Abstract
Tumor progression is accompanied by fibrosis, a condition of excessive extracellular matrix accumulation, which is associated with diminished antitumor immune infiltration. Here we demonstrate that tumor-associated macrophages (TAMs) respond to the stiffened fibrotic tumor microenvironment (TME) by initiating a collagen biosynthesis program directed by transforming growth factor-β. A collateral effect of this programming is an untenable metabolic milieu for productive CD8+ T cell antitumor responses, as collagen-synthesizing macrophages consume environmental arginine, synthesize proline and secrete ornithine that compromises CD8+ T cell function in female breast cancer. Thus, a stiff and fibrotic TME may impede antitumor immunity not only by direct physical exclusion of CD8+ T cells but also through secondary effects of a mechano-metabolic programming of TAMs, which creates an inhospitable metabolic milieu for CD8+ T cells to respond to anticancer immunotherapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
RNAseq data described here have been deposited in Gene Expression Omnibus and is publicly available (referenced accession numbers: GSE157290 and GSE184398), and data for the breast invasive carcinoma, glioblastoma multiforme, kidney renal clear cell carcinoma, lung adenocarcinoma and pancreatic adenocarcinoma projects were acquired from TCGA repository (https://portal.gdc.cancer.gov/). Source data for Figs. 1b–d, 3f,h,i,j, 4a–d,h,j, 5e,g,h, 6b–f,h and Fig. 7b–f,i and Extended Data Figs. 1a,b,d, 2, 3a,d,e, 4a,c–i, 5a,e,f,g and 6b have been provided as source data files. Minimally processed metabolomics datasets are provided in Supplementary Table 2. All other data supporting the findings of this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
Open-source R scripts for metabolomics analysis can be found at https://rdrr.io/github/graeberlab-ucla/MetabR/.
References
Galon, J. et al. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313, 1960–1964 (2006).
Tumeh, P. C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014).
Bagaev, A. et al. Conserved pan-cancer microenvironment subtypes predict response to immunotherapy. Cancer Cell 39, 845–865 (2021).
Sun, X. et al. Tumour DDR1 promotes collagen fibre alignment to instigate immune exclusion. Nature 599, 673–678 (2021).
Nicolas-Boluda, A. et al. Tumor stiffening reversion through collagen crosslinking inhibition improves T cell migration and anti-PD-1 treatment. eLife 10, e58688 (2021).
Acerbi, I. et al. Human breast cancer invasion and aggression correlates with ECM stiffening and immune cell infiltration. Integr. Biol. (Camb.) 7, 1120–1134 (2015).
Maller, O. et al. Tumour-associated macrophages drive stromal cell-dependent collagen crosslinking and stiffening to promote breast cancer aggression. Nat. Mater. 20, 548–559 (2021).
Tauriello, D. V. F. et al. TGFβ drives immune evasion in genetically reconstituted colon cancer metastasis. Nature 554, 538–543 (2018).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Chakravarthy, A., Khan, L., Bensler, N. P., Bose, P. & De Carvalho, D. D. TGF-β-associated extracellular matrix genes link cancer-associated fibroblasts to immune evasion and immunotherapy failure. Nat. Commun. 9, 4692 (2018).
Sahai, E. et al. A framework for advancing our understanding of cancer-associated fibroblasts. Nat. Rev. Cancer 20, 174–186 (2020).
Scharping, N. E. et al. Mitochondrial stress induced by continuous stimulation under hypoxia rapidly drives T cell exhaustion. Nat. Immunol. 22, 205–215 (2021).
Buck, M. D., Sowell, R. T., Kaech, S. M. & Pearce, E. L. Metabolic instruction of immunity. Cell 169, 570–586 (2017).
Geiger, R. et al. l-Arginine modulates T cell metabolism and enhances survival and anti-tumor activity. Cell 167, 829–842 (2016).
Canale, F. P. et al. Metabolic modulation of tumours with engineered bacteria for immunotherapy. Nature 598, 662–666 (2021).
Ma, E. H. et al. Metabolic profiling using stable isotope tracing reveals distinct patterns of glucose utilization by physiologically activated CD8+ T cells. Immunity 51, 856–870 (2019).
Rodriguez, P. C. et al. l-Arginine consumption by macrophages modulates the expression of CD3ζ chain in T lymphocytes. J. Immunol. 171, 1232–1239 (2003).
Reinfeld, B. I. et al. Cell-programmed nutrient partitioning in the tumour microenvironment. Nature 593, 282–288 (2021).
Gabrilovich, D. I., Ostrand-Rosenberg, S. & Bronte, V. Coordinated regulation of myeloid cells by tumours. Nat. Rev. Immunol. 12, 253–268 (2012).
Grzywa, T. M. et al. Myeloid cell-derived arginase in cancer immune response. Front. Immunol. 11, 938 (2020).
Wei, Z., Oh, J., Flavell, R. A. & Crawford, J. M. LACC1 bridges NOS2 and polyamine metabolism in inflammatory macrophages. Nature https://doi.org/10.1038/s41586-022-05111-3 (2022).
Menjivar, R. E. et al. Arginase 1 is a key driver of immune suppression in pancreatic cancer. eLife https://doi.org/10.7554/eLife.80721 (2023).
Li, S. et al. Metabolism drives macrophage heterogeneity in the tumor microenvironment. Cell Rep. 39, 110609 (2022).
Mouw, J. K. et al. Tissue mechanics modulate microRNA-dependent PTEN expression to regulate malignant progression. Nat. Med. 20, 360–367 (2014).
Broz, M. L. et al. Dissecting the tumor myeloid compartment reveals rare activating antigen-presenting cells critical for T cell immunity. Cancer Cell 26, 638–652 (2014).
Afik, R. et al. Tumor macrophages are pivotal constructors of tumor collagenous matrix. J. Exp. Med. 213, 2315–2331 (2016).
Schnoor, M. et al. Production of type VI collagen by human macrophages: a new dimension in macrophage functional heterogeneity. J. Immunol. 180, 5707–5719 (2008).
Karsdal, M. A. (ed.) Biochemistry of Collagens, Laminins and Elastin: Structure Function and Biomarkers https://doi.org/10.1016/C2015-0-05547-2 (Elsevier, 2016).
Combes, A. J. et al. Discovering dominant tumor immune archetypes in a pan-cancer census. Cell 185, 184–203 (2022).
Yu, X. et al. The cytokine TGF-β promotes the development and homeostasis of alveolar macrophages. Immunity 47, 903–912 (2017).
Paszek, M. J. et al. Tensional homeostasis and the malignant phenotype. Cancer Cell 8, 241–254 (2005).
Peranzoni, E. et al. Macrophages impede CD8 T cells from reaching tumor cells and limit the efficacy of anti-PD-1 treatment. Proc. Natl Acad. Sci. USA 115, E4041–E4050 (2018).
Sinha, P., Clements, V. K. & Ostrand-Rosenberg, S. Reduction of myeloid-derived suppressor cells and induction of M1 macrophages facilitate the rejection of established metastatic disease. J. Immunol. 174, 636–645 (2005).
Leone, R. D. & Powell, J. D. Metabolism of immune cells in cancer. Nat. Rev. Cancer 20, 516–531 (2020).
Lim, A. R., Rathmell, W. K. & Rathmell, J. C. The tumor microenvironment as a metabolic barrier to effector T cells and immunotherapy. eLife 9, e55185 (2020).
Bantug, G. R., Galluzzi, L., Kroemer, G. & Hess, C. The spectrum of T cell metabolism in health and disease. Nat. Rev. Immunol. 18, 19–34 (2018).
Rossiter, N. J. et al. CRISPR screens in physiologic medium reveal conditionally essential genes in human cells. Cell Metab. 33, 1248–1263 (2021).
Leney-Greene, M. A., Boddapati, A. K., Su, H. C., Cantor, J. R. & Lenardo, M. J. Human plasma-like medium improves T lymphocyte activation. iScience 23, 100759 (2020).
Cantor, J. R. et al. Physiologic medium rewires cellular metabolism and reveals uric acid as an endogenous inhibitor of UMP synthase. Cell 169, 258–272 (2017).
Tharp, K. M. et al. Adhesion-mediated mechanosignaling forces mitohormesis. Cell Metab. 33, 1322–1341 (2021).
Sullivan, M. R. et al. Quantification of microenvironmental metabolites in murine cancers reveals determinants of tumor nutrient availability. eLife 8, e44235 (2019).
Benedetti, E. et al. A multimodal atlas of tumour metabolism reveals the architecture of gene–metabolite covariation. Nat. Metab. 5, 1029–1044 (2023).
Kumar, V., Patel, S., Tcyganov, E. & Gabrilovich, D. I. The nature of myeloid-derived suppressor cells in the tumor microenvironment. Trends Immunol. 37, 208–220 (2016).
Raber, P., Ochoa, A. C. & Rodríguez, P. C. Metabolism of l-arginine by myeloid-derived suppressor cells in cancer: mechanisms of T cell suppression and therapeutic perspectives. Immunol. Invest. 41, 614–634 (2012).
Schwörer, S. et al. Proline biosynthesis is a vent for TGFβ-induced mitochondrial redox stress. EMBO J. 39, e103334 (2020).
Durante, W., Liao, L., Reyna, S. V., Peyton, K. J. & Schafer, A. I. Transforming growth factor-β1 stimulates l-arginine transport and metabolism in vascular smooth muscle cells: role in polyamine and collagen synthesis. Circulation 103, 1121–1127 (2001).
Yadav, P. et al. Reciprocal inflammatory signals establish profibrotic cross-feeding metabolism. Preprint at bioRxiv https://doi.org/10.1101/2023.09.06.556606 (2023).
Tran, D. H. et al. Mitochondrial NADP+ is essential for proline biosynthesis during cell growth. Nat. Metab. 3, 571–585 (2021).
Torrino, S. et al. Mechano-induced cell metabolism promotes microtubule glutamylation to force metastasis. Cell Metab. 33, 1342–1357 (2021).
Steggerda, S. M. et al. Inhibition of arginase by CB-1158 blocks myeloid cell-mediated immune suppression in the tumor microenvironment. J. Immunother. Cancer 5, 101 (2017).
Stekovic, S. et al. Alternate day fasting improves physiological and molecular markers of aging in healthy, non-obese humans. Cell Metab. 30, 462–476 (2019).
Argüello, R. J. et al. SCENITH: a flow cytometry-based method to functionally profile energy metabolism with single-cell resolution. Cell Metab. 32, 1063–1075 (2020).
Mak, T. W. et al. Glutathione primes T cell metabolism for inflammation. Immunity 46, 675–689 (2017).
Yarosz, E. L. & Chang, C.-H. The role of reactive oxygen species in regulating T cell-mediated immunity and disease. Immune Netw. 18, e14 (2018).
Engelhardt, J. J. et al. Marginating dendritic cells of the tumor microenvironment cross-present tumor antigens and stably engage tumor-specific T cells. Cancer Cell 21, 402–417 (2012).
Zaitsev, A. et al. Precise reconstruction of the TME using bulk RNA-seq and a machine learning algorithm trained on artificial transcriptomes. Cancer Cell 40, 879–894 (2022).
Pakshir, P. et al. Dynamic fibroblast contractions attract remote macrophages in fibrillar collagen matrix. Nat. Commun. 10, 1850 (2019).
Pickup, M. W., Mouw, J. K. & Weaver, V. M. The extracellular matrix modulates the hallmarks of cancer. EMBO Rep. 15, 1243–1253 (2014).
Ricard-Blum, S. The collagen family. Cold Spring Harb. Perspect. Biol. 3, a004978 (2011).
Papanicolaou, M. et al. Temporal profiling of the breast tumour microenvironment reveals collagen XII as a driver of metastasis. Nat. Commun. 13, 4587 (2022).
Simões, F. C. et al. Macrophages directly contribute collagen to scar formation during zebrafish heart regeneration and mouse heart repair. Nat. Commun. 11, 600 (2020).
Wishart, A. L. et al. Decellularized extracellular matrix scaffolds identify full-length collagen VI as a driver of breast cancer cell invasion in obesity and metastasis. Sci. Adv. 6, eabc3175 (2020).
Thompson, S. B. et al. Formin-like 1 mediates effector T cell trafficking to inflammatory sites to enable T cell-mediated autoimmunity. eLife 9, e58046 (2020).
Davidson, M. D., Burdick, J. A. & Wells, R. G. Engineered biomaterial platforms to study fibrosis. Adv. Healthc. Mater. 9, e1901682 (2020).
Carey, S. P., Martin, K. E. & Reinhart-King, C. A. Three-dimensional collagen matrix induces a mechanosensitive invasive epithelial phenotype. Sci. Rep. 7, 42088 (2017).
Özdemir, B. C. et al. Depletion of carcinoma-associated fibroblasts and fibrosis induces immunosuppression and accelerates pancreas cancer with reduced survival. Cancer Cell 25, 719–734 (2014).
Germano, G. et al. Role of macrophage targeting in the antitumor activity of trabectedin. Cancer Cell 23, 249–262 (2013).
Mantovani, A., Allavena, P., Marchesi, F. & Garlanda, C. Macrophages as tools and targets in cancer therapy. Nat. Rev. Drug Discov. https://doi.org/10.1038/s41573-022-00520-5 (2022).
Dröge, W. et al. Suppression of cytotoxic T lymphocyte activation by l-ornithine. J. Immunol. 134, 3379–3383 (1985).
Lercher, A. et al. Type I interferon signaling disrupts the hepatic urea cycle and alters systemic metabolism to suppress T cell function. Immunity 51, 1074–1087 (2019).
Adler, M. et al. Principles of cell circuits for tissue repair and fibrosis. iScience 23, 100841 (2020).
Nguyen-Chi, M. et al. Identification of polarized macrophage subsets in zebrafish. eLife 4, e07288 (2015).
Foster, D. S., Jones, R. E., Ransom, R. C., Longaker, M. T. & Norton, J. A. The evolving relationship of wound healing and tumor stroma. JCI Insight 3, e99911 (2018).
Timblin, G. A. et al. Mitohormesis reprogrammes macrophage metabolism to enforce tolerance. Nat. Metab. 3, 618–635 (2021).
Mills, C. D., Kincaid, K., Alt, J. M., Heilman, M. J. & Hill, A. M. M-1/M-2 macrophages and the Th1/Th2 paradigm. J. Immunol. 164, 6166–6173 (2000).
Van De Velde, L.-A. et al. T cells encountering myeloid cells programmed for amino acid-dependent immunosuppression use Rictor/mTORC2 protein for proliferative checkpoint decisions. J. Biol. Chem. 292, 15–30 (2017).
Eming, S. A., Murray, P. J. & Pearce, E. J. Metabolic orchestration of the wound healing response. Cell Metab. 33, 1726–1743 (2021).
Wouters, O. Y., Ploeger, D. T. A., van Putten, S. M. & Bank, R. A. 3,4-Dihydroxy-l-phenylalanine as a novel covalent linker of extracellular matrix proteins to polyacrylamide hydrogels with a tunable stiffness. Tissue Eng. Part C Methods 22, 91–101 (2016).
Kaukonen, R., Jacquemet, G., Hamidi, H. & Ivaska, J. Cell-derived matrices for studying cell proliferation and directional migration in a complex 3D microenvironment. Nat. Protoc. 12, 2376–2390 (2017).
Tharp, K. M. et al. Actomyosin-mediated tension orchestrates uncoupled respiration in adipose tissues. Cell Metab. 27, 602–615 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Lin, Y. C. et al. A global network of transcription factors, involving E2A, EBF1 and Foxo1, that orchestrates B cell fate. Nat. Immunol. 11, 635–643 (2010).
Eisen, M. B., Spellman, P. T., Brown, P. O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl Acad. Sci. USA 95, 14863–14868 (1998).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Cameron, A. M. et al. Inflammatory macrophage dependence on NAD+ salvage is a consequence of reactive oxygen species-mediated DNA damage. Nat. Immunol. 20, 420–432 (2019).
Kersten, K. et al. Spatiotemporal co-dependency between macrophages and exhausted CD8+ T cells in cancer. Cancer Cell 40, 624–638 (2022).
Abe, M. et al. An assay for transforming growth factor-β using cells transfected with a plasminogen activator inhibitor-1 promoter-luciferase construct. Anal. Biochem. 216, 276–284 (1994).
Acknowledgements
We thank E. Benedetti and E. Reznik for access to the Cancer Atlas of Metabolic Profiles datasets. This work was supported by 1F32CA236156-01A1, 5T32CA108462-15 and the Sandler Program for Breakthrough Biomedical Research (postdoctoral independence award) to K.M.T.; R35 CA242447-01A1, R01CA192914 and R01CA222508-01 to V.M.W.; and the National Institutes of Health Shared Instrumentation Grant S10 OD016387. Quantitative analysis of amino acids in interstitial fluid was performed by H. Shah and R. AminiTabrizi at the University of Chicago Comprehensive Cancer Center Metabolomics Platform, which receives financial support from the University of Chicago Comprehensive Cancer Center Support Grant (P30-CA014599).
Author information
Authors and Affiliations
Contributions
Conceptualization: K.M.T. and V.M.W. Methodology: K.M.T. Investigation: K.M.T., K.K., O.M., G.A.T., A.M., C.S., J.t.H., F.P.C., I.B. R.E.M. and M.-K.H. Formal analysis: K.M.T., K.K., A.M., C.S., B.S., A.J.C., J.t.H., M.-K.H. and A.J.I. Data curation: K.M.T. Funding acquisition: V.M.W. and K.M.T. Project administration: K.M.T. Software: C.S. Supervision: V.M.W. Validation: K.M.T., K.K., G.A.T. and C.S. Visualization: K.M.T. Writing—original draft: K.M.T. Writing—review and editing: K.M.T., V.M.W., K.K., M-K.H., M.P.d.M., R.G., G.A.T. and A.J.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks Costas Lyssiotis and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Tumor progression is associated with ECM synthetic myeloid programming.
a. H&E-stained histological sections of 8 week and 11 week FVB/N PyMT mammary tumors, quantification of the grade of stromal fibrosis (n = 10 or 9 mice). b. Quantification of pathological assessment for area of hyperplasia, DCIS with early invasion (DCIS) and advanced invasion (invasion) within H&E-stained histological sections of 8 week and 11 week FVB/N PyMT mammary tumors (n = 10 or 9 mice). c. Representative second-harmonic generation (SHG) images of collagen fibers in 11 week old FVB/N PyMT mammary tumors, treated with anti-CSF1 blocking antibody or IgG control weekly from 4 weeks of age until 11 weeks of age (n = 3 mice), (scale bar: 100 µm). d. Relative expression of macrophage polarization-associated gene expression of TAMs derived from 11 week old FVB/N PyMT mammary tumors, relative to TAMs derived from 8 week old PyMT mammary tumors, (n = 5 mice). e. Representative immunofluorescence microscopy of Collagen VI (white) and DNA (blue) of 11 week old FVB/N PyMT mammary tumors treated with or without LOX-inhibition, representative of the effect observed in 2 independent experiments (n = 4 mice) (scale bar: 100 µm). f. Representative immunofluorescence microscopy of F4/80 (red) and DNA (blue) in 11 week old FVB/N PyMT mammary tumors, representative localization of every tumor-stroma border assessed (n = >10 mice from > 10 independent experiments) (scale bar: 40 µm). g. Representative immunofluorescence microscopy of Collagen VI (white) and DNA (blue) of BMDMs treated with or without 5 ng/mL IL4 and 1 ng/mL TGFβ1 on fibroblast synthesized ECM surfaces for 24 h, representative of the effect observed in 3 independent experiments (scale bar: 40 µm). Data shown represent ± SEM via two-tailed unpaired Student t test (b).
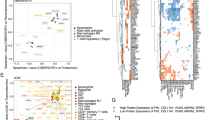
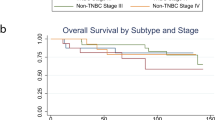
Extended Data Fig. 2 ECM-synthetic stage II and IIIA tumors are associated with poor survival.
Kaplan-Meier survival curves of 2506 stage II and IIIA breast tumors, stratified for the top and bottom quartile expression level of the genes comprising the top GO category identified in d (Cox Proportional Hazard model: p = 0.0364, z = 2.092432; LogRank: p = 0.0346).
Extended Data Fig. 3 TGFβ1 signaling and production is mechanosensitive.
a. Relative gene expression of IL4-polarized BMDMs cultured on soft (400 Pa) or stiff (60k Pa) collagen I-coated polyacrylamide hydrogel surfaces treated with 0, 0.1, 1, or 10 ng/mL TGFβ1 for 4 h, qPCR-ΔΔCT normalized to BMDMs treated without IL4 (housekeeping gene: 18 s), (n = 3 independent experiments). b. Representative immunofluorescence microscopy for RETNLA (red) and DNA (blue) of IL4-polarized BMDMs cultured on soft (400 Pa) or stiff (60k Pa) collagen I-coated polyacrylamide hydrogel surfaces, treated with 1 ng/mL TGFβ1 with or without 10 µg/mL TGFβ1-blocking antibody (1D.11) for 24 h, representative of the effect observed in 3 independent experiments (scale bar: 20 µm). c. Representative immunofluorescence microscopy for phosphorylated-SMAD2/3 (red) and DNA (blue) of IL4-polarized BMDMs cultured on soft (400 Pa) or stiff (60k Pa) collagen I-coated polyacrylamide hydrogel surfaces, treated with 1 ng/mL TGFβ1 with or without 10 µg/mL TGFβ1-blocking antibody (1D.11) for 24 h, representative of the effect observed in 3 independent experiments (scale bar: 40 µm). d. Active TGFβ1 in culture medium after 48 h of culture of IL4-polarized BMDMs cultured on soft (400 Pa) or stiff (60k Pa) collagen I-coated polyacrylamide hydrogel surfaces. (n = 4 independent experiments) e. Relative luminescent intensity of TGFβ1-reporter (PAI-1 Luc) expressing mink lung epithelial cells88 when cultured for 24 h in 1:1 conditioned media from IL4-polarized BMDMs cultured on soft (400 Pa) or stiff (60k Pa), (n = 9, 3 independent experiments of 3 technical replicates). f. Representative immunofluorescence microscopy of f4/80 (red), and DNA (blue) of stiff collagen orthotopic C57BL/6J PyMT mammary after 3 weeks of growth in Tgfbr2MyeKO or control animals, (n = 3 mice) (scale bar: 100 µm). Data shown represent ± SEM via two-tailed unpaired Student t test (d-e).
Extended Data Fig. 4 Arginine, TGFβ1, and ECM stiffness affect myeloid ECM synthesis and metabolism.
a. Relative arginine concentrations of stiff collagen orthotopic C57BL/6J PyMT mammary tumors in Tgfbr2MyeKO or control animals for 3 weeks, LC-MS analysis, (n = 7 or 6 mice). b. Representative immunofluorescence microscopy of Collagen VI (orange) and DNA (blue) of IL4-polarized WT or Arg1KO BMDMs cultured with or without 0.4 mM arginine on stiff (60k Pa) collagen I-coated polyacrylamide hydrogel surfaces treated with 1 ng/mL TGFβ1 for 24 h, representative of the effect observed in 3 independent experiments (scale bar:40 µm). c. Relative hydroxyproline (Pro-OH) concentration in BMDMs cultured on soft (400 Pa) or stiff (60k Pa) collagen I-coated polyacrylamide hydrogel surfaces with or without 1 ng/mL TGFβ1 for 24 h, measured via LC-MS, (n = 3 independent experiments). d. Fractional contributions (all isotopologues, m + 5 or m + 3) of 13C derived from 13C5-glutamine in BMDMs cultured on soft (400 Pa) or stiff (60k Pa) collagen I-coated polyacrylamide hydrogel surfaces in medium containing 12C5-glutamine, treated with 1 ng/mL TGFβ1 for 22 h, swapped for fresh medium containing 13C5-glutamine for 2 h, BMDMs were harvested and measured via LC-MS, (n = 3 independent experiments). e. Fractional contributions (all isotopologues, m + 5 or m + 3) of 13C derived from 13C5-glutamine in WT or Tgfbr2MyeKO BMDMs cultured on soft (400 Pa) or stiff (60k Pa) collagen I-coated polyacrylamide hydrogel surfaces in medium containing 12C5-glutamine, treated with or without 1 ng/mL TGFβ1 for 22 h, swapped for fresh medium containing 13C5-glutamine for 2 h, BMDMs were harvested and measured via LC-MS, (n = 3 independent experiments). f. Cellular respirometry of BMDMs cultured with or without 1 ng/mL TGFβ1 for 24 h or 100 ng/mL LPS for 1 h along with sequential additions via injection ports of Oligomycin [1 µM final], FCCP [1 µM final], and Antimycin A/Rotenone [1 µM final] during respirometry measurements, measured with an SeahorseXF24, (n = 5 wells, repeated 3 times) g-i. Relative g. NADH h. NAD i. NADH/NAD ratio in BMDMs cultured on soft (400 Pa) or stiff (60k Pa) collagen I-coated polyacrylamide hydrogel surfaces with or without 1 ng/mL TGFβ1 for 24 h, measured via LC-MS, (n = 3). Data shown represent ± SEM via two-tailed unpaired Student t test (a, d, and f) or one-way ANOVA with Tukey test for multiple comparisons (c, e, g-i) and **P < 0.01 via, ns indicates statistically not significant.
Extended Data Fig. 5 Arginine improves CTL tumor infiltration.
a. LC-MS based metabolomics of medium after 3 h of culture of Arg-ECN or ECN metabolizing a modified M9 medium, lacking ammonia, supplemented with ornithine [3 mM], (n = 3 independent experiments). b. Using the BRCA1 dataset from the Cancer Atlas of Metabolic Profiles42 we compared the levels of arginine and proline in 61 breast tumor to 47 normal adjacent tissues. c. Graphical description of the experimental setup for c-d. d. Tumor mass after 3 weeks of growth in C57BL/6J mice gavaged daily with 2 g/kg glycine, ornithine, arginine, or water, (n = 8 mice) e. Relative serum metabolites derived from retro-orbital isolated blood from mice containing PyMT tumors after 3 weeks of growth in C57BL/6J mice gavaged daily with 2 g/kg glycine, ornithine, arginine, or water (100 µL), 4 h prior to isolation of blood/serum, measured with LC-MS (n = 8 mice pooled and measured as 2 technical replicates). f. Quantitation of cleaved-caspase 3+ %-area per field view of tumor border zone of 3 week old tumors from C57BL/6J mice gavaged daily with 2 g/kg glycine, ornithine, arginine, or water (n = 5 mice). g. Quantitation of CD8+ %-area per field view from the core of 3 week old tumors from C57BL/6J mice gavaged daily with 2 g/kg glycine, ornithine, arginine, or water (n = 4 mice). h. Representative thesholded-area mask for CD8+ from the core of 3 week old tumors from C57BL/6J mice gavaged daily with 2 g/kg glycine, ornithine, arginine, or water (n = 4 mice) (scale bar: 100 µm). i. Representative immunofluorescence microscopy for CD8+ from the core of 3 week old tumors from C57BL/6J mice gavaged daily with 2 g/kg glycine, ornithine, arginine, or water (n = 4 mice) (scale bar: 100 µm). j. Representative immunofluorescence microscopy of CD8+ morphologies observed in orthotopic PyMT mammary tumors in Tgfbr2MyeKO mice gavaged daily with 2 g/kg ornithine (n = 5 mice) (scale bar: 10 µm). Data shown represent ± SEM via two-tailed unpaired Student t test (a) or one-way ANOVA with Tukey test for multiple comparisons (c-e, & f-g).
Extended Data Fig. 6 Ornithine alters CTL metabolism.
a. Heat map of relative metabolite levels of CD3/CD28-activated CD8+ CTLs cultured for 24 h in medium containing a molar ratio of 1:1 [0.5 mM], 3:1 [1.5 mM], and 9:1 [4.5 mM] ornithine:arginine, LC-MS analysis. (n = 3 independent experiments). b. Graphical description of the experimental setup and fractional contribution of 13C5-ornithine to intracellular ornithine in CD3/CD28-activated CD8+ CTLs cultured for 24 h with and without 1 mM Arginine (12C) present in medium, LC-MS analysis, (n = 3 independent experiments). c. Heat map of differentially abundant metabolite levels in b, LC-MS analysis. (n = 3 independent experiments). d. Representative immunofluorescence microscopy of OVA-PyMT tumor cells challenged with GFP+-OTI CTLs (green) for 24 h in medium containing a molar ratio of 1:1 [0.5 mM] or 3:1 [1.5 mM] ornithine:arginine, cleaved-caspase 3 (red) and DNA (blue), representative of the effect observed in 4 independent experiments (scale bar: 40 µm).
Supplementary information
Supplementary Information
Supplementary Fig. 1: flow and RNAseq gating strategy.
Supplementary Data 1
TCGA case list used.
Supplementary Tables
Supplementary Table 1: qPCR primers. Supplementary Table 2: raw/minimally processed LC–MS datasets.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tharp, K.M., Kersten, K., Maller, O. et al. Tumor-associated macrophages restrict CD8+ T cell function through collagen deposition and metabolic reprogramming of the breast cancer microenvironment. Nat Cancer 5, 1045–1062 (2024). https://doi.org/10.1038/s43018-024-00775-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-024-00775-4
This article is cited by
-
Using a pan-cancer atlas to investigate tumour associated macrophages as regulators of immunotherapy response
Nature Communications (2024)
-
Fibrotic tumors tune metabolism for immune evasion
Nature Cancer (2024)
-
Spatial proteomic profiling elucidates immune determinants of neoadjuvant chemo-immunotherapy in esophageal squamous cell carcinoma
Oncogene (2024)