Abstract
Similar to the physiological importance of gut microbiomes, recent works have shown that insect ectomicrobiotas can mediate defensive colonization resistance against fungal parasites that infect via cuticle penetration. Here we show that engineering the entomopathogenic fungus Metarhizium robertsii with a potent antibacterial moricin gene from silkworms substantially enhances the ability of the fungus to kill mosquitos, locusts, and two Drosophila species. Further use of Drosophila melanogaster as an infection model, quantitative microbiome analysis reveals that engineered strains designed to suppress insect cuticular bacteria additionally disrupt gut microbiomes. An overgrowth of harmful bacteria such as the opportunistic pathogens of Providencia species is detected that can accelerate insect death. In support, quantitative analysis of antimicrobial genes in fly fat bodies and guts indicates that topical fungal infections result in the compromise of intestinal immune responses. In addition to providing an innovative strategy for improving the potency of mycoinsecticides, our data solidify the importance of both the ecto- and endo-microbiomes in maintaining insect wellbeing.
Similar content being viewed by others
Introduction
Entomopathogenic fungi (EPF) that infect insects via cuticle penetration play critical roles in regulating insect populations1. Representative species such as the ascomycete Metarhizium anisopliae, M. acridum, and Beauveria species have been developed as ecofriendly mycoinsecticides2,3. To boost fungal virulence for cost-effective applications, different strategies have been explored by genetic engineering of EPF species4,5. For example, overexpression of endogenous protease or chitinase genes significantly increased the virulence of either Metarhizium or Beauveria to facilitate fungal penetration of insect cuticles6,7. Transgenic expression of insect peptide hormones such as the caterpillar diuretic hormone increased the virulence of B. bassiana against the wax moth Galleria mellonella by interfering with insect physiological homeostasis8. The use of insect-specific neurotoxin peptides from scorpions or spiders for gene synthesis and engineering Metarhizium could also substantially augment fungal performance by paralyzing insects before their death9,10,11. Genetic engineering of EPF expressing double-strand RNAs or miRNAs targeting insect immune defensive genes also boosted fungal virulence against different insects12,13. Alternative strategy remains to be determined to improve fungal biocontrol potency.
Insect gut microbiotas have been well demonstrated in connection with host development and health14,15. Similar to the protective roles of human skin and plant phyllosphere microbiomes16,17, recent works have indicated that insect cuticular bacteria can mediate defensive colonization resistance against fungal parasites18,19,20,21,22. To counterattack, EPF have evolved strategies such as the secretion of a defensin-like gene by Beauveria bassiana and the production of potent antibiotics by Metarhizium species to suppress host cuticular bacteria23,24. It is implicative therefore that genetic engineering of EPF with a potent antimicrobial peptide (AMP) gene may boost fungal infection performance.
In this study, we selected two antibacterial peptides for experiments, i.e., the artificially designed peptide guavanin 2 and silkworm (Bombyx mori) moricin, which have been reported with high bactericidal activities against both the Gram-positive (G+) and -negative (G−) bacteria25,26. After peptide syntheses and antibacterial assays, moricin peptide was selected for gene synthesis and transgenic transformation of M. robertsii. It was evident that the virulence of engineered strains was significantly increased against four insect species when compared with the wild-type (WT) strain. In addition to suppressing insect cuticular bacteria as designed, the engineered strains more severely disrupted insect gut microbiomes than the parental strain.
Results
Engineering Metarhizium with the moricin gene significantly increases fungal virulence against different insects
We first synthesized each mature peptide for testing their minimum inhibitory concentrations (MICs) against the G+/G− bacteria isolated from different insect cuticles. The results showed that moricin, but not guavanin 2, could efficiently suppress bacterial proliferations more or less similar to ampicillin. Both were non-active against the yeasts Saccharomyces cerevisiae and Candida albicans (Table S1).
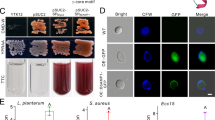
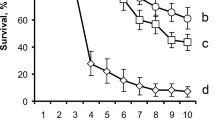
The moricin gene was then synthesized to generate the overexpression mutants (OMs) by transformation of the WT strain of Metarhizium robertsii. Two independent mutants, OM2 and OM7, were selected for further experiments based on their higher expression levels of moricin than the other mutants (Fig. S1a). Consistent with the above non-antifungal activity of moricin, there was no phenotypic difference between WT and mutants (Fig. 1a). We next performed topical fungal infections against four insect species using the WT and engineered strains. Relative to the WT strain, both OM2 (log-rank test, p = 0.0009) and OM7 (p = 0.008) killed the spotted-wing pest Drosophila suzukii significantly more quickly than did the WT strain (Fig. 1b). The estimation of the median lethal time (LT50) indicated that both OM2 (LT50 = 132 ± 4.39 h) and OM7 (LT50 = 132 ± 6.81 h) exhibited a 27% faster killing speed than the WT (LT50 = 168 ± 15.26 h). Similarly, both mutants were able to kill the mosquito Culex sinensis (a 43% increase), the desert locust Locusta migratoria (17%), and the fruit fly D. melanogaster (22%) significantly faster (p < 0.01) than the WT strain (Fig. 1c–e). No statistical difference was detected between OM2 and OM7 in killing insects.
a No phenotypic difference was observed between the WT and mutants after growing on potato dextrose agar for 2 weeks. Bar, 1 cm. b–e Overexpression of moricin in M. robertsii significantly enhanced fungal virulence against D. suzukii (b), L. migratoria (c), C. sinensis (d), and D. melanogaster (e). The plotted values are the means ± SEMs. Log-rank tests were conducted to determine the difference between the WT and individual mutants: **p < 0.01; ***p < 0.001; ****p < 0.0001; ns, not significant. Control (CK) insects were treated with 0.05% Tween 20. f Fungal load difference between treatments assayed by qPCR analysis of fungal DNA copies. D. melanogaster females were topically infected by different strains for 108 h before analysis. One-way ANOVA followed by Tukey’s test was conducted to determine the significance of difference among samples with the p values of 0.01 (capital) and 0.05 (lowercase).
Our probit estimation of the median lethal concentration (LC50) revealed that both OM2 (LC50 = 5.02 × 106 conidia/ml) and OM7 (LC50 = 5.37 × 106 conidia/ml) required fewer (>35%) spores than the WT did (LC50 = 7.26 × 106 conidia/ml) to cause 50% mortality of fruit flies. Further using D. melanogaster as a target, our quantification of fungal loads unveiled that the mutant strains colonized the flies more quickly (one-way ANOVA, p < 0.01) than the WT strain did at 108 h post-treatment (Fig. 1f). In contrast to the results of the above topical assays, both the injection assays to bypass fruit fly cuticles and topical infection of axenic flies did not result in any difference in survival rates between the WT and mutants (p > 0.05) (Fig. S1b, c).
Engineered strains exhibit a greater ability to outcompete bacteria for germination
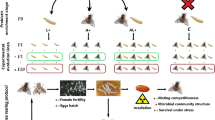
The following isolation of cuticular bacteria showed that the mosquitos were dominantly inhabited by Enterococcus bacteria; the locusts were dominated by Pseudomonas; and the spotted-wing drosophila flies were dominated by Paenibacillus and Lactiplantibacillus (Table S2). Our scanning electron microscopy (SEM) observations confirmed the presence of bacterial biofilms on the cuticles of D. suzukii adults and locusts (Fig. 2a, b), which were similar to the cuticular inhabitation of bacteria on D. melanogaster27. Taking together with the isolated bacteria from D. melanogaster27, we performed the germination of fungal spores in the absence or presence of G+ or G− bacteria isolated from different insects. Both WT and mutant spores showed similar germination rates, reaching approximately 80% after 12 h in Luria-Bertani (LB) broth (Figs. 2c, S2). When germinating fungal spores in the presence of either G+ or G− bacterial cells, however, the OM2 and OM7 strains exhibited significantly higher germination rates (p < 0.01) than the WT strain (Figs. 2d, S2). For example, in the presence of the G+ Lactiplantibacillus plantarum cells (an initial concentration at 0.05 OD600), the WT spores only germinated at a rate of 26.7% whereas both OM2 and OM7 had a similar germination rate of approximately 50% at 12 h post-inoculation. In the presence of G− bacteria such as Pseudomonas oryzihabitans from locusts or Elizabethkingia miricola from mosquitos, both OM2 and OM7 also similarly germinated better (p < 0.01) than the WT strain did (Fig. 2d). The dual overlay assays confirmed that the transgenic strains could better curb the proliferation of L. plantarum than the WT strain (Fig. S3a). The data indicated therefore that the engineered strains were better than the WT in suppressing insect cuticular bacteria.
a, b SEM images showing the presence of abundant bacterial cells on the body surfaces of D. suzukii (a) and L. migratoria (b). Two panels were randomly selected and shown for D. suzukii abdomen (left) and leg (right) parts and for different L. migratoria abdomen parts. Bar, 2 μm. c No obvious difference between the WT and transgenic strains in spore germination in LB broth without bacterial cells. d Substantial increase of the M. robertsii WT and mutant spore germinations after co-culturing for 12 h in LB broth with different bacteria. Values are the mean ± SD. One-way ANOVA followed by Tukey’s test was conducted and different letters labeled above columns show the difference with the p values of < 0.01 (capital) and < 0.05 (lowercase). Different insects are Dmel D. melanogaster; Lmig L. migratoria; Csin C. sinensis; Dsus D. suzukii. OD600 values are indicated in parenthesis for the final use of bacterial cell amount in each competing assay.
Fly ecto- and endo-microbiome dysbioses caused by fungal infections
Next, we used the females of fruit flies to isolate cuticular bacteria and count the number of bacterial colony-forming units (CFUs) after topical fungal infections. As a result, fungal treatments significantly (p < 0.01) reduced cuticular bacterial numbers compared to the mock control, and the engineered strains suppressed the proliferation of fly surface bacteria significantly more (p < 0.01) than the WT strain did (Fig. 3a). We also examined fly gut bacteria by counting CFUs after fungal infections and found that, in contrast, the gut bacterial numbers were significantly increased (p < 0.01) in the treated flies than that of mock control (Figs. 3b, S3b).
a, b Bacterial number variations on cuticles (a) and in guts (b) of flies after fungal infections. Cuticular bacteria were isolated for CFU counting from D. melanogaster females 16 h post fungal inoculations, and gut bacteria were isolated from female flies 108 h post fungal infections. Mock control insects were treated with 0.05% Tween 20. c Quantitative microbiome analysis showing the decrease of fly cuticular bacterial loads after fungal treatments. d Quantitative microbiome analysis showing the increases of fly gut bacterial loads after fungal treatments. Differential reduction in fly cuticular (e) and gut (f) bacterial diversity after fungal infections. Venn diagram analysis showing the fly cuticular (g) and gut (h) bacterial taxa divergently shared among the treatments. The cuticular bacteria isolated from D. melanogaster females after topical infection for 16 h, and gut bacteria isolated from the flies after being treated for 108 h were used for analyses. There were six replicates for each treatment. For (a, b, e, and f) one-way ANOVA followed by Tukey’s test was conducted to determine the significance of difference among samples with the p values of 0.01 (capital) and 0.05 (lowercase).
Considering that OM2 and OM7 were similar to each other in killing insects and outcompeting different bacteria, we further performed quantitative microbiome analyses using WT and OM7 strains. Treatments with WT or OM7 spores resulted in 2.3-fold and 2.8-fold reductions, respectively, in cuticular bacterial numbers compared to the mock control (Fig. 3c). In contrast, the number of fly gut bacteria increased by 40% following infection with the WT strain, while OM7 treatment caused a 5-fold increase in the number of bacteria. In particular, dominant increases of the G− Providencia bacteria were observed following WT (1.7-fold) and OM7 (7.2-fold) infections (Fig. 3d). Consistently, it was interesting to find that moricin was largely non-active against this bacterium (MIC > 128 μg/ml) by using an isolated Providencia strain for inhibition assay. Both the WT and OM7 infections substantially (p < 0.01) reduced the diversities of both cuticular and gut bacteria (Fig. 3e, f). Our Venn diagram analysis revealed that the cuticular bacterial taxa were mostly shared among treatments (Fig. 3g). In contrast, OM7 infection sharply reduced the abundance of gut bacterial species, resulting in fewer shared taxa across treatments (Fig. 3h). Overall, topical fungal infections caused the dysbioses of both the ecto- and endo-microbiomes of insects.
Fungal infections result in immune compromise of fly gut immunity
Drosophila evolved the diptericin genes DptA and DptB to mediate specific resistance against the Providencia and Acetobacter bacteria, respectively28. To corroborate the finding of a Providencia surge in fly guts, we performed a reverse transcription (RT)-quantitative PCR (RT-qPCR) analysis of different AMP genes in fruit-fly fat bodies (FBs) and guts after fungal infections. Largely consistent with the observation, the expression of DptA was not induced (p < 0.001) in the guts of the OM7-treated insects when compared to the WT strain 48 h post-infection (HPI), and its expression became similar in the WT- and OM7-treated fly guts 72 HPI (Fig. 4a). DptB expression in guts was lower (p < 0.05) in the OM7-treated insects than that of the WT-infected flies 48 HPI and then decreased to a similar level between WT and OM7 72 HPI (Fig. 4b). The antifungal drosomycin Drs and baramicin BaraA29 genes increased first but more substantially decreased (p < 0.05) in the guts after OM7 infection 72 HPI (Fig. 4c, d). In FBs, both DptA and DptB were substantially induced 48 HPI when compared to mock control and substantially downregulated 72 HPI (Fig. 4e, f). Otherwise, relative to mock controls, both Drs and BaraA were significantly upregulated (p < 0.01) in fly FBs after infections by both strains for up to 72 h. In comparison, OM7 triggered a stronger (p < 0.05) expression of Drs and BaraA than the WT strain did 72 HPI (Fig. 4g, h).
Expression of DptA (a), DptB (b), Drs (c), and BaraA (d) in fly guts after infections by the WT and OM7 strains of M. robertsii for different times. Expression of DptA (e), DptB (f), Drs (g), and BaraA (h) in fly fat bodies (FBs) after infections by the WT and OM7 strains of M. robertsii for different times. Values are the means ± SDs. There were three replicates for each sample. Pairwise two-tailed Student’s t-test was conducted between WT and OM7: ns, not significant; *p < 0.05; ***p < 0.001.
Discussion
In this study, we aimed to increase EPF performance by engineering to express an exogenous AMP gene to suppress insect cuticular defensive microbiotas. As designed, the engineering of M. robertsii with the silkworm moricin gene substantially augmented fungal virulence against different insects during topical infections. No obvious difference was observed between the parental WT and engineered strains during injection of nonaxenic flies or during topical infection of axenic flies. Unexpectedly, our quantitative microbiome data revealed that topical fungal infections sharply dysregulated fly gut microbiomes. In particular, the flies infected by the transgenic strain resulted in a more extensive surge of the pathogenic bacteria Providencia spp. in fly guts than by the WT strain, which might additionally facilitate the engineered strains to kill insects.
Guavanin 2 was designed and derived from a guava glycine-rich peptide by showing potent bactericidal activity against the G+ and G− mammalian pathogens such as Listeria ivanovii and Pseudomonas aeruginosa25. It was unexpected to find that the synthetic peptide had no obvious activity against the bacterial species isolated from insect cuticles, suggesting that this AMP might have specificity against different bacteria. Cationic moricin has a rather broad bactericidal activity30. Both the MIC tests and fungal spore germination assays confirmed its antibacterial activity against different G+/G− bacteria isolated from insects. However, we found that this AMP was non-active against the G− Providencia, which could be due to the specific lipopolysaccharides produced by this bacterial genus31. It has also been found that the Drosophila diptericin and drosocin isoforms have specific activities against alternative bacterial species28,32. We found that in contrast to the altered gut bacterial communities, fly surface bacterial taxa were largely shared after the treatments by the WT and mutant strains. Apart from the function of the bacteriostatic antibiotic helvolic acid produced and accumulated in M. robertsii spores23, the selective antibacterial activity of moricin could be a plausible reason, however, which requires further investigations.
As expected and designed, the natural infection virulence of the engineered strains OM2 and OM7 expressing moricin was significantly increased against different insects. Taken together with the suppression of insect cuticular bacterial loads by topical fungal infections, the data supported the role of insect ectomicrobiotas in defending hosts against fungal parasite infections18,19. In addition to better-suppressing insect ectomicrobiomes, it was unexpected to find that the engineered strain caused a more severe dysbiosis of the infected fly gut microbiomes than the WT strain. The axenic and nonaxenic mosquitos (Anopheles stephensi) topically infected by B. bassiana showed that the former died much slower than those nonaxenic insects, which was largely due to the overgrowth of the opportunistic pathogenic bacterium Serratia marcescens in nonaxenic guts and its translocation into insect hemolymph33. The axenic and nonaxenic bark beetle (Dendroctonus valens) larvae infected by B. bassiana resulted in a similar finding that the latter died much quicker than those axenic insects due to fungal dysregulation of gut microbiota, e.g., the overgrowth of Erwinia bacteria34. Likewise, the dysbiosis of honeybee gut microbiota could be induced by the attack of the microsporidian parasite Nosema ceranae35. It is common therefore that fungal infection of insects can cause dysbiosis of gut microbiotas that further facilitates insect death.
On top of the above understanding, it was intriguing to find that the moricin-engineered strain triggered a more extensive proliferation of bacteria in fly guts than the WT M. robertsii did, especially the overgrowth of Providencia. Providencia spp. are pathogenic to Drosophila and resistant to most host AMPs28,31. In addition to the evident non-effectiveness of moricin to Providencia, we found that the expression of DptA (being able to inhibit Providencia) was not induced in the guts of flies infected by the engineered strain, which would facilitate the surge of Providencia in the OM7-treated fly guts. Providing that EPF like M. robertsii infects and colonize insects by propagation in host body cavities36, moricin expressed by the engineered strains might not be able to enter host guts to inhibit different bacteria. Taken together, the observation would help explain why the engineered strain had a better performance than the WT in killing insects apart from the additive suppressing of cuticular bacteria. Since the engineered strains could be comparably quicker in killing locusts, mosquitos, and spotted-wing drosophila, a similar trend of bacterial overgrowth in guts might also occur in those insects infected by the WT and engineered strains, however, which requires a further investigation.
The study in Drosophila showed that fly gut immune responses against bacteria are regulated by the IMD and JAK-STAT pathways but not the Toll pathway37. Our examination of the selected AMP genes in fly guts and FBs demonstrated that the antifungal genes Drs and BaraA were also upregulated in guts after topical fungal infections, suggesting that the fungus-induced Toll pathway might also contribute to maintaining gut microbiomes. Unlike the expression pattern of antifungal genes in FBs, the antibacterial DptA and DptB were substantially induced in FBs at the early stage of fungal infection (i.e., 48 HPI) and then quickly reduced at 72 HPI. It was reported that the fungus-induced antibacterial AMPs facilitated M. rileyi to outcompete the translocated bacteria in the hemocoel of the cotton bollworm (Helicoverpa armigera) larvae38. In this respect, the early upregulations of DptA and DptB in fly FBs might involve in eliminating hemolymph bacteria. As evidenced at 72 HPI, antifungal genes were downregulated in guts but upregulated in FBs would suggest that fungal infection triggered a compromise in local (intestinal) immunity that can facilitate the dysbiosis of gut microbiomes.
In summary, we report that engineering M. robertsii with a caterpillar antibacterial gene significantly enhanced fungal potency against different insects by disrupting both the cuticular and gut microbiotas. Besides the technical advance, our data highlight that both the ecto- and endo-microbiomes are functionally essential in protecting insect hosts against fungal parasite infections.
Methods
Fungal and bacterial strains
The wild-type ARSEF 2575 and its derived mutant strains of M. robertsii36 were maintained on potato dextrose agar (PDA; BD Difco, Franklin Lakes, NJ) at 25 °C. The fungi were also cultured in Sabouraud dextrose broth (SDB; BD Difco) for different experiments. Other fungi like Saccharomyces cerevisiae AH109 and Candida albicans SC5314 were cultured in YPD (yeast extract, 10 g/l; peptone, 20 g/l, and dextrose, 20 g/l) broth for MIC assays. Bacterial species isolated from the body surface of C. sinensis, L. migratoria, and D. suzukii (Table S2) and those previously isolated from D. melanogaster27 were subjected to different assays. The isolated bacterial strains were cultured in LB broth at 37 °C or in de Man, Rogosa, and Sharpe (MRS) broth (Oxoid, Hampshire, UK) at 30 °C.
Microscopic examination of insect cuticular bacteria
We observed the microbes on the body surfaces of D. suzukii and L. migratoria using a Field-Emission Scanning Electron Microscope (Merlin Compact VP, Zeiss, Köln, Germany)39. The adults of D. suzukii 6 days post molting and the 3rd instar nymphs of L. migratoria 2 days post molting were freeze-killed and subjected to SEM analysis as described before27.
Peptide synthesis and MIC assay
The mature peptides of guavanin 2 (RQYMRQIEQALRYGYRISRR)25 and silkworm moricin (AKIPIKAIKTVGKAVGKGLRAINIASTANDVFNFLKPKKRKH)26 were synthesized by a company (Sangon Biotech, Shanghai, China) and dissolved in sterile phosphate buffer as stock solutions (128 μg/ml). MIC assays were performed as described before by serial dilutions in LB or MRS media39. The fresh bacterial cells were suspended in media at the concentration of 0.01 OD600 (~106 CFUs/ml). Equal volume (50 μl each) of different concentrations of AMPs and bacterial suspensions were mixed in 96-well plates. The plates were incubated at 30 °C in a rotary shaker at 200 rpm for 18 h to determine the MICs of moricin and guavanin 2 against each selected bacterium and yeast. The wells containing blank media were included as controls.
Peptide gene synthesis and transgenic expression in M. robertsii
Since silkworm moricin has better antibacterial activities than guavanin 2 against both G+ and G− bacteria, the former was synthesized (Sangon Biotech) by fusion with the Mcl1 mRNA 5′-untranslated region and its signal peptide that we used before10. The product was amplified with the primers TransMorF/R (Table S3), purified, and cloned into the binary vector pDHt-Bar-tef 40 for the transformation of the M. robertsii WT strain. The drug-resistant colonies were subsequently transferred to new PDA plates for single spore isolation and verification via PCR. To screen the high-expression transformants, we prepared the cDNA samples after growing the isolates in SDB for 36 h for RNA isolations. RT-qPCR was conducted with the ReverTra Ace quantitative PCR RT master mix (Toyobo, Japan). The β-tubulin gene of M. robertsii was used as a reference41. Two independent mutants (OM2 and OM7) were selected for further analysis.
Insect survival assays
Following the obtaining of the Metarhizium transgenic mutants, four insect species were then used for comparative virulence assays of the WT and OM mutants, including D. suzukii (male adults), the mosquito C. sinensis (female adults), L. migratoria (the 3rd instar nymphs) and D. melanogaster W1118 (female adults). After trial experiments, different concentrations of spore suspensions were prepared in 0.05% Tween 20 for topical infection of D. suzukii (3 × 106 conidia/ml) and D. melanogaster (5 × 106 conidia/ml) by immersion for 30 s. Spraying spore suspension was used to treat C. sinensis (3 × 107 conidia/ml) and L. migratoria (5 × 106 conidia/ml). The females of D. melanogaster were also used for injection assays (50 nl per insect; 1 × 107 conidia/ml) using a microinjector (Nanoject III; Drummond, Broomall, PA). The axenic D. melanogaster adults were also prepared for topical infections27. Thus, the serial concentrations of the WT and mutant spore suspensions, i.e., 5 × 105, 1 × 106, 5 × 106, 1 × 107, and 5 × 107 conidia/ml, were prepared for topical infection of D. melanogaster (6 days post eclosion) to determine the LC50 values of the WT and mutant strains. Except for the injection assays, the topically treated insects were maintained under a high humidity condition [relative humidity (RH) > 95%] for 16 h post treatments42. The data were analyzed by probit estimation of LC50 values. Insect survival was recorded every 12 h. At least 60 insects were used for each treatment, and the experiments were repeated twice. Representative results from one experimental batch were used.
After topical infection of D. melanogaster, the flies were collected (30 flies per replicate) at 108 h post-inoculation to determine fungal loads. The samples were homogenized in liquid nitrogen for DNA extraction using a DNeasy Blood & Tissue kit (QIAGEN). Quantitative PCR analysis was conducted using the primers for amplification of the Rpl32 gene of M. robertsii by referring to the Rpl32 gene of D. melanogaster43. There were six replicates for each sample.
Antagonistic activity assays against insect cuticular bacteria
To corroborate the virulence increase of transgenic fungus, we performed the antagonistic activity assays of the WT and mutant strains of M. robertsii in competition with different bacteria. In addition to the previously isolated cuticular bacteria from D. melanogaster27, body-surface bacteria were also isolated from the adults of D. suzukii and C. sinensis, and from the 3rd instar nymph of L. migratoria by washing with sterile phosphate buffer solution (PBS) and plating. Individual colonies were transferred for PCR amplification of the bacterial 16S rRNA genes using the universal primers 27F/1492R (Table S3). Different G+ or G− bacterial species were then randomly selected to examine the ability of the WT and mutants of M. robertsii to counteract bacterial inhibitions.
The fresh bacterial cells were collected from the overnight LB cultures, washed twice with sterile water, and re-suspended in LB. Fungal spores were collected from the 2-week-old PDA plates and suspended in 0.05% Tween 20. After trial experiments, fungal spores (at a final concentration of 5 × 106 conidia ml−1) were co-inoculated in the LB medium (10 ml) with the cells of different bacteria (Acetobacter persici, Lactiplantibacillus plantarum, Pseudomonas oryzihabitans, Paenibacillus glucanolyticus, Enterococcus thailandicus, and Elizabethkingia miricola each at a final OD600 = 0.05; Lactobacillus brevis and Glutamicibacter arilaitensis at a final OD600 = 0.01). The mixed samples were incubated in a rotary shaker at 28 °C and 200 rpm for 12 h to determine the spore germination rates39. There were six replicates for each sample. We also performed dual-culture overlay experiments using agar strips (2 × 0.5 cm each) inoculated with L. plantarum cells and transferred to a new plate (two strips per plate). Each plate was subsequently overlaid with 10 ml malt extract soft agar (BD Difco) containing the WT or mutant spores at a final concentration of 5 × 106 conidia/ml. The plates were incubated at 30 °C for 3 days to determine the size of the bacterial inhibition zone against Metarhizium growth27.
Cuticle and gut microbiome analysis
To compare the effect of moricin transgenesis on the fungal manipulation of insect ectomicrobiome structures, we used D. melanogaster females (6 days post eclosion) for topical infection with the WT and mutant spore suspensions (each containing 5 × 106 conidia/ml in 0.05% Tween 20). Treatment of flies with 0.05% Tween 20 was included as a mock control. After being maintained at an RH > 95% for 16 h, the files (10 files per sample) were anesthetized with CO2 and washed with sterile PBS (pH, 7.4). The wash buffers were individually diluted, plated on LB agar, and incubated at 37 °C for 2 days for counting the bacterial CFUs. The guts of the flies topically treated above after 108 h were dissected in sterile PBS and collected (5 guts per sample) in 1.5 ml microtubes containing sterile microbeads and 50 μl PBS. The samples were homogenized at 40 Hz for 30 s, and diluted for plating on LB agars for CFU counting.
The collected cuticular and gut bacteria were also concentrated by centrifugation at 12,000 rpm and 4 °C for 20 min for quantitative microbiome analysis27. To this end, PCR products were obtained from bacterial cells using the 16S universal primers 515F/806R (Table S3). After quality checks, the products were purified and used to generate libraries for amplicon sequencing (HiSeq 2500, Illumina, San Diego, CA) by a service company (Biozeron; Shanghai, China). Sequencing reads were normalized with the program Mothur (ver. 1.45.3)44, and clustered into individual operational taxonomic units (OTUs) with a cutoff value of 97% identity. The α-diversity of the Shannon H index was estimated based on the detected OTU taxa with the program Mothur. There were six replicates for each treatment.
Quantitative analysis of AMP gene expression
We performed the RT-qPCR analysis of different AMP genes in fruit-fly FBs and guts after fungal infection for different durations. Thus, the females of D. melanogaster were collected 6 days post molting for topical infections with the spore suspensions of WT and OM7 (5 × 106 conidia/ml each in 0.05% Tween 20). After treatment for 48 and 72 h, the flies were collected (10 each) to dissect FBs and guts for RNA extractions with TRIzol reagent (Thermo Fisher Scientific, Waltham, MA), respectively40. The flies newly treated with 0.05% Tween were used as a mock control. Four AMP genes were selected for examinations including the antifungal drosomycin Drs and baramicin BaraA29,45 and the antibacterial diptericins DptA and DptB28. There were three replicates for each sample and the experiments were repeated twice.
Statistics and reproducibility
Different statistical tests were conducted for different experiments. The log-rank test of difference in insect survival was performed with the program GraphPad Prism (ver. 10.1.1). One-way ANOVA analysis was conducted with the Turkey test using GraphPad Prism to determine the significance of differences in CFU numbers, α-diversity index, and fungal spore germinations between treatments or samples. Two-tailed Student’s t-tests were performed to determine the difference in AMP expression in fly FBs or guts between WT and OM7 after the same treatment duration. Sample sizes, animal numbers, replicate and or repeat numbers are indicated in relevant experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All sequencing data have been deposited in the NCBI SRA database under accession numbers PRJNA1002827 and PRJNA1002832 for the Drosophila cuticle and gut bacterial replicons, respectively. The data that support the findings of this study are available as Supplementary Information. Supplementary Data 1 represents the numerical source data for graphs Figs. 1b–e, 2c,d, 3, 4.
References
Hong, S., Shang, J., Sun, Y., Tang, G. & Wang, C. S. Fungal infection of insects: molecular insights and prospects. Trends Microbiol. 32, 302–316 (2024).
Jiang, Y. & Wang, J. The registration situation and use of mycopesticides in the world. J. Fungi 9, 940 (2023).
Wang, C. S. & Wang, S. B. Insect pathogenic fungi: genomics, molecular interactions, and genetic improvements. Annu. Rev. Entomol. 62, 73–90 (2017).
Lovett, B., Bilgo, E., Diabate, A. & St Leger, R. A review of progress toward field application of transgenic mosquitocidal entomopathogenic fungi. Pest Manag. Sci. 75, 2316–2324 (2019).
St Leger, R. J. & Wang, C. Genetic engineering of fungal biocontrol agents to achieve greater efficacy against insect pests. Appl. Microbiol. Biotechnol. 85, 901–907 (2010).
St Leger, R., Joshi, L., Bidochka, M. J. & Roberts, D. W. Construction of an improved mycoinsecticide overexpressing a toxic protease. Proc. Natl Acad. Sci. USA 93, 6349–6354 (1996).
Fang, W. et al. Cloning of Beauveria bassiana chitinase gene Bbchit1 and its application to improve fungal strain virulence. Appl. Environ. Microbiol. 71, 363–370 (2005).
Fan, Y., Borovsky, D., Hawkings, C., Ortiz-Urquiza, A. & Keyhani, N. O. Exploiting host molecules to augment mycoinsecticide virulence. Nat. Biotechnol. 30, 35–37 (2012).
Fang, W. et al. Development of transgenic fungi that kill human malaria parasites in mosquitoes. Science 331, 1074–1077 (2011).
Wang, C. & St Leger, R. J. A scorpion neurotoxin increases the potency of a fungal insecticide. Nat. Biotechnol. 25, 1455–1456 (2007).
Lovett, B. et al. Transgenic Metarhizium rapidly kills mosquitoes in a malaria-endemic region of Burkina Faso. Science 364, 894–897 (2019).
Cui, C. et al. Expression of mosquito miRNAs in entomopathogenic fungus induces pathogen-mediated host RNA interference and increases fungal efficacy. Cell Rep. 41, 111527 (2022).
Wang, Y. et al. Integration of dsRNA against host immune response genes augments the virulence of transgenic Metarhizium robertsii strains in insect pest species. Microb. Biotechnol. 14, 1433–1444 (2021).
Motta, E. V. S. & Moran, N. A. The honeybee microbiota and its impact on health and disease. Nat. Rev. Microbiol. 22, 122–137 (2024).
Zheng, R. et al. Holobiont perspectives on tripartite interactions among microbiota, mosquitoes, and pathogens. ISME J. 17, 1143–1152 (2023).
Harris-Tryon, T. A. & Grice, E. A. Microbiota and maintenance of skin barrier function. Science 376, 940–945 (2022).
Sohrabi, R., Paasch, B. C., Liber, J. A. & He, S. Y. Phyllosphere microbiome. Annu. Rev. Plant Biol. 74, 539–568 (2023).
Shang, J., Hong, S. & Wang, C. S. Fights on the surface prior to fungal invasion of insects. PLoS Pathog. 20, e1011994 (2024).
Hong, S., Sun, Y., Chen, H., Zhao, P. & Wang, C. S. Fungus–insect interactions beyond bilateral regimes: the importance and strategy to outcompete host ectomicrobiomes by fungal parasites. Curr. Opin. Microbiol. 74, 102336 (2023).
Janke, R. S. et al. Bacterial ectosymbionts in cuticular organs chemically protect a beetle during molting stages. ISME J. 16, 2691–2701 (2022).
Batey, S. F. D., Greco, C., Hutchings, M. I. & Wilkinson, B. Chemical warfare between fungus-growing ants and their pathogens. Curr. Opin. Chem. Biol. 59, 172–181 (2020).
Zhao, P. et al. From phyllosphere to insect cuticles: silkworms gather antifungal bacteria from mulberry leaves to battle fungal parasite attacks. Microbiome 12, 40 (2024).
Sun, Y. L., Hong, S., Chen, H. M., Yin, Y. & Wang, C. S. Production of helvolic acid in Metarhizium contributes to fungal infection of insects by bacteriostatic inhibition of the host cuticular microbiomes. Microbiol. Spectr. 10, e0262022 (2022).
Sun, Y. L., Chen, B., Li, X. L., Yin, Y. & Wang, C. S. Orchestrated biosynthesis of the secondary metabolite cocktails enables the producing fungus to combat diverse bacteria. mBio 13, e0180022 (2022).
Porto, W. F. et al. In silico optimization of a guava antimicrobial peptide enables combinatorial exploration for peptide design. Nat. Commun. 9, 1490 (2018).
Hara, S. & Yamakawa, M. Moricin, a novel type of antibacterial peptide isolated from the silkworm, Bombyx mori. J. Biol. Chem. 270, 29923–29927 (1995).
Hong, S., Sun, Y., Sun, D. & Wang, C. S. Microbiome assembly on Drosophila body surfaces benefits the flies to combat fungal infections. iScience 25, 104408 (2022).
Hanson, M. A., Grollmus, L. & Lemaitre, B. Ecology-relevant bacteria drive the evolution of host antimicrobial peptides in Drosophila. Science 381, eadg5725 (2023).
Hanson, M. A. et al. The Drosophila Baramicin polypeptide gene protects against fungal infection. PLoS Pathog. 17, e1009846 (2021).
Yi, H. Y., Chowdhury, M., Huang, Y. D. & Yu, X. Q. Insect antimicrobial peptides and their applications. Appl. Microbiol. Biotechnol. 98, 5807–5822 (2014).
Shaka, M., Arias-Rojas, A., Hrdina, A., Frahm, D. & Iatsenko, I. Lipopolysaccharide-mediated resistance to host antimicrobial peptides and hemocyte-derived reactive-oxygen species are the major Providencia alcalifaciens virulence factors in Drosophila melanogaster. PLoS Pathog. 18, e1010825 (2022).
Hanson, M. A., Kondo, S. & Lemaitre, B. Drosophila immunity: the Drosocin gene encodes two host defence peptides with pathogen-specific roles. Proc. Biol. Sci. 289, 20220773 (2022).
Wei, G. et al. Insect pathogenic fungus interacts with the gut microbiota to accelerate mosquito mortality. Proc. Natl Acad. Sci. USA 114, 5994–5999 (2017).
Xu, L. T. et al. Gut microbiota in an invasive bark beetle infected by a pathogenic fungus accelerates beetle mortality. J. Pest Sci. 92, 343–351 (2019).
Paris, L. et al. Honeybee gut microbiota dysbiosis in pesticide/parasite co-exposures is mainly induced by Nosema ceranae. J. Invertebr. Pathol. 172, 107348 (2020).
Shang, J., Song, S. & Wang, C. Metarhizium robertsii. Trends Parasitol. 40, 192–193 (2024).
Buchon, N., Broderick, N. A., Poidevin, M., Pradervand, S. & Lemaitre, B. Drosophila intestinal response to bacterial infection: activation of host defense and stem cell proliferation. Cell Host Microbe 5, 200–211 (2009).
Wang, J. L. et al. An entomopathogenic fungus exploits its host humoral antibacterial immunity to minimize bacterial competition in the hemolymph. Microbiome 11, 116 (2023).
Hong, S., Sun, Y. L., Chen, H. M. & Wang, C. S. Suppression of the insect cuticular microbiomes by a fungal defensin to facilitate parasite infection. ISME J. 17, 1–11 (2023).
Lu, M. et al. Suppression of Drosophila antifungal immunity by a parasite effector via blocking GNBP3 and GNBP-like 3, the dual receptors for β-glucans. Cell Rep. 43, 113642 (2024).
Huang, A., Lu, M., Ling, E., Li, P. & Wang, C. S. A M35 family metalloprotease is required for fungal virulence against insects by inactivating host prophenoloxidases and beyond. Virulence 11, 222–237 (2020).
Shang, J. M., Shang, Y. F., Tang, G. R. & Wang, C. S. Identification of a key G-protein coupled receptor in mediating appressorium formation and fungal virulence against insects. Sci. China Life Sci. 64, 466–477 (2021).
Shang, J. M. et al. Sensing of a spore surface protein by a Drosophila chemosensory protein induces behavioral defense against fungal parasitic infections. Curr. Biol. 33, 276–286 (2023).
Schloss, P. D. Reintroducing Mothur: 10 Years Later. Appl. Environ. Microbiol. 86, e02343–02319 (2020).
Huang, J. et al. A Toll pathway effector protects Drosophila specifically from distinct toxins secreted by a fungus or a bacterium. Proc. Natl Acad. Sci. USA 120, e2205140120 (2023).
Acknowledgements
The authors thank Zhiping Zhang and Xiaoyan Gao for helping with the scanning electron microscopy analysis. This work was supported by the National Natural Science Foundation of China (No. 32230087 and 32302430) and the National Key R&D Program of China (2022YFD1400700 and 2022YFD1400500).
Author information
Authors and Affiliations
Contributions
C.W. conceived and designed the study. S.H., H.G., and H.C. performed the experiments. S.H. and C.W. processed, analyzed, and visualized the data. S.H. and C.W. wrote the manuscript. C.W. provided supervision and received funding support.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Yongjun Zhang, Andreas Vilcinskas, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Ben Bessieres. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hong, S., Gao, H., Chen, H. et al. Engineered fungus containing a caterpillar gene kills insects rapidly by disrupting their ecto- and endo-microbiomes. Commun Biol 7, 955 (2024). https://doi.org/10.1038/s42003-024-06670-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06670-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.