3% of 3R (space group R3m). As shown in Fig. 7, the predicted synthesizability likelihood of the hexagonal P63/mmc is 0.82, which we label as synthesizable. The formation of this hexagonal structure is consistent with the Pauling’s guideline because it has a cation and anion coordination of 4 and 2, respectively. Additionally, we predict three tetragonal structures as synthesizable, consistent with the experimentally reported metastable tetragonal structures31.
Discussion
The predictive power of the presented synthesizability framework lies in its ability to combine generality and accuracy, due to the highly non-linear and flexible design of the neural networks used in the feature learning and classification tasks. However, generality and accuracy are gained at the expense of losing the interpretability of the features of synthesizability. The learned features by the CNN or CAE (which are black-box models) are too complicated and not easily understandable in terms of common physical parameters. Translating these complex features into simpler and more understandable features can be achieved through additive feature attribution methods, such as layer-wise relevance propagation32, which is the subject of a future study by the authors.
The framework of our model enables synthesizability prediction across any crystal structure type for any given chemical composition. This distinguishes our model from most of the existing predictive models for synthesizability, which are limited to either a specific crystal structure type or a given chemical composition. To further illustrate this capability of our model, in what follows, we compare it with two existing methods, namely an energy-based threshold model by Aykol et al.7 and a deep network model by Ryan et al.14.
The crystal synthesizability model by Aykol et al.7, hereafter referred to as the stability skyline model, proposes the DFT energy of the amorphous solid phase as the thermodynamic upper limit on the energy scale of synthesizable crystalline materials. The stability skyline model introduces a simple energy-based metric for crystal synthesizability, and because it provides an upper limit for the thermodynamic scale of synthesizability, it can accurately predict high energy crystal anomalies (although it misses high energy synthesizable crystals such as high-pressure phases). However, this model is expected to perform poorly in predicting low-energy crystal anomalies. More specifically, all crystals with an energy lower than the amorphous solid energy are predicted as synthesizable. An earlier high-throughput study13 calculated the DFT energies on a large-scale data set of inorganic crystalline phases and revealed that many low-energy compounds are absent in experimental databases while many high-energy polymorphs are observed or synthesized experimentally. This suggests that the thermodynamic energy scale cannot act as the sole reliable synthesizability metric, i.e., it is an oversimplified descriptor for crystal synthesizability. This stems from the fact that thermodynamic stability cannot be individually mapped into synthesizability, where a range of other parameters, such as kinetic limitations, synthesis routes, and synthesis precursors, can affect the synthesizability likelihood of a given crystalline solid. The advantage of our model is that it can predict low-energy unsynthesizable crystals and high-energy synthesizable crystals according to more complicated chemical and structural patterns, which is not achievable by the upper limit energy approach. In Fig. 8a, we compare the prediction outcomes of our model with those from the stability skyline model. Both the synthesizable crystals and crystal anomalies predicted by the CAE+MLP classifier span over a range of energies above the ground state. The stability skyline model predicts the crystals below the skyline limit as synthesizable and those above it as unsynthesizable, with some categorical exceptions described in ref. 7. As shown in Fig. 8b, our model projects that same energy distribution above the ground state for the predicted crystal anomalies and synthesizable crystals as the stability skyline model7 (compare Fig. 8b with Fig. 2B of ref. 7). This indicates that our model can correctly predict the exceptions in the stability skyline model, which are low-energy crystal anomalies and high-energy synthesizable crystals. The synthesizable crystals distribution shows a higher peak at low energies compared to anomalies, which show a more uniform distribution at higher energy values. Additionally, in the stability skyline model, a new amorphous limit must be calculated for each composition, which limits its predictability to a given chemical composition.
a The energy above the ground state for a total number of 815 crystalline materials studied in ref. 7, alongside the synthesizability predictions of the CAE+MLP classifier. The horizontal bars indicate the amorphous energy limit calculated in ref. 7, above which the stability skyline model predicts a crystal as an anomaly with some exceptions (more details can be found in ref. 7). The lowest computed amorphous energy is represented by the bold black line, while higher amorphous energies are indicated by thin gray lines. The number in front of each legend label indicates the abundance of samples in each class. All the energy values are extracted from the Materials Project database27. b The probability distribution function for the energy above the ground state for the predicted synthesizable crystals and anomalies (based on our model), as well as the amorphous limit obtained from ref. 7.
Aside from energy-based models, some studies use machine learning to predict the probability of crystal formation. The study by Ryan et al.14 predicts the probability of an individual crystallographic site to be occupied by various elements in the periodic table. While their model cannot explicitly predict the synthesizability of a hypothetical crystal structure, it can predict the probability of element substitution for each crystallographic site within an observed crystal structure. This implicitly confines their model’s predictive power to a specific crystal structure and its minor structural variations. This arises from the architecture of their model, in which crystal information is represented by multi-perspective atomic fingerprints, which serve as descriptors in a neural network model. More specifically, the atomic fingerprint can encode the local topology in the vicinity of each site by translating the high-dimensional position-composition space into a lower dimension radial distance space, which can lead to information loss. Our model surpasses this limitation by combining a more global structural and chemical pattern representation using voxel-wise images. Additionally, feature learning through the convolutional encoder used in this study is a suitable design because of the inherent long-range transnational symmetry of crystals.
Methods
Crystal Data Collection
The COD (2019) database is used to collect all the crystallographic information files (CIFs) for synthesizable crystals. For crystal anomaly samples, the CIF files are generated using the Crystal Structure Prototype Database (CSPD) toolkit33. Only CIF files that can be successfully parsed by the Atomic Simulation Environment (ASE) package34,35 are considered as readable and are included in our data set. Crystals with the minimum inter-atomic separation below 0.947 Å are excluded from our data set (due to the model’s resolution constraint described below); these are mostly crystals with hydrogen atoms as ligands. Crystal files with partial site occupancy, as a result of configuration disorder or positional disorder, are translated into supercell structures without point disorders by utilizing the Supercell software package36. Anomalies are selected by eliminating the experimentally synthesized structures in the COD out of the generated structures by the CSPD for the 0.1% most-studied compositions in the literature, as shown in Fig. 2. More information about anomaly crystal structures is presented in Supplementary Table 1 and Supplementary Fig. 1.
Crystal representation by three-dimensional Images
All the CIF files are parsed using the ASE software package and then are converted into three-dimensional cubic images using our in-house code. To translate the CIF files into three-dimensional images, each unit cell is replicated as many times as needed to fill a cube with a side length of 70 Å. This choice of cube length ensures that more than 96% of crystals in the COD are replicated at least twice along the largest lattice vector. Supplementary Figure 3a illustrates the distribution of the three lattice constants for all the readable crystal files in the COD. The distribution of crystals with the largest lattice constant higher than 35 Å is slightly less than 4%. We digitize the cubic structures to (128 × 128 × 128) voxel three-channeled images. Each voxel has three channels, which are the atomic number, the periodic table row number, and the group number of the chemical element occupying the voxel or zero otherwise. We consider the group number of 3.5 for the elements of the lanthanides and actinides. The channels are normalized by dividing them by the highest atomic number (118), the highest row number (7), and the highest group number (18) plus the number one (1). To avoid a voxel occupation by more than one chemical component, we exclude crystal structures with the nearest neighbor distance less than 0.947 Å, which mostly contains lingering hydrogen atoms. The nearest neighbor distance distribution for all the crystals in the COD is shown in Supplementary Fig. 3b. Although higher image resolutions are possible by increasing the number of voxels, the selected resolution ensures sufficient accuracy in representing the crystal structures while keeping the computational costs of convolution tasks feasible (see Supplementary Note 3). Supplementary Figure 4 illustrates the translation of diopside (CaMgO6Si2) crystal information into a three-dimensional image color-coded by its chemical attributes as an example (see Supplementary Note 4 for more information).
CAE design
The encoder and decoder blocks of the CAE learn the latent representation of crystal images and reconstruct it based on the latent representation, respectively. As shown in Supplementary Fig. 6, the design of the CAE consists of three layers for the encoding and four layers for the decoding, which is implemented using the Keras python package37. Each encoding layer consists of a convolutional sub-layer with a rectified linear unit (ReLu) as the activation function and a filter with a size of (3 × 3 × 3). The filter convolves within the three-dimensional image and constructs a new set of images based on the filter’s feature maps. A max-pooling sub-layer follows the convolutional sub-layer with a pool size of (4 × 4 × 4) for the first two layers and (2 × 2 × 2) for the third layer. The max-pooling sub-layer reduces the size of an image by the rate of the pooling size. The first two convolutional sub-layers in the encoding block output 32 channels while the last one outputs 64. The decoder has a reverse layer architecture with up-sampling sub-layers adopted instead of max-pooling sub-layers. The order of the sub-layers in the layers remain intact. Each max-pooling or up-sampling sub-layer is followed by a dropout layer with a 30% dropout rate, which is the probability that outputs of the preceding layer are dropped out. The dropout layers regularize the auto-encoder and make the fit network more general. At the end of the decoder, there is an extra layer compared to the encoding block. This layer has a convolutional sub-layer outputting three channels followed by a sigmoid activation function to reconstruct images with three normalized channels. This last layer does not have an up-sampling sub-layer. Supplementary Figure 6 depicts the detailed design of the CAE. See Supplementary Note 6 for more information. The convolution filters are trained by minimizing the difference between the original and reconstructed images (or the reconstruction error), measured by the loss function. The stochastic Adam optimization method38 is used to minimize the per-voxel binary-cross-entropy loss function of the following form:
where xi and \({\hat{x}}_{i}\) represent the ith voxel of the original and reconstructed images, respectively. The CAE constitutes 281,923 fitting parameters. Supplementary Figure 8 illustrates the loss function variation during the training. See Supplementary Note 7 for more information.
CNN design
The CNN encoder consists of three hidden layers. Each layer consists of a convolutional sub-layer with a ReLu activation function and a (3 × 3 × 3) filter which outputs 32 channels. A max-pooling sub-layer follows the convolutional sub-layer with a pool size of (4 × 4 × 4) for all the three layers. The latent space from the CNN is the input layer for the connected MLP classifier described below (see Supplementary Fig. 7).
We utilize the stochastic Adam optimizer38 to minimize the binary-cross-entropy loss between true labels and predicted labels according to the following equation:
where yi and \({\hat{y}}_{i}\in (0,1)\) represent the true labels and the predicted values for the ith crystal sample, respectively, and N is the total number of crystal samples in the training set. The CNN classifier (i.e., the connected CNN and the MLP classifier) constitutes 61,703 fitting parameters. Supplementary Figure 8a, b illustrates the loss function and accuracy convergence of the CNN classifier during the training process for the training and validation sets.
The MLP classifier design
We use an MLP classifier to classify crystal anomalies versus synthesizable crystals. The input layer is the latent space representation from the encoder block of the CNN or the CAE. The classifier has three fully-connected hidden layers each with 13 nodes (see Supplementary Fig. 7).
Packages
We have used the following Python libraries and software packages for completing this work: ASE35, Keras37, TensorFlow39, Scikit-Learn40, CSPD33, Supercell Program36, Pandas41. For visualization of CIFs and three-dimensional crystal images, we used the Visualization for Electronic and STructure Analysis (VESTA)42 and the Visual Molecular Dynamics (VMD)43 packages, respectively.
Data availability
The crystal images generated and/or analyzed in this study are available in the GitHub repository https://github.com/kadkhodaei-research-group/XIE-SPP.
Code availability
The codes generated in this study are available in the GitHub repository https://github.com/kadkhodaei-research-group/XIE-SPP.
References
Stein, A., Keller, S. W. & Mallouk, T. E. Turning down the heat: design and mechanism in solid-state synthesis. Science 259, 1558–1564 (1993).
Price, S. L. Why don’t we find more polymorphs? Acta Crystallogr. B Struct. Sci. Cryst. Eng. Mater. 69, 313–328 (2013).
Maddox, J. Crystals from first principles. Nature 335, 201–201 (1988).
Oganov, A. R. Modern Methods of Crystal Structure Prediction (Wiley, 2011).
Tanaka, I., Rajan, K. & Wolverton, C. Data-centric science for materials innovation. MRS Bull. 43, 659–663 (2018).
Curtarolo, S. et al. The high-throughput highway to computational materials design. Nat. Mater. 12, 191–201 (2013).
Aykol, M., Dwaraknath, S. S., Sun, W. & Persson, K. A. Thermodynamic limit for synthesis of metastable inorganic materials. Sci. Adv. 4, eaaq0148 (2018).
Aykol, M. et al. Network analysis of synthesizable materials discovery. Nat. Commun. 10, 2018 (2019).
Raccuglia, P. et al. Machine-learning-assisted materials discovery using failed experiments. Nature 533, 73–76 (2016).
Kim, E. et al. Materials synthesis insights from scientific literature via text extraction and machine learning. Chem. Mater. 29, 9436–9444 (2017).
Huo, H. et al. Semi-supervised machine-learning classification of materials synthesis procedures. npj Comput. Mater. 5, 62 (2019).
Tang, B. et al. Machine learning-guided synthesis of advanced inorganic materials. Mater. Today 41, 72–80 (2020).
Sun, W. et al. The thermodynamic scale of inorganic crystalline metastability. Sci. Adv. 2, e1600225 (2016).
Ryan, K., Lengyel, J. & Shatruk, M. Crystal structure prediction via deep learning. J. Am. Chem. Soc. 140, 10158–10168 (2018).
Hautier, G., Fischer, C. C., Jain, A., Mueller, T. & Ceder, G. Finding natures missing ternary oxide compounds using machine learning and density functional theory. Chem. Mater. 22, 3762–3767 (2010).
Hautier, G., Fischer, C., Ehrlacher, V., Jain, A. & Ceder, G. Data mined ionic substitutions for the discovery of new compounds. Inorg. Chem. 50, 656–663 (2011).
Takahashi, K. & Takahashi, L. Creating machine learning-driven material recipes based on crystal structure. J. Phys. Chem. Lett. 10, 283–288 (2019).
Kim, K. et al. Machine-learning-accelerated high-throughput materials screening: discovery of novel quaternary Heusler compounds. Phys. Rev. Mater. 2, 123801 (2018).
Schmidt, J. et al. Predicting the thermodynamic stability of solids combining density functional theory and machine learning. Chem. Mater. 29, 5090–5103 (2017).
Tshitoyan, V. et al. Unsupervised word embeddings capture latent knowledge from materials science literature. Nature 571, 95–98 (2019).
Quirós, M., Gražulis, S., Girdzijauskaitė, S., Merkys, A. & Vaitkus, A. Using SMILES strings for the description of chemical connectivity in the Crystallography Open Database. J Cheminformatics 10, 23 (2018).
Merkys, A. et al. COD::CIF::Parser: an error-correcting CIF parser for the Perl language. J. Appl. Crystallogr. 49, https://doi.org/10.1107/S1600576715022396 (2016).
Gražulis, S., Merkys, A., Vaitkus, A. & Okulič-Kazarinas, M. Computing stoichiometric molecular composition from crystal structures. J. Appl. Crystallogr. 48, 85–91 (2015).
Gražulis, S. et al. Crystallography open database (COD): an open-access collection of crystal structures and platform for world-wide collaboration. Nucleic Acids Res. 40, D420–D427 (2012).
Gražulis, S. et al. Crystallography Open Database—an open-access collection of crystal structures. J. Appl. Crystallogr. 42, 726–729 (2009).
Downs, R. T. & Hall-Wallace, M. The American mineralogist crystal structure database. Am. Mineralogist 88, 247–250 (2003).
Jain, A. et al. The Materials Project: a materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
Zhou, F., Cococcioni, M., Marianetti, C. A., Morgan, D. & Ceder, G. First-principles prediction of redox potentials in transition-metal compounds with LDA+U. Phys. Rev. B 70, 235121 (2004).
Ong, S. P. et al. The materials application programming interface (API): a simple, flexible and efficient API for materials data based on REpresentational state transfer (REST) principles. Comput. Mater. Sci. 97, 209–215 (2015).
Ashby, M. F. In Materials Selection in Mechanical Design, Ch. 5, 4th edn 97–124 (ed. Ashby, M. F.) (Butterworth-Heinemann, 2011). http://www.sciencedirect.com/science/article/pii/B9781856176637000059
Krishnan, U., Kaur, M., Singh, K., Kumar, M. & Kumar, A. A synoptic review of mos2: synthesis to applications. Superlattices Microstruct. 128, 274–297 (2019).
Lundberg, S. M. & Lee, S.-I. In Advances in Neural Information Processing Systems (eds Guyon, I. et al.) Vol. 30. 4765–4774 (Curran Associates, Inc., 2017).
Su, C. et al. Construction of crystal structure prototype database: methods and applications. J. Phys. Condens. Matter 29, 165901 (2017).
Bahn, S. R. & Jacobsen, K. W. An object-oriented scripting interface to a legacy electronic structure code. Comput. Sci. Eng. 4, 56–66 (2002).
Larsen, A. H. et al. The atomic simulation environment—a python library for working with atoms. J. Phys. Condens. Matter 29, 273002 (2017).
Okhotnikov, K., Charpentier, T. & Cadars, S. Supercell program: a combinatorial structure-generation approach for the local-level modeling of atomic substitutions and partial occupancies in crystals. J. Cheminformatics 8, 17 (2016).
Chollet, F. et al. Keras. https://keras.io (2015).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980 (2014).
Abadi, M. et al. TensorFlow: large-scale machine learning on heterogeneous systems. Software available from tensorflow.org. https://www.tensorflow.org/ (2015).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
pandas-dev/pandas: Pandas. https://doi.org/10.5281/zenodo.3509134Feb (2020).
Momma, K. & Izumi, F. Vesta: a three-dimensional visualization system for electronic and structural analysis. J. Appl. Crystallogr. 41, 653–658 (2008).
Humphrey, W., Dalke, A. & Schulten, K. VMD—visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Acknowledgements
This research is based upon work supported by the National Science Foundation (NSF) under Award Numbers DMR-1954621 and DMR-2119308 as well as by Sara Kadkhodaei’s startup grant at UIC. We used the Extreme Science and Engineering Discovery Environment (XSEDE) resources through allocation TG-MAT200013 and resources at the Electronic Visualization Laboratory (EVL) at UIC available through the NSF Award CNS-1828265. Additionally, we would like to thank Dr. A. Mohammadian at UIC for kindly sharing his computer resources at his lab.
Author information
Authors and Affiliations
Contributions
A.D. conducted all the calculations and developed different components of the presented method. Z.K. contributed to developing the idea of crystal representation by three-dimensional images and provided insight about the image processing and machine learning algorithms. S.K. supported and supervised the study. All authors analyzed and discussed the results and co-wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Communications Materials thanks Edward Kim and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Milica Todorović and Aldo Isidori.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Davariashtiyani, A., Kadkhodaie, Z. & Kadkhodaei, S. Predicting synthesizability of crystalline materials via deep learning. Commun Mater 2, 115 (2021). https://doi.org/10.1038/s43246-021-00219-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43246-021-00219-x
This article is cited by
-
Accelerating the prediction of stable materials with machine learning
Nature Computational Science (2023)
-
Formation energy prediction of crystalline compounds using deep convolutional network learning on voxel image representation
Communications Materials (2023)
-
Predicting the synthesizability of crystalline inorganic materials from the data of known material compositions
npj Computational Materials (2023)
-
Machine learned synthesizability predictions aided by density functional theory
Communications Materials (2022)
-
Perovskite synthesizability using graph neural networks
npj Computational Materials (2022)