Abstract
Companion planting of white clover (Trifolium repens L.) with orchard grass (Dactylis glomerata L.), a famous hay grass, improves the forage quality of orchard grass. Microbiome profiling techniques can reveal the specific role of white clover companion planting with orchard grass. This study aimed to explore the microbiome distribution and gene functions of rhizosphere and non-rhizosphere soil via companion planting systems of white clover and orchard grass. From metagenomics sequencing analysis, we confirmed the significant role of white clover on soil environment modeling during companion planting with orchard grass. Twenty-eight biomarkers of rhizosphere soil organisms were identified during companion planting, including Proteobacteria, Betaproteobacteria, Flavobacteriia, and Caulobacterales. The number of gene functions of nitrogen and carbon fixation in companion planting was higher than that in single plants, indicating new functional flora for companion planting. We characterized specific rhizosphere effects, typical biomarker flora, and potential regulatory mechanisms for white clover-related companion planting by metagenomics analyses.
Similar content being viewed by others
Introduction
White clover (Trifolium repens L.) is native to Europe and Asia1,2 and is widely used in animal husbandry owing to its highly palatable and nutritious forage for live stocks1,3,4. White clover has been widely used as a typical companion planting species to facilitate the growth of multiple plant types, including green manure, crops, and orchard grass (Dactylis glomerata L.)3. Orchard grass is a widely known hay grass type with high yields and sugar content5. As an external plant in other regions, such as Eastern United States6, this grass is the most widely used forage for animal husbandry due to its low cost and high efficiency. Companion planting of white clover and orchard grass improves the forage quality of orchard grass and is used in modern animal husbandry and agricultural applications7,8,9.
There are advantages and disadvantages to companion planting of white clover with forage feed plants such as orchard grass5,10. Among the advantages are: (1) white clover contributes to establishing a pure plant stand in a region competitively inhibiting the growth of other weeds except for the dominant forage feed plants; (2) white clover is a general plant with effective nitrogen-fixing capacity which can provide nitrogen for grass growth near their roots; (3) white clover can tolerate close grazing, reducing the planting and management costs10. Among the disadvantages are: (1) white clover is not durable in high-traffic areas; (2) white clove-dependent plant stand purring is slow. Sometimes, white clover requires reseeding every few years to help purify the line of forage plants10. Although there are multiple advantages to companion planting, the biological foundations of the improved efficiency have not been fully revealed. The nitrogen-fixing capacity of white clover may be a major mechanism, indicating that pathways related to nitrogen fixation, together with its microenvironment and root-related environment (rhizosphere soil microenvironment), may be crucial for reaping the companion planting effects of white clover.
Microorganisms in the soil are crucial for the growth and development of plants. They play an important role in soil ecology and regeneration11. Using the principle of allelopathy between plants and arranging matching plants or intercropping reasonably not only improves vegetable yield and quality but effectively reduces pests and diseases, improves the soil ecological environment, and reduces recurrent cropping obstacles12,13. Intercropping of Cowpea/sorghum and peanut/corn activates soil phosphorus and increases soil phosphatase activity14,15. The root exudates of accompanying plants further promote the growth of seedlings and plants by inhibiting the growth of pathogenic microorganisms and regulating the soil’s microecological environment16.
White clover and orchard grass are two typical and traditional model plants for companion planting17. However, the detailed mechanisms of the interactions between white clover and orchard grass are unclear. In this study, we applied soil microbiome analyses to reveal the microbiome profiling around white clover, orchard grass, and companion planting roots. Furthermore, it aimed to identify key microorganisms, which were contributed to nitrogen fixation or other companion planting-dependent growth efficiency associated biological pathways. We identified key regulators involved in these biological processes and demonstrated the potential biological regulatory mechanisms and foundations at the metagenomics level.
Materials and methods
Experimental site and treatments
Experiments were performed in October 2019 at the Dayangdian Experimental Station of Anhui Agricultural University (31° 58′ N, 117° 24′ E), Hefei City, Anhui Province, Southeast China. The study involved three groups: white Clover (Trifolium repens L.) alone, orchard grass (Dactylis glomerata L.) alone, and their companion planting. The land area used in the experiment measured 4 × 4 m2 in each plot, and the plants were seeded in a row. Specifically, 10 and 20 kg ha−1 of white clover and orchard grass were sown, respectively; in companion planting, 7.5 kg ha−1 white clover and 5 kg ha−1 orchard grass were sown. Climate, soil physicochemical characteristics, and field management were according to the protocol described by previous research5.
Sample collecting
Eighteen samples from six different soil regions around white clover, orchard grass, or the companion planting groups were collected for analyses, with three replicates each. For comparison, samples were classified into non-rhizosphere or rhizosphere groups. The plant was uprooted, and the rhizosphere soil samples were carefully collected using a sterile brush, marked as WC.R (rhizosphere soil of white clover), OG.R (rhizosphere soil of orchard grass), and MIX.R (rhizosphere soil of mixed plant). The non-rhizosphere soil samples were collected from the between rows at a depth of 10 cm, and marked WC.T (non-rhizosphere soil of white clover), OG.T (non-rhizosphere soil of orchard grass), and MIX.T (non-rhizosphere soil of mixed plant). Samples were collected under the same conditions following the same method to be consistent. All samples were collected and placed in liquid nitrogen immediately for quick freezing. The samples were stored at − 80 ℃ in the lab for further sequencing.
Metagenomics sequencing
Metagenomics sequencing was performed to recognize the microorganism profiling in the rhizosphere and non-rhizosphere soil environments.
DNA extraction
Total DNA was extracted from the soil contents using the DNeasy PowerSoil kit (Qiagen, Hilden, Germany) following the manufacturer’s instructions. The concentration and integrity of DNA were measured on a NanoDrop2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA) and by agarose gel electrophoresis, respectively.
Fragmenting DNA
In brief, the gDNA sample (1 µg) was sheared by transferring 52.5 µL of each DNA sample to the corresponding Covaris tube. Fragmented DNA (50 µL) was transferred from each Covaris tube to the corresponding well of the new 0.3 mL PCR plate.
Clean up for fragmented DNA
In brief, 80 µL well-mixed AMPure XP (A63881, BECKMAN COULTER, CA, USA) beads were added to each well. The entire volume was gently piped up and down at least ten times to mix thoroughly, and the plate was incubated at room temperature for 5 min. The PCR plate was placed on the magnetic stand at room temperature until the liquid appeared clear. All of the supernatant was removed and discarded from each well. With the plate on the magnetic stand, 200 µL of freshly prepared 80% EtOH was added to each well containing the sample without disturbing the beads. The plate was incubated at room temperature for 30 s, and subsequently, the supernatant was removed and discarded from each well. The plate was kept at room temperature for 5 min to dry, and the plate was removed from the magnetic stand. The dried pellet was resuspended in each well with 62.5 µL of resuspension buffer. The entire volume was gently pipetted up and down ten times to mix thoroughly. The plate was incubated at room temperature for 2 min. The plate was placed on the magnetic stand at room temperature until the liquid appeared clear. The clear supernatant (60 µL) is transferred from each well to the corresponding well of a new 0.2 mL PCR plate.
Repair ends and select Library size
ERP2 or ERP3 (40 µL, A63881, BECKMAN COULTER, CA, USA) was added to each well. The entire volume was gently pipetted up and down to mix thoroughly. The sealed plate was placed on the pre-heated thermal cycler and incubated at 30 °C for 30 min.
Adenylate 3′ ends
ATL (12.5 µL, A63881, BECKMAN COULTER, CA, USA) was thawed and added to each well of the ALP plate. The entire volume was gently pipetted up and down ten times to mix thoroughly. The sealed plate was placed on the pre-heated thermal cycler and incubated at 37 °C for 30 min and 70 °C for 5 min. The sealed plate was placed on the pre-heated thermal cycler and incubated at 30 °C for 10 min. STL (5 µL) was added to each well of the plate to inactivate the ligation reaction. The entire volume was gently pipetted up and down ten times to mix thoroughly.
Ligate adapters
RSB (2.5 µL, A63881, BECKMAN COULTER, CA, USA) was added to each well of the plate. LIG2 (2.5 µL) and the DNA Adapter Index (2.5 µL) were successively added to each well of the plate. The entire volume was gently pipetted up and down ten times to mix thoroughly.
Enriching DNA fragments
The thermal cycler (C1000 Touch, Bio-rad, Hercules, CA, USA) was pre-programmed as follows: the pre-heat lid option was chosen and set to 100 °C; 95 °C for 3 min; 8 cycles at 98 °C 20 s; 60 °C 15 s; 72 °C 30 s; 72 °C for 5 min, and hold at 4 °C. PPC (5 µL) was added per well in the PCR plate. EPM (20 µL) was added to each well of the PCR plate. The sealed PCR plate was placed in the pre-programmed thermal cycler.
Data preprocessing
The quality of raw reads was assessed using the FASTQC software (version 0.11.9)18 (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Adapter sequences and low-quality sequences were trimmed using Trimmomatic (version 0.33)19 with the following parameters: LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 HEADCROP:2 MINLEN:36. The taxonomic assignment of the treated reads was performed using kraken2 software20 (v2.0.8-beta) (https://github.com/BenLangmead/aws-indexes?tab=readme-ov-file) and Bracken21. The linear discriminant analysis effect size (LEfSe) method was used to identify microorganisms with significant differences across different groups. First, the difference function between different groups was detected through the rank sum test, and linear discriminant analysis (LDA) was used to reduce dimensionality and evaluate the effect of the difference function, resulting in the calculation of the LDA score. LEfSe multi-level species difference analysis was used to identify significant biomarker flora. Biological biomarkers of the rhizosphere and non-rhizosphere of three cultivated plants at different classification levels reached as high as 745 with an LDA threshold of 2. The alpha diversity analysis was conducted using the vegan package in R22. Principal Co-ordinate Analysis (PCoA) was conducted following a previously described protocol23.
Metagenome assembly and gene function analyses
Reads from all samples were co-assembled with MEGAHIT embedded in the metaWRAP Assembly24. Genes were predicted using MetaGeneMark and MetaEuk. Gene functions were annotated using eggNOG (v5.0)25, containing information such as GO, KEGG26, KOG, and CAZy terms. The kraken2 software (v2.0.8-beta) (https://github.com/BenLangmead/aws-indexes?tab=readme-ov-file) was used to assign taxonomic annotations to each sample with self-built data base with archaea (964), bacteria (172,595), fungi (300), humans, protozoa (94) and viruses (9362) from NCBI. The genome, plasmid sequence (3137), and net library (nucleic acid sequence database used by NCBI BLAST) were compared and analyzed, and bracken combined with the taxonomic information database from NCBI was used to obtain the species annotation and abundance. Statistical analyses on the sample abundance were performed at different taxonomic levels: Domain, Kingdom, Phylum, Class, Order, Family, Genus, and Species. Statistical values for each sample and abundance were used to construct the abundance spectrum for the corresponding taxonomic level.
Results
Clean metagenomics sequencing data for microbiome profiling
We performed shotgun metagenomic sequencing analysis for 18 samples, yielding 402 390 072 sequence reads with a length of 150 bp. Low-quality bases were removed. More than 99.99% of the sequences were retained and the total data volume was 261 GB. The data sequence volume after cleaning was sufficient for subsequent microbial species and gene function analyses. The detailed results before and after preprocessing are shown in Table S1.
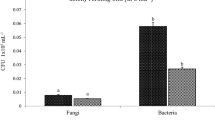
Half of the sequences could not be aligned with the existing known species sequences, as shown in Table S2. Specifically, 41–52% of the sequences were microbial. Among the microorganisms identified by comparison, the vast majority were bacteria, with an average content of more than 96%, followed by fungi, with a content of 2.92–3.13%; protozoa and viruses were very low (Table S2, Figs. S1, S2). The microbiome assembly information is described in Table S3.
Higher alpha diversity for non-rhizosphere compared with rhizosphere samples
The coverage rate of each sample library reached almost 100%, indicating that the sequencing result could represent all organisms of the sample. No significant differences among groups were observed in alpha diversity parameters, including the comprehensive richness index, Chao1 index, ACE index, and Simpson index. Compared with rhizosphere samples, non-rhizosphere samples showed a relatively higher alpha diversity. However, no significant differences were identified between rhizosphere samples around different plant groups. Therefore, the distribution of soil microorganisms correlated significantly with the matched plant parts and root-related distances (Table 1).
Alteration in the microbial community structure evaluated by ecological similarity analysis reveals white clover as a dominant regulator
PCoA is a visualization method to assess the similarity or difference in data, which can reveal the most important coordinates in the matrix. PCoA can show the differences between individuals or groups. At the genus level, we found a clear difference between the microbial composition of the rhizosphere and non-rhizosphere regions. As for the rhizosphere microbiome, the microbial composition of MIX samples was similar to white clover samples and orchard grass samples, with the same trend as the non-rhizosphere microbiome. However, the non-rhizosphere microbial composition of MIX samples was more similar to white clover samples (Figs. 1 and 2). The rhizosphere microbial composition was affected by companion planting but the non-rhizosphere microbial composition was more affected by white clover than orchard grass samples.
PCoA analyses at genus level comparing different groups. Diverse genus distributions were observed in samples with different locations (rhizosphere and non-rhizosphere) and with/without companion planting. WC.R, OG.R, and MIX.R mean the rhizosphere soil samples of white clover, orchard grass, and MIX, respectively. WC.T, OG.T, and MIX.T mean the non-rhizosphere soil samples of white clover, orchard grass, and MIX, respectively.
PCoA analyses at species level comparing different groups. Diverse species distributions were observed in samples with different locations (rhizosphere and non-rhizosphere) and with/without companion planting. WC.R, OG.R, and MIX.R mean the rhizosphere soil samples of white clover, orchard grass, and MIX, respectively. WC.T, OG.T, and MIX.T mean the non-rhizosphere soil samples of white clover, orchard grass, and MIX, respectively.
Significant biomarker flora identified by comparing experimental groups
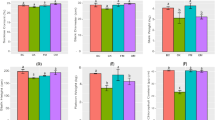
At the phylum level (Fig. 3A), compared with the non-rhizosphere soil samples, the rhizosphere soil samples showed higher relative abundances of Proteobacteria and Gemmatimonadetes and lower relative abundances of Actinobacteria and Nitrospirae. At the genus level (Fig. 3B), compared with the non-rhizosphere soil samples, the rhizosphere soil samples had higher relative abundance of Gemmatirosa, Rhizobacter, Variovorax, and Sphingomonas, and lower relative abundances of Nitrospira, Lysobacter, Solirubrobacter, Gaiella, and Phycicoccus. The differences among microbial communities of several cultivated groups are shown (Fig. S3). To identify the biomarker with large differences, the LDA threshold was set at 3. Cultivated plants under the three conditions showed 91 significant biomarkers between the rhizosphere and non-rhizosphere samples at different taxonomic levels (Table S2). The biomarkers of rhizosphere samples for orchard grass included Verrucomicrobia at the phylum level, Oxalobacteraceae at the class level, Opitutales at the order level, Opitutae at the family level, Polaromonas, Janthinobacterium, and Ramlibacter at the genus level, and Ramlibacter tataouinensis at the species level (Fig. S4). The biomarkers of rhizosphere samples for white clover included Bacteroidetes at the phylum level, Sphingomonadales at the order level, Erythrobacteraceae and Sphingomonadaceae at the family level, and Altererythrobacter, Pedobacter, Novosphingobium, Stenotrophomonas and Sphingomonas at the genus level. There were 28 biomarkers of rhizosphere organisms in the companion planting group, including Proteobacteria, Betaproteobacteria and Flavobacteriia, Caulobacterales, Yellow Flavobacteriales, Micrococcales, Pseudomonadales and Burkholderiales, Intrasporangiaceae, Micrococcaceae, Nocardia Nocardioidaceae, Microbacteriaceae, Flavobacteriaceae, Caulobacteraceae, Pseudomonadaceae and Comamonadaceae, Rhodobacter Genus (Rhodoferax), Microbacterium (Microbacterium), Sphingopyxis, Arthrobacter, Nocardioides, Flavobacterium, Massilia, Variovorax, and Pseudomonas, Pseudomonas fluorescens, Mucilaginibacter ginsenosidivorax, and Variovorax paradoxus. Archaea and fungi in the rhizosphere soil of the companion planting group were less than those in the rhizosphere soil of a single plant, and those in the rhizosphere soil were significantly less compared with the non-rhizosphere soil (Fig. S5). Caulobacteraceae belonging to the class of α-Proteobacteria, Rhodoferax belonging to the class of β-Proteobacteria, Rhodoferax, Massilia, Variovorax paradoxus, and Pseudomonas fluorescens (Pseudomonas fluorescens) of the γ-proteobacteria in the rhizosphere soil of the companion planting group were significantly higher compared with the rhizosphere soil of single cultivated plants.
The microbiota community composition of rhizosphere soil and non-rhizosphere soil samples at different levels. WC.R, OG.R, and MIX.R mean the rhizosphere soil samples of white clover, orchard grass, and MIX, respectively. WC.T, OG.T, and MIX.T mean the non-rhizosphere soil samples of white clover, orchard grass, and MIX, respectively.
Nitrogen and carbon fixation-related microbes enriched by white clover-mediated companion planting effects
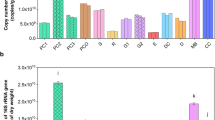
Nitrogen and carbon fixation-related pathways were differentially enriched, as evidenced by differential gene functions in microorganisms in the comparison between single and companion planting groups. The number of rhizosphere microbial carbon fixation pathway genes in the companion planting groups was significantly higher than that of a single plant, as shown in Fig. 4. The top 10 terms of the carbon fixation pathway accounted for more than 60% of the total number of genes in the carbon fixation pathway. Among them, more than 10% was due to Burkholderiales and Rhizobiales (Table S4). Similar to the carbon fixation pathway, the number of nitrogen metabolism pathway genes in the rhizosphere microbial nitrogen metabolism pathway of white clover cultivated alone was slightly higher than that of orchard grass. In contrast, the number of genes in the rhizosphere microbial nitrogen metabolism pathway of the companion planting group was slightly higher. The number of genes for carbon fixation pathways and nitrogen metabolism in MIX was higher than in the single planting of white clover or orchard grass (Fig. 5). The number of genes among the top 10 terms related to the nitrogen metabolism pathway accounted for nearly 60% of the total nitrogen metabolism pathways, of which more than 10% were due to Burkholderiales and Rhizobiales. The highest contribution of the genus of first-level microorganisms was by soybean rhizobia (Bradyrhizobium); the number of genes in this genus affecting the carbon fixation pathway was 4.44%.
KEGG pathways enrichment results for rhizosphere soil microbiome biomarker: WC.R, OG.R, and MIX.R mean the rhizosphere soil samples of white clover, orchard grass, and MIX, respectively. WC.T, OG.T, and MIX.T mean the non-rhizosphere soil samples of white clover, orchard grass, and MIX, respectively.
Gene enrichment number for significant carbon-fixation and nitrogen metabolism pathways: (A) carbon fixation pathways; (B) nitrogen metabolism pathways. WC.R, OG.R, and MIX.R mean the rhizosphere soil samples of white clover, orchard grass, and MIX, respectively. WC.T, OG.T, and MIX.T mean the non-rhizosphere soil samples of white clover, orchard grass, and MIX, respectively. The analytic workflow, significance threshold selection and reference database are described in the “Materials and methods” Section. In part (A) or (B), noting with different lowercase letters means significant difference (P < 0.05).
Discussion
Different alpha diversity indexed across sampling environments
Although the non-rhizosphere sample had relatively higher alpha diversity than the rhizosphere samples, there were no significant differences in alpha diversity indexes between soil samples around different plants. Consistently, plants, especially those with nitrogen-fixing capacity, may shape the microbiome around their roots27,28,29, resulting in altered microbiome diversity between rhizosphere and non-rhizosphere regions. Similarities observed from the comparison between plants may indicate that, at least for white clover, companion planting facilitates the dominant plant’s growth, and this is not by the regulation of microbiome diversity.
Alteration in microbial community structure patterns with or without companion planting assessed based on ecological similarity
Alpha diversity analysis showed that microbial community structure differences across sample groups were common. As shown in Figs. 1 and 2, the microbiome diversity away from the root (non-rhizosphere regions) tends to be higher at both genus and species levels, closer to uncontrolled randomized distributions. In the rhizosphere regions, the similarities between white clover alone and companion planting are higher than those between orchard grass only and companion planting. Such similarity indicates that white clover dominates the soil microbiome alteration and modification in companion planting conditions and, therefore, contributes more to the effects of companion planting.
Identification of significant biomarker flora across experimental groups
We integrated the discussion on microbial composition abundance and biomarker flora to provide a comprehensive overview of the selection of companion planting-associated microbes. Multiple nitrogen fixation-associated classes, orders, genera, or species were identified, indicating the specific role of nitrogen’s effects on companion plants, in line with previous reports30,31. For a detailed discussion, we identified the typical rhizosphere microbiome for orchard grass and white clover and companion planting separately, including Caulobacteraceae, Rhodoferax, Massilia, and Variovorax paradoxus and Pseudomonas fluorescens. These microbes are associated with nitrogen fixation-related biological processes32,33,34,35, indicating that nitrogen fixation may be a key regulatory effect at the biochemical level mediating the effects of companion planting with white clover.
Functional interpretation for white clover-mediated effects of companion planting
Based on the initial analyses of the three groups, differential biological processes were observed between companion planting and single planting groups in rhizosphere samples.
Specifically, we focused on the carbon fixation- and nitrogen fixation-associated pathways for further analyses. A higher number of gene functions related to carbon fixation and nitrogen metabolism pathways was observed, indicating the specific role of such two pathways in the rhizosphere and the improved effects of companion planting. According to recent studies, functionally, carbon fixation and nitrogen metabolism are associated with the production and growth status of planting5, which is consistent with our results. Both these metabolism pathways are associated with companion planting with white clover, confirming the specific contribution of white clover in the activation of these pathways.
In this study, we performed a comprehensive analysis of the rhizosphere and non-rhizosphere soil microbiome of white clover only, orchard grass only, and companion planting. The comparison between rhizosphere and non-rhizosphere samples revealed that the effects of companion planting showed a differential efficiency in rhizosphere or non-rhizosphere soil. Functional exploration and biomarker flora identification confirmed the contribution of nitrogen fixation on companion planting effects with white clover. We verified the rhizosphere effects and recognized key biomarker flora and related biological processes of companion planting with white clover.
Data availability
The data that support the findings of this study have been deposited https://db.cngb.org/search/project/CNP0004951/.
References
Chestnutt, D. & Lowe, J. White clover/grass relationships: agronomy of white clover/grass swards: A review. In Occasional Symposium of the British Grassland Society, pp. 191–213 (1970).
Jahufer, M., Cooper, M., Ayres, J. & Bray, R. Identification of research to improve the efficiency of breeding strategies for white clover in Australia—a review. Aust. J. Agric. Res.53, 239–257 (2002).
Davies, W. E. White clover breeding: A review. 99–122 (1970).
Brock, J., Caradus, J. & Hay, M. Fifty years of white clover research in New Zealand. In Proceedings of the New Zealand Grassland Association, 25–39 (1989).
Chen, L. et al. Comparative Analysis of soil microbiome profiles in the companion planting of white clover and orchard grass using 16S rRNA gene Sequencing data. Front. Plant Sci.11, 1431 (2020).
Gold, A. L. Habitat use of black bears in the northeast cascades of Washington. Biol. Conserv.113(1), 55–62 (1997).
Maldonado Peralta, M. D. L. Productivity of orchard grass (Dactylis glomerata L.) alone and associated with perennial ryegrass (Lolium perenne L.) and white clover (Trifolium repens L). Revista Brasileira De Zootecnia46, 890–895 (2017).
Rojas García, A. R. et al. Tiller population in orchard grass (Dactylis glomerata L.) grown alone and associated with perennial ryegrass (Lolium perenne L.) and white clover (Trifolium repens L). Revista mexicana de ciencias Pecuarias8, 419–428 (2017).
Seguin, P., Pomerleau-Lacasse, F., Drapeau, R., Lajeunesse, J. & St‐Pierre‐Lepage, S. Productivity and persistence of kura clover and white clover mixtures with grasses. Agron. J.112, 2891–2904 (2020).
Heshmati, S. Effect of white clover and perennial ryegrass genotype on yield and forage quality of grass-clover and grass-clover-forb mixtures. Georg-August-Universität Göttingen (2018).
Liu, Z. et al. Nitrogen fertilizers affect microbial hitchhiking to the plant roots. J. Agric. Food Chem.72(9), 4639–4648 (2024).
Fu, X., Li, C., Zhou, X., Liu, S. & Wu, F. Physiological response and sulfur metabolism of the V. dahliae-infected tomato plants in tomato/potato onion companion cropping. Sci. Rep.6, 36445 (2016).
Gao, D. et al. Wheat cover crop promoted cucumber seedling growth through regulating soil nutrient resources or soil microbial communities?. Plant. Soil.418(1–2), 1–17 (2017).
Makoi, J. H. J. R., Chimphango, S. B. M. & Dakora, F. D. Elevated levels of acid and alkaline phosphatase activity in roots and rhizosphere of cowpea (Vigna unguiculata L. Walp.) genotypes grown in mixed culture and at different densities with sorghum (Sorghum bicolor L). Crop Pasture Sci.61(4), 279–286 (2010).
Kirk, J. L. et al. Methods of studying soil microbial diversity. J. Microbiol. Methods58(2), 169–188 (2004).
Dai, C. C. et al. Effects of intercropping of peanut with the medicinal plant atractylodes lancea on soil microecology and peanut yield in subtropical China. Agrofor. Syst. (2013).
Davis, W. E. P. Effects of Using oats as a Companion crop with Orchard Grass, Dactylis glomerata L. and white clover, Trifolium Repens L. sown for pasture. Can. J. Plant Sci.42(4), 582–588 (1962).
Andrews, S. FastQC: A quality control tool for high throughput sequence data. (2017). (2010).
Bolger, A., Giorgi, F. & Trimmomatic,. A flexible read trimming tool for illumina NGS data. Bioinformatics30, 2114–2120 (2014).
Garcia, B. J. et al. Kraken2 metagenomic virus database. Oak Ridge National Lab. (ORNL), Oak Ridge, TN (United States) (2020).
Lu, J., Breitwieser, F. P., Thielen, P. & Salzberg, S. L. Bracken: Estimating species abundance in metagenomics data. PeerJ Comput. Sci.3, e104 (2017).
Oksanen, J. et al. Package ‘vegan’. Community ecology package, version 2, 1–295 (2013).
Avershina, E., Frisli, T. & Rudi, K. De novo semi-alignment of 16S rRNA gene sequences for deep phylogenetic characterization of next generation sequencing data. Microbes Environ.28(2), 211–216 (2013).
Li, D., Liu, C. M., Luo, R., Sadakane, K. & Lam, T. W. MEGAHIT: An ultra-fast single-node solution for large and complex metagenomics assembly via succinct de bruijn graph. Bioinformatics31, 1674–1676 (2015).
Huerta-Cepas, J. et al. eggNOG 5.0: A hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res.47, D309–D314 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res.51(D1), D587–D592 (2023).
Da Silva Fonseca, E. et al. The microbiome of eucalyptus roots under different management conditions and its potential for biological nitrogen fixation. Microb. Ecol.75, 183–191 (2018).
Stanton, D. E., Batterman, S. A., Von Fischer, J. C. & Hedin, L. O. Rapid nitrogen fixation by canopy microbiome in tropical forest determined by both phosphorus and molybdenum. Ecology100, e02795 (2019).
Mahmud, K., Makaju, S., Ibrahim, R. & Missaoui, A. Current progress in nitrogen fixing plants and microbiome research. Plants9, 97 (2020).
Fustec, J., Lesuffleur, F., Mahieu, S. & Cliquet, J. B. Nitrogen rhizodeposition of legumes. A review. Agron. Sustain. Dev.30, 57–66 (2010).
Mutiga, S. K., Gohole, L. S. & Auma, E. O. Effects of integrating companion cropping and nitrogen application on the performance and infestation of collards by Brevicoryne brassicae. Entomol. Exp. Appl.134, 234–244 (2010).
De La Fuente, L. et al. Inoculation with pseudomonas fluorescens biocontrol strains does not affect the symbiosis between rhizobia and forage legumes. Soil Biol. Biochem.34, 545–548 (2002).
Guenet, B., Leloup, J., Hartmann, C., Barot, S. & Abbadie, L. A new protocol for an artificial soil to analyse soil microbiological processes. Appl. Soil. Ecol.48, 243–246 (2011).
Nacke, H. et al. Pyrosequencing-based Assessment of bacterial community structure along different management types in German forest and grassland soils. PloS One6, e17000 (2011).
Inokuchi, R., Kuma, K. I., Miyata, T. & Okada, M. Nitrogen-assimilating enzymes in land plants and algae: Phylogenic and physiological perspectives. Physiol. Plant.116, 1–11 (2002).
Funding
This work was supported by grants from the National Natural Science Foundation of China (32172769, 31872418).
Author information
Authors and Affiliations
Contributions
LC and YZ designed the experiments. DL and YW performed the experiments and collected samples. DL, YW, XL, ZZ and GW analyzed the data. DL, LC, and YZ wrote and revised the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, D., Wang, Y., Li, X. et al. Exploring microbial diversity and function in companion planting systems of white clover and orchard grass. Sci Rep 14, 21609 (2024). https://doi.org/10.1038/s41598-024-72705-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-72705-4
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.