Abstract
Thirty-five 9-O-berberrubine carboxylate derivatives were synthesized and evaluated for yeast α-glucosidase inhibitory activity. All compounds demonstrated better inhibitory activities than the parent compounds berberine (BBR) and berberrubine (BBRB), and a positive control, acarbose. The structure–activity correlation study indicated that most of the substituents on the benzoate moiety such as methoxy, hydroxy, methylenedioxy, benzyloxy, halogen, trifluoromethyl, nitro and alkyl can contribute to the activities except multi-methoxy, fluoro and cyano. In addition, replacing benzoate with naphthoate, cinnamate, piperate or diphenylacetate also led to an increase in inhibitory activities except with phenyl acetate. 9, 26, 27, 28 and 33 exhibited the most potent α-glucosidase inhibitory activities with the IC50 values in the range of 1.61–2.67 μM. Kinetic study revealed that 9, 26, 28 and 33 interacted with the enzyme via competitive mode. These four compounds were also proved to be not cytotoxic at their IC50 values. The competitive inhibition mechanism of these four compounds against yeast α-glucosidase was investigated using molecular docking and molecular dynamics simulations. The binding free energy calculations suggest that 26 exhibited the strongest binding affinity, and its binding stability is supported by hydrophobic interactions with D68, F157, F158 and F177. Therefore, 9, 26, 28 and 33 would be promising candidates for further studies of antidiabetic activity.
Similar content being viewed by others
Introduction
Diabetes mellitus is a chronic metabolic disorder characterized by uncontrolled hyperglycemia due to inadequate production or ineffective use of insulin in the body which can result in many consequences such as neuropathy, nephropathy, stroke and cardiovascular disease1. In 2021, there are approximately 537 million adults diagnosed to have diabetes in the world and the number is predicted to reach 783 million by 20452. In addition, 6.7 million death due to diabetes was recorded and at least 966 billion dollars were expensed for treatment of diabetes2. Type 2 diabetes, the repercussion of excess body weight and physical inactivity, accounts for more than 95% of the people living with diabetes. This type of diabetes was diagnosed only in adults but now it tends to appear frequently in children3. A potential therapeutic approach for diabetes mellitus, particularly in type 2 diabetes, is to decline postprandial hyperglycemia by using α-glucosidase inhibitors to prevent carbohydrate digestion. α-Glucosidase located in the brush-border surface membrane of intestinal cells, is an essential hydrolytic enzyme in the carbohydrates digestion process, it degrades oligosaccharide to monosaccharide4. The production of these absorbable glucose results in postprandial hyperglycemia in patients with diabetes. Some α-glucosidase inhibitors such as acarbose, miglitol and voglibose, which are carbohydrate mimetics, have been used in the clinic for the treatment of type 2 diabetes. Nevertheless, because of their side effects including flatulence, diarrhea, stomach ache and liver damage, it is necessary to develop new, efficient and benign α-glucosidase inhibitors for the treatment of diabetes mellitus5. Recently, there have been several research about α-glucosidase inhibitors6,7,8,9.
Berberine (BBR), a quaternary ammonium salt from the protoberberine group of isoquinoline plant alkaloids, is used for medicinal purposes in 3000 years-long histories of the Indian and Chinese. Berberine was found as an active component in the root, rhizome and stem bark of many well-known medicinal plants such as Hydrastis canadensis (goldenseal), Coptis chinensis (Coptis or goldenthread), Berberis aquifolium (Oregongrape), Berberis vulgaris (barberry) and Berberis aristata (Treeturmeric)10. Berberine possesses a wide range of biological activities such as antimicrobial11, anti-Alzheimer12, antidiabetic13, antihypertensive14, anticancer15, and anti-inflammatory activity16. Nevertheless, efficient applications of berberine are hindered by its poor bioavailability which is less than 5%17,18. It is attributed to its poor aqueous solubility (~ 1.8 mg/mL at 20 °C) since berberine has a temperature-dependent aqueous solubility which increases with an increase in temperature19. In addition, berberine is a hydrophilic compound with a log p value of − 1.5 which makes berberine lipophobic with limited membrane permeability, and this leads to low gastrointestinal absorption20. Therefore, to improve the bioavailability of berberine, its new derivatives have to be designed and synthesized with their enhanced aqueous solubility and permeability through intestinal membranes.
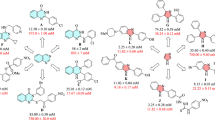
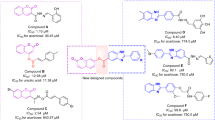
There have been several reports about the antidiabetic activity of berberine. It was proved that the antihyperglycemic activity of berberine in the Caco-2 cell line is partly based on its α-glucosidase inhibitory activity21,22. Maltase and sucrase activities were also suppressed by berberine23. Additionally, treatment of berberine could considerably attenuate the activities of intestinal disaccharidases in STZ-induced diabetic rats and normal rats24,25. In addition, to enhance the antidiabetic activity of berberine, the introduction of lipophilic substituents has been proved to be a potent strategy since it can improve the pharmacological activities and bioavailability of berberine12,26,27,28. A series of 9-O-berberrubine carboxylates possessing the 9-O-lipophilic group were reported to show low cytotoxicity and good hypoglycemic activity against HepG2 cells29. Compound 29 demonstrated the most substantial increase in hypoglycemic activity with glucose consumption (GC) of 6.73 mM compared with berberine (GC = 5.04 mM) while its cytotoxicity was lower than berberine (Fig. 1). On the other hand, it was reported that yeast α-glucosidase inhibitory activity of xanthone derivatives was considerably improved by adding an aromatic ring to their structure30,31. 3-Arylacyloxyxanthone (X2) was one of the outstanding candidates with the IC50 value of 10.6 μM which was superior to 1,3-dihydroxyxanthone (X1) (IC50 = 145 μM)32. It was claimed that the π-stacking and hydrophobic effects of the additional benzene rings to the α-glucosidase enzyme were the key factors for enhancing inhibitory activity. Therefore, 9-O-berberrubine carboxylate derivatives were designed with the 9-substitution of lipophilic moiety for evaluation of yeast α-glucosidase inhibitory activity (Fig. 1). In this study, baker’s yeast α-glucosidase was used to evaluate inhibitory activity of compounds. Further studies involving rat, human intestinal enzymes and cell-based experiments need to be performed to confirm their antidiabetic activity33,34.
Results and discussion
Synthesis
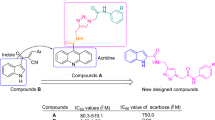
The preparation of BBRB and 9-O-berberrubine carboxylates (1–35) is shown in Fig. 2. BBRB could be efficiently synthesized in the yield of 90% by selective demethylation of BBR chloride at the 9-OCH3 group. Berberine was heated at 190 °C for 1–2 h29. Various carboxylic acids such as benzoic acid, naphthoic acid, cinnamic acid, phenylacetic acid and piperic acid were used to form an ester with berberrubine. Acid chloride derivatives were prepared in situ at room temperature in CH2Cl2 solvent from the corresponding carboxylic acid using the reagents such as triphenylphosphine and trichloroacetonitrile. Then, BBRB and 4-picoline were added to the mixture, and the reaction was refluxed for 8 h to obtain 9-O-berberrubine ester derivatives (1–35) in low-to-moderate yield35,36. 1H-, 13C-NMR and HR-MS spectra of compounds can be seen in the Supplementary Information.
α-Glucosidase inhibitory activity
The inhibitory activity of berberrubine derivatives toward yeast α-glucosidase was evaluated using a protocol similar to those described in the literature37. The percentage inhibition at 50 μM and IC50 values of 1–35, together with BBR, BBRB and acarbose, for comparison are displayed in Tables 1 and 2.
A series of 9-O-berberrubine benzoate derivatives with a wide range of substituents on the benzoate moiety were synthesized and assessed for their α-glucosidase inhibitory activities, the structures and biological results of twenty-six compounds (1–26) are presented in Table 1. 9-O-Berberrubine benzoate (1) showed better % inhibition than BBR and BBRB, i.e., 1 demonstrated 58% inhibition while BBR and BBRB showed no inhibitory activity at 50 μM, which indicated that the introduction of a benzene ester could boost the inhibitory activity as expected. Moreover, 1–35 displayed IC50 values in the range of 1.61–32.84 μM which were higher inhibitory activities than acarbose (IC50 93.60 μM)—a marketed drug for type 2 diabetes. The effect of substituents such as electron-donating, electron-withdrawing and alkyl groups on the additional phenyl ring was then evaluated.
2–4 exhibited higher inhibition with the IC50 values of 19.52, 11.62 and 18.44 μM, respectively compared with 1 (25.32 μM), indicating that the inhibitory activity increased when the methoxy group was substituted to the benzoate aromatic ring. In addition, the methoxy group at the 3-position could improve the activity better than those at the 2- or 4-position. 5 bearing 3-OH group possessed the IC50 value of 7.51 μM which was more active than 3 bearing the 3-OMe group. It could be rationalized by taking into account the hydrogen bonding or other electrostatic interaction with the enzyme31. Hydroxy group is known as an H-bonding donor/acceptor; therefore, it can efficiently form hydrogen-bonding interaction with α-glucosidase enzyme while the methoxy group only acts as an H-bonding acceptor38. Some studies on α-glucosidase inhibitors also proved the importance of the hydroxy group in enhancing inhibitory activity39,40. For multi-methoxy substituent, 6 bearing 3,4-diOCH3 and 8 bearing 3,4,5-triOCH3 groups displayed comparable or even lower activity than mono-methoxy substituent (2–4) with the IC50 values of 19.12 and 23.87 μM, respectively, indicating that addition of more methoxy groups was not a proper way to strengthen the activity. Surprisingly, inhibitory activity was significantly amplified by inserting the 3,4-methylenedioxy group into the phenyl moiety. 7 demonstrated a potent IC50 value of 6.44 μM, which might result from the compactness of the methylenedioxy group compared to the dimethoxy group. Another aromatic ring was added by replacing the 2-methoxy group with the 2-benzyloxy group of benzoate moiety, leading to dramatic enhancement of α-glucosidase inhibitory activity of 9 (IC50 = 2.29 μM) compared with 2 (IC50 = 19.52 μM), which re-emphasized the crucial role of phenyl moiety for the inhibitory activity.
Halogen and other electron-withdrawing groups were also found to have a considerable contribution to the activity. 10–12 bearing chloro substituent at 2-, 3- and 4-position, respectively possessed the IC50 values of 12.21, 9.43 and 10.58 μM, which were better than 1. Because of the potential of chloro group for the activity, other halogen groups such as fluoro, bromo and iodo together with trifluoromethyl, nitro and cyano were introduced. It is worth noting that the fluoro substituent at 2-, 3- or 4-position of 13–15 completely abolished the activity with the IC50 values of 24.89, 32.84 and 21.29 μM, respectively. Nevertheless, 16 and 17 possessing 2-Br and 2-I groups showed better inhibitory activity with the IC50 values of 6.45 and 9.24 μM compared with 10 possessing 2-Cl, especially 16. It has been reported that the chemical softness is an important factor in alleviating α-glucosidase inhibitory activity; thus, chloro, bromo and iodo substituents tend to enhance the activity better than fluoro substituent31,41. On the other hand, 18 and 19 bearing strong EWGs such as 2-CF3 and 2-NO2 groups exhibited the IC50 values of 15.73 and 10.54 μM, which were comparable to 10. Interestingly, the activity of 20 bearing the 4-CN group was diminished with the IC50 value of 26.53 μM. 21 with 2,6-diCl substituents showed significantly high inhibitory activity compared with 10 with the IC50 value of 5.86 μM, indicating the potential of multi-halogen benzoate moiety for α-glucosidase inhibitory activity.
22 and 23 possessing 2- and 4-CH3 groups displayed higher activities than 1, their IC50 values are 18.12 and 17.50 μM, respectively. Replacing the 4-CH3 group with the 4-C2H5 group rendered the amelioration of the inhibitory activity in 24 with the IC50 value of 12.52 μM. The considerably higher IC50 value of 3.53 μM was obtained by 25 possessing 4-C(CH3)3 substituent, implying that the hydrophobic effect from branched alkyl groups is essential for α-glucosidase inhibitory activity. In addition to π-stacking and hydrophobic interaction from aromatic rings which can increase the activity30,31,32, other structural moieties such as the nonconjugated π-system can also be used to modulate the inhibitory activity by hydrophobic interaction42. To prove the hypothesis, 26 bearing two lipophilic tert-butyl groups at 3,5-positions were evaluated for the activity. 26 exhibited even higher inhibitory activity than 25 with the IC50 value of 1.61 μM.
Besides, other kinds of 9-O-berberrubine carboxylates (27–35) were utilized to evaluate their structure-relationship correlation and their biological results are displayed in Table 2. 9-O-Berberrubine 1-naphthoate (27) and 9-O-berberrubine 2-naphthoate (28) demonstrated dramatically high activities with the IC50 values of 2.67 and 2.63 μM. Their inhibitory activities were much higher than 1 due to the π-stacking and hydrophobic effect between extended π-system of the naphthalene ring and α-glucosidase enzyme30,38,42. In addition, their identical IC50 values indicated that there is no preference between linearly and angularly fused aromatic rings to elicit the activity. 9-O-Berberrubine cinnamate (29) and 9-O-berberrubine α-methyl cinnamate (30) exhibited higher activities than 1 with the IC50 values of 15.15 and 10.34 μM. The increase in their inhibitory activities results from the π-stacking and hydrophobic effect of π-conjugated systems of the styryl moiety. The methyl group at α-position of 30 seemed to amplify the hydrophobic effect leading to the higher activity of 30 compared with 29. 9-O-Berberrubine 3,4-methylenedioxycinnamate (31) also showed higher inhibitory activity than 7 with the IC50 of 4.64 μM. 9-O-Berberrubine piperate (33) bearing one more conjugated double bond than 31 displayed even better activity than 31 with the IC50 value of 1.95 μM, which confirmed that π-stacking and hydrophobic effect of long π-conjugated systems involving aromatic rings and double bonds could be utilized to modulate α-glucosidase inhibitory activity. However, there was one exception that 9-O-berberrubine 2,6-dichlorocinnamate (32) showed lower activity than 9-O-berberrubine 2,6-dichlorobenzoate (21), the IC50 value of 32 was 15.96 μM. 9-O-Berberrubine 2,6-dichlorophenylacetate (34) exhibited comparable inhibitory activity to 21 with the IC50 value of 7.30 μM indicating that the conjugation between carbonyl and phenyl groups was not important for the activity. 9-O-Berberrubine diphenylacetate (35) displayed substantially higher inhibitory activity than 9-O-berberrubine benzoate (1) with the IC50 value of 6.19 μM, which re-emphasized that inserting a benzene ring can strengthen the activity. From the analysis of the structure–activity relationship which is also summarized in Fig. 3, five compounds 9, 26, 27, 28 and 33 exhibited the strongest inhibitory activities with the IC50 values of 2.29, 1.61, 2.67, 2.63 and 1.95 μM, respectively. Four compounds (9, 26, 28 and 33) were selected for kinetic study because of the resemblance of structures and activities between 27 and 28. Due to several differences in size and amino acid sequences among yeast, rat and human intestinal α-glucosidase enzymes, additional research relating to rat, human intestinal enzymes and cell-based experiments need to be carried out to corroborate the antidiabetic activity of the potent compounds.43.
Kinetic study
To further investigate how these 9-O-berberrubine carboxylates interact with yeast α-glucosidase, the inhibition types of four potential compounds, 9, 26, 28 and 33 were studied by using Lineweaver–Burk plot analysis. As shown in Fig. 4, the double reciprocal plots showed straight lines with the same Vmax, indicating that 9, 26, 28 and 33 are competitive inhibitors of α-glucosidase. The inhibition constants (Ki) were 29.64 μM for 9, 10.67 μM for 26, 25.27 μM for 28 and 8.64 μM for 33.
Cytotoxicity
To determine the toxicities of the 9-O-berberrubine carboxylate derivatives, the most four potent compounds were tested with HEK-293 cells. The results from three independent experiments showed that 9, 26, 28 and 33 displayed CC50 values of 30.46, 18.16, 20.86 and 17.73 μM, respectively (Fig. 5), indicating that these compounds are not cytotoxic at their IC50 values in the range of 2.63–1.61 μM.
Molecular mechanism of potent compounds
To understand the competitive inhibition mechanism at the atomistic level suggested by the kinetic study, 9, 26, 28, and 33 were docked to the active site of yeast α-glucosidase using the Autodock Vina 1.2.3 program. In this study, the 3D structure of α-glucosidase in the molecular docking study was performed using the same amino sequence as we used in the experimental study, which is a different sequence compared to the available structure in the protein databank (PDB ID: 3A4A). These sequences show 84.9% similarity based on needle pairwise alignment as shown in the Supplementary Information, resulting in a slightly different residue number. Figure 6A shows that the binding pattern for these four compounds is somewhat diverse. The warhead of 9 and 28 inserts to the inner pocket, while that of 26 and 33 points out of the binding pocket. The van der Waals (vdW), π–π, S-π, and alkyl-π interactions are significantly contributed for 9-O-berberrubine carboxylates binding in particular at the core structure. Their theoretical binding affinities are likely comparable, i.e., − 10.40, − 10.87, − 10.46, and − 11.39 kcal/mol for 9, 26, 28, and 33, respectively. By comparing to previous studies44,45, the binding interaction energies of the native inhibitor (acarbose) and the glyceollin were of − 8.70 and − 10.30 kcal/mol, where the electrostatic interaction was identified as a major contribution in the acarbose/α-glucosidase complex46,47. Our compounds shared same interacting residues with acarbose (residues with an asterisk in Fig. 6A). The results suggested that our compounds are potential to be the novel yeast α-glucosidase inhibitor.
(A) Conformation and interaction of four potent 9-O-berberrubine carboxylates 9, 26, 28, and 33 binding to yeast α-glucosidase at the active site resulted from molecular docking study. Black arrow and R212 are used as a reference point for this picture. The residue with asterisk refered to the interacted residue was found in the acarbose/α-glucosidase complex. (B) MM/GBSA and QM-GBSA binding free energies and Pearson correlation coefficient for the four simulated complexes using the 200–300 ns MD trajectories. (C) Water distribution function around 26 via 3D-RISM calculation with g(r) > 5.0 on the minimized structure and the last MD snapshot. The red and white color represent water oxygen and hydrogen distributions. (D) MM/GBSA per-residue energy decomposition for 26/α-glucosidase complex. Key binding residues with binding free energy ≤ − 1.0 kcal/mol are labelled.
To elucidate the structural dynamics of 9-O-berberrubine carboxylates/α-glucosidase complexes, all-atom molecular dynamics (MD) simulation was performed for 300 ns. A total of 100 snapshots extracted from the last 100 ns was analyzed in terms of binding strength based on MM/GBSA and QM-MM/GBSA with AM1, PM3, and PM6 methods treated on the compound molecule only. The plot of binding free energies predicted from all calculations in Fig. 6B reveals that 26 shows the greatest binding affinity among the four studied compounds. Additionally, Pearson correlation coefficient among the calculated binding free energies for these potent compounds shows that our prediction based on four different methods is highly correlated to each other (r2 ≥ 0.97). Importance of water molecules in complexation was then studied by 3D-RISM solvation free energy calculation. The surrounding waters around 26 before and after MD simulation were compared in Fig. 6C. The 3D-RISM solvation free energy difference of ligand between minimized (249.19 kcal/mol) and production phases (232.84 kcal/mol) is about − 16 kcal/mol48,49. The rearrangement of 26 upon simulation increased the solvent molecules as the bridge of interaction between interacted residues and ligand. The MM/GBSA per-residue decomposition free energy analysis on 26/α-glucosidase complex in Fig. 6D revealed that D68, F157, F158 and F177 in the active site were important for the binding of 26.
Conclusion
In conclusion, a series of 9-O-berberrubine carboxylates have been synthesized and evaluated as yeast α-glucosidase inhibitors. All compounds showed higher inhibitory activities compared with BBR, BBRB and acarbose. The analysis of structure–activity relationship revealed that hydrogen bonding, π-stacking, hydrophobic effect and chemical softness are essential factors modulating the activity. 9, 26, 27, 28 and 33 displayed the strongest activities with the IC50 values in the range of 2.67–1.61 μM. 9, 26, 28 and 33 were found to inhibit yeast α-glucosidase via a competitive mechanism. Additionally, cytotoxicity results verified the safety of these four compounds at their IC50 values because their CC50 values were higher than 15 μM. The theoretical binding affinities of these four compounds (− 10.40 to − 11.39 kcal/mol) were higher than those of acarbose and glyceollin (− 8.47 and − 10.30 kcal/mol). These ligands showed a high binding stability over the 300-ns MD simulations. Among the outstanding compounds, 26 was the most susceptible compound toward yeast α-glucosidase. Overall, these potent 9-O-berberrubine carboxylates could be utilized for further studies of antidiabetic activity.
Methods
General information
The reagents (chemicals) were purchased from TCI and used without further purification. α-Glucosidase from Saccharomyces cerevisiae (EC.3.2.1.2.0) and p-NPG (p-nitrophenyl-α-D-glucopyranoside), and acarbose (positive control) were purchased from Sigma Aldrich. Nuclear magnetic resonance (NMR) spectroscopy was performed on JEOL JNM-ECZ500R/S1 spectrometer at 500 MHz (500 MHz for 1H NMR, 125 MHz for 13C NMR). Chemical shifts were reported in parts per million (ppm, d) referenced to the residual solvent signals (DMSO-d6: δH = 2.50, δC = 39.5 ppm; Methanol-d4: δH = 3.31, δC = 49.0 ppm). Proton coupling patterns were described as singlet (s), doublet (d), triplet (t), quartet (q), multiplet (m), and broad (br). HRESIMS were determined on a micrOTOF-Q II 10,335. All reactions were monitored by thin-layer chromatography (TLC) on silica gel G1P4S6. Anhydrous solvents were purchased from commercial suppliers.
General procedure for pyrolysis of berberine
The pyrolysis of berberine (BBR) (3.0 g, 8.068 mmol) was performed at 190 °C for 1–2 h to produce berberrubine (BBRB).
9-hydroxy-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (BBRB): Dark red powder (2.598 g, 90%); 1H NMR (500 MHz, CD3OD): δ ppm 9.24 (1H, s), 8.00 (1H, s), 7.50 (d, J = 8.5 Hz, 1H), 7.39 (s, 1H), 6.89 (d, J = 8.0 Hz, 1H), 6.81 (s, 1H), 6.01 (s, 2H), 4.59 (t, J = 6.0 Hz, 2H), 3.85 (s, 3H), 3.10 (t, J = 6.5 Hz, 2H); 13C NMR (125 MHz, CD3OD): 162.8, 150.9, 150.7, 149.5, 147.2, 135.8, 133.7, 130.7, 124.1, 122.8, 121.4, 119.8, 109.2, 105.8, 103.3, 98.8, 56.7, 55.7, 28.9.
General procedure for synthesizing compounds 1–35
Trichloroacetonitrile (0.3 mL, 3 mmol) was added to a mixture of carboxylic acid (1.5 mmol) and triphenylphosphine (0.787 g, 3 mmol) in CH2Cl2 (20 mL) at room temperature. The mixture was stirred for 1–2 h. After that, berberrubine (0.358 g, 1 mmol) and 4-picoline (0.292 mL, 3 mmol) were added to the above mixture and the reaction mixture was refluxed for 8 h. Then, the organic layer was extracted with 1 M HCl and saturated NaHCO3, dried over anhydrous MgSO4, and evaporated in vacuo. The residue was purified by silica gel column chromatography (chloroform/methanol) to afford the products (1–35).
9-(benzoyloxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (1): Yellow powder (0.168 g, 36%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.92 (1H, s), 9.02 (1H, s), 8.26 (1H, d, J = 9.5 Hz), 8.21 (2H, m), 8.20 (1H, d, J = 8.0 Hz), 7.78 (1H, t, J = 7.5 Hz), 7.76 (1H, d, J = 5.0 Hz), 7.63 (1H, s), 7.62 (1H, t, J = 8.0 Hz), 7.01 (1H, s), 6.10 (2H, s), 4.84 (2H, t, J = 6.5 Hz), 3.95 (3H, s), 3.12 (2H, t, J = 6.0 Hz); 13C NMR (125 MHz, DMSO-d6): 163.5, 150.5, 150.0, 147.8, 144.6, 138.2, 134.7, 133.6, 133.0, 130.9, 130.5, 129.2, 128.0, 127.0, 125.9, 121.3, 120.7, 120.4, 108.5, 105.6, 102.2, 57.3, 55.3, 26.2.
10-methoxy-9-((2-methoxybenzoyl)oxy)-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (2): Yellow powder (0.023 g, 5%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.85 (1H, s), 9.05 (1H, s), 8.32 (1H, d, J = 9.0 Hz), 8.25 (1H, d, J = 9.0 Hz), 8.18 (1H, dd, J = 8.0, 2.0 Hz), 7.82 (1H, s), 7.75 (1H, td, J = 7.8, 1.5 Hz), 7.32 (1H, d, J = 8.5 Hz), 7.18 (1H, t, J = 8.0 Hz), 7.08 (1H, s), 6.17 (2H, s), 4.92 (2H, t, J = 6.5 Hz), 4.03 (3H, s), 3.92 (3H, s), 3.20 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 161.8, 160.3, 150.8, 150.1, 147.8, 144.6, 138.2, 135.8, 133.9, 132.8, 132.7, 131.1, 126.8, 126.0, 121.2, 120.7, 120.5, 120.4, 116.8, 113.1, 108.6, 105.6, 102.2, 57.4, 56.2, 55.6, 26.3; HRMS (ESI) calcd for C27H22ClNO6 [M–Cl]+ 456.14416, found 456.1442.
10-methoxy-9-((3-methoxybenzoyl)oxy)-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (3): Yellow powder (0.148 g, 30%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.98 (1H, s), 9.08 (1H, s), 8.34 (1H, d, J = 9.0 Hz), 8.28 (1H, d, J = 9.5 Hz), 7.86 (1H, d, J = 8.0 Hz), 7.82 (1H, s), 7.73 (1H, t, J = 2.5 Hz), 7.62 (1H, t, J = 8.0 Hz), 7.42 (1H, dd, J = 8.5, 3.0 Hz), 7.09 (1H, s), 6.18 (2H, s), 4.91 (2H, t, J = 5.5 Hz), 4.03 (3H, s), 3.89 (3H, s), 3.20 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 163.6, 159.8, 150.7, 150.3, 148.0, 144.7, 138.4, 133.8, 133.3, 131.2, 130.7, 129.5, 127.4, 126.1, 123.0, 121.5, 120.9, 120.6, 115.3, 108.7, 105.8, 102.4, 57.6, 55.9, 55.6, 26.4.
10-methoxy-9-((4-methoxybenzoyl)oxy)-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (4): Yellow powder (0.338 g, 69%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.92 (1H, s), 9.04 (1H, s), 8.29 (1H, d, J = 9.5 Hz), 8.24 (1H, d, J = 9.0 Hz), 8.20 (2H, d, J = 8.5 Hz), 7.78 (1H, s), 7.20 (2H, d, J = 8.5 Hz), 7.06 (1H, s), 6.15 (2H, s), 4.90 (2H, t, J = 6.0 Hz), 4.00 (3H, s), 3.91 (3H, s), 3.18 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 164.4, 163.2, 150.6, 150.1, 147.8, 144.6, 138.2, 133.9, 133.1, 132.9, 131.0, 127.0, 125.9, 121.5, 120.8, 120.4, 120.1, 114.6, 108.5, 105.7, 102.3, 57.4, 56.0, 55.4, 26.3.
9-((3-hydroxybenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (5): Yellow powder (0.168 g, 35%); 1H NMR (500 MHz, DMSO-d6): δ ppm 10.29 (1H, s), 9.96 (1H, s), 9.09 (1H, s), 8.33 (1H, d, J = 9.5 Hz), 8.26 (1H, d, J = 9.5 Hz), 7.82 (1H, s), 7.69 (1H, m), 7.68 (1H, d, J = 2.0 Hz), 7.47 (1H, t, J = 8.0 Hz), 7.25 (1H, m), 7.08 (1H, s), 6.17 (2H, s), 4.92 (2H, t, J = 6.0 Hz), 4.02 (3H, s), 3.20 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 163.5, 158.0, 150.4, 150.0, 147.7, 144.5, 138.1, 133.7, 133.0, 130.9, 130.1, 129.0, 127.0, 125.9, 121.7, 121.3, 121.0, 120.7, 120.4, 116.8, 108.4, 105.6, 102.2, 57.3, 53.3, 26.2; HRMS (ESI) calcd for C26H20ClNO6 [M–Cl]+ 442.12851, found 442.1297.
9-((3,4-dimethoxybenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (6): Brown powder (0.32 g, 61%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.89 (1H, s), 9.03 (1H, s), 8.26 (1H, d, J = 9.0 Hz), 8.20 (1H, d, J = 9.0 Hz), 7.86 (1H, dd, J = 8.5, 2.0 Hz), 7.76 (1H, s), 7.63 (1H, d, J = 2.5 Hz), 7.18 (1H, d, J = 8.5 Hz), 7.02 (1H, s), 6.12 (2H, s), 4.87 (2H, t, J = 6.5 Hz), 3.96 (3H, s), 3.86 (3H, s), 3.82 (3H, s), 3.14 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 163.2, 154.2, 150.6, 150.0, 148.7, 147.8, 144.5, 138.1, 133.8, 133.0, 130.9, 126.9, 125.9, 125.0, 121.4, 120.7, 120.4, 119.8, 112.6, 111.5, 108.4, 105.6, 102.2, 57.3, 56.0, 55.8, 55.3, 26.2.
9-((benzo[d][1,3]dioxole-5-carbonyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (7): Brown powder (0.188 g, 37%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.92 (1H, s), 9.04 (1H, s), 8.31 (1H, d, J = 9.0 Hz), 8.25 (1H, d, J = 9.0 Hz), 7.88 (1H, dd, J = 8.0, 1.5 Hz), 7.81 (1H, s), 7.69 (1H, d, J = 2.0 Hz), 7.20 (1H, d, J = 8.0 Hz), 7.08 (1H, s), 6.23 (2H, s), 6.17 (2H, s), 4.90 (2H, t, J = 6.0 Hz), 4.01 (3H, s), 3.19 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 163.5, 153.1, 150.9, 150.5, 148.3, 148.2, 144.8, 138.6, 134.0, 133.4, 131.4, 127.5, 127.4, 126.2, 121.8, 121.7, 121.0, 120.7, 110.1, 109.2, 108.9, 106.0, 103.0, 102.6, 57.7, 55.9, 26.6.
10-methoxy-9-((3,4,5-trimethoxybenzoyl)oxy)-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (8): Brown powder (0.069 g, 12%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.96 (1H, s), 9.07 (1H, s), 8.32 (1H, d, J = 9.5 Hz), 8.27 (1H, d, J = 9.5 Hz), 7.81 (1H, s), 7.53 (2H, s), 7.08 (1H, s), 6.17 (2H, s), 4.92 (2H, t, J = 6.0 Hz), 4.02 (3H, s), 3.91 (6H, s), 3.81 (3H, s), 3.19 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 163.1, 153.1, 150.6, 150.1, 147.8, 144.6, 143.0, 138.3, 133.6, 133.1, 131.0, 127.2, 125.9, 122.9, 121.3, 120.8, 120.5, 108.6, 107.9, 105.7, 102.3, 60.5, 57.4, 56.4, 55.4, 26.3.
9-((2-(benzyloxy)benzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (9): Yellow powder (0.171 g, 30%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.86 (1H, s), 9.08 (1H, s), 8.32 (1H, d, J = 9.5 Hz), 8.26 (1H, d, J = 9.5 Hz), 8.24 (1H, dd, J = 8.0, 2.0 Hz), 7.82 (1H, s), 7.75 (1H, td, J = 9.0, 2.0 Hz), 7.52 (2H, d, J = 7.0 Hz), 7.42 (1H, d, J = 9.0 Hz), 7.33 (2H, t, J = 7.5 Hz), 7.27 (1H, t, J = 7.0 Hz), 7.21 (1H, t, J = 7.0 Hz), 7.07 (1H, s), 6.17 (2H, s), 5.31 (2H, s), 4.85 (2H, t, J = 5.5 Hz), 4.01 (3H, s), 3.17 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 161.8, 159.1, 150.5, 150.0, 147.8, 144.4, 138.1, 136.8, 135.7, 133.8, 133.1, 132.9, 130.9, 128.4, 127.8, 127.2, 126.8, 126.0, 121.4, 120.8, 120.6, 120.4, 117.2, 114.3, 108.5, 105.6, 102.2, 69.8, 57.3, 55.4, 26.2; HRMS (ESI) calcd for C33H26ClNO6 [M–Cl]+ 532.17546, found 532.1761.
9-((2-chlorobenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (10): Yellow powder (0.077 g, 16%); 1H NMR (500 MHz, DMSO-d6): δ ppm 10.00 (1H, s), 9.09 (1H, s), 8.43 (1H, dd, J = 8.0, 2.0 Hz), 8.33 (1H, d, J = 9.0 Hz), 8.28 (1H, d, J = 9.5 Hz), 7.81 (1H, s), 7.79 (1H, td, J = 8.0, 2.0 Hz), 7.75 (1H, dd, J = 7.5, 1.5 Hz), 7.66 (1H, td, J = 7.5, 2.0 Hz), 7.08 (1H, s), 6.17 (2H, s), 4.94 (2H, t, J = 6.0 Hz), 4.05 (3H, s), 3.21 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 161.6, 150.3, 150.1, 147.8, 144.5, 138.3, 135.0, 134.0, 133.3, 133.2, 133.1, 131.7, 131.0, 127.8, 127.3, 127.0, 126.0, 121.1, 120.8, 120.4, 108.5, 105.7, 102.2, 57.5, 55.4, 26.2; HRMS (ESI) calcd for C26H19Cl2NO5 [M–Cl]+ 460.09463, found 460.09953.
9-((3-chlorobenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (11): Yellow powder (0.12 g, 24%); 1H NMR (500 MHz, DMSO-d6): δ ppm 10.02 (1H, s), 9.08 (1H, s), 8.33 (1H, d, J = 9.5 Hz), 8.28 (1H, d, J = 9.5 Hz), 8.26 (1H, d, J = 4.0 Hz), 8.21 (1H, dd, J = 8.0, 3.0 Hz), 7.93 (1H, dd, J = 8.0, 3.0 Hz), 7.81 (1H, s), 7.74 (1H, t, J = 8.0 Hz), 7.08 (1H, s), 6.17 (2H, s), 4.90 (2H, t, J = 6.0 Hz), 4.02 (3H, s), 3.20 (2H, t, J = 6.0 Hz); 13C NMR (125 MHz, DMSO-d6): 162.4, 150.4, 150.1, 147.8, 144.6, 138.3, 134.5, 133.9, 133.3, 133.1, 131.3, 131.0, 130.1, 129.9, 129.2, 127.3, 125.9, 121.1, 120.7, 120.4, 108.5, 105.7, 102.2, 57.4, 55.4, 26.2; HRMS (ESI) calcd for C26H19Cl2NO5 [M–Cl]+ 460.09463, found 460.09963.
9-((4-chlorobenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (12): Brown powder (0.034 g, 7%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.97 (1H, s), 9.06 (1H, s), 8.30 (1H, d, J = 9.0 Hz), 8.24 (1H, d, J = 9.0 Hz), 8.23 (2H, d, J = 8.5 Hz), 7.78 (1H, s), 7.74 (2H, d, J = 8.0 Hz), 7.04 (1H, s), 6.14 (2H, s), 4.86 (2H, t, J = 7.0 Hz), 3.98 (3H, s), 3.16 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 162.7, 150.4, 150.0, 147.7, 144.5, 139.6, 138.2, 133.3, 133.0, 132.3, 130.9, 129.3, 127.2, 126.8, 125.9, 121.2, 120.7, 120.4, 108.4, 105.6, 102.2, 57.3, 55.3, 26.2.
9-((2-fluorobenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (13): Yellow powder (0.207 g, 43%); 1H NMR (500 MHz, DMSO-d6): δ ppm 10.01 (1H, s), 9.10 (1H, s), 8.34 (1H, d, J = 9.5 Hz), 8.28 (2H, m), 7.89 (1H, m), 7.82 (1H, s), 7.53 (1H, t, J = 8.0 Hz), 7.51 (1H, t, J = 7.0 Hz), 7.08 (1H, s), 6.17 (2H, s), 4.92 (2H, t, J = 6.5 Hz), 4.04 (3H, s), 3.21 (2H, t, J = 6.0 Hz); 13C NMR (125 MHz, DMSO-d6): 163.5, 161.4, 161.1, 161.0, 150.9, 150.5, 148.2, 145.0, 138.7, 137.5, 137.4, 133.7, 133.5, 133.4, 131.4, 127.7, 126.4, 125.6, 125.5, 121.6, 121.2, 120.9, 118.1, 118.0, 116.9, 116.8, 109.0, 106.1, 102.7, 57.9, 55.8, 26.7; HRMS (ESI) calcd for C26H19ClFNO5 [M–Cl]+ 444.12418, found 444.1247.
9-((3-fluorobenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (14): Yellow powder (0.278 g, 58%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.96 (1H, s), 9.04 (1H, s), 8.25 (1H, d, J = 9.0 Hz), 8.20 (1H, d, J = 9.0 Hz), 8.04 (1H, d, J = 7.5 Hz), 7.96 (1H, dt, J = 9.0, 2.5 Hz), 7.73 (1H, s), 7.68 (1H, dd, J = 13.5, 8.5 Hz), 7.65 (1H, td, J = 8.5, 3.0 Hz), 6.99 (1H, s), 6.08 (2H, s), 4.84 (2H, t, J = 6.0 Hz), 3.94 (3H, s), 3.12 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 163.1, 162.3, 161.1, 150.4, 150.0, 147.7, 144.5, 138.2, 133.2, 133.0, 131.5, 131.4, 130.9, 130.3, 130.2, 127.2, 126.7, 125.9, 121.8, 121.6, 121.0, 120.7, 120.3, 117.1, 116.9, 108.4, 105.6, 102.2, 57.4, 55.3, 26.2; HRMS (ESI) calcd for C26H19ClFNO5 [M–Cl]+ 444.12418, found 444.1247.
9-((4-fluorobenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (15): Yellow powder (0.099 g, 21%); 1H NMR (500 MHz, DMSO-d6): δ ppm 10.01 (1H, s), 9.10 (1H, s), 8.34 (3H, m), 8.28 (1H, d, J = 9.5 Hz), 7.83 (1H, s), 7.55 (2H, t, J = 8.5 Hz), 7.09 (1H, s), 6.18 (2H, s), 4.91 (2H, t, J = 6.5 Hz), 4.03 (3H, s), 3.20 (2H, t, J = 6.0 Hz); 13C NMR (125 MHz, DMSO-d6): 167.0, 164.9, 162.5, 150.4, 150.0, 147.8, 144.5, 138.2, 133.6, 133.5, 133.4, 133.0, 130.9, 127.1, 125.9, 124.6, 121.2, 120.7, 120.4, 116.5, 116.3, 108.5, 105.6, 102.2, 57.3, 55.3, 26.2; HRMS (ESI) calcd for C26H19ClFNO5 [M–Cl]+ 444.12418, found 444.1247.
9-((2-bromobenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (16): Brown powder (0.165 g, 30%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.97 (1H, s), 9.06 (1H, s), 8.42 (1H, d, J = 7.0 Hz), 8.33 (1H, d, J = 9.0 Hz), 8.28 (1H, d, J = 9.5 Hz), 7.93 (1H, d, J = 8.5 Hz), 7.81 (1H, s), 7.70 (1H, t, J = 8.5 Hz), 7.69 (1H, t, J = 8.5 Hz), 7.08 (1H, s), 6.17 (2H, s), 4.93 (2H, s), 4.05 (3H, s), 3.20 (2H, t, J = 5.5 Hz); 13C NMR (125 MHz, DMSO-d6): 162.2, 150.4, 150.2, 147.9, 144.5, 138.4, 135.1, 133.4, 133.3, 133.2, 131.0, 129.0, 128.3, 127.4, 126.0, 122.5, 121.1, 120.8, 120.5, 108.6, 105.7, 102.3, 57.5, 55.5, 26.3; HRMS (ESI) calcd for C26H19ClBrNO5 [M–Cl]+ 504.04411, found 504.0447.
9-((2-iodobenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (17): Yellow powder (0.2 g, 34%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.99 (1H, s), 9.08 (1H, s), 8.41 (1H, d, J = 8.0 Hz), 8.35 (1H, d, J = 9.5 Hz), 8.28 (1H, d, J = 9.0 Hz), 8.22 (1H, dd, J = 7.5, 1.5 Hz), 7.83 (1H, s), 7.71 (1H, t, J = 7.5 Hz), 7.47 (1H, td, J = 7.5, 2.0 Hz), 7.10 (1H, s), 6.18 (2H, s), 4.94 (2H, t, J = 6.0 Hz), 4.06 (3H, s), 3.21 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 162.8, 150.4, 150.2, 147.9, 144.5, 142.0, 138.4, 134.8, 133.5, 133.2, 132.8, 131.9, 131.1, 128.7, 127.3, 126.0, 121.2, 120.8, 120.5, 108.6, 105.7, 102.3, 96.8, 57.5, 55.5, 26.3; HRMS (ESI) calcd for C26H19ClINO5 [M–Cl]+ 552.03024, found 552.0307.
10-methoxy-9-((2-(trifluoromethyl)benzoyl)oxy)-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (18): Yellow powder (0.206 g, 39%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.96 (1H, s), 9.13 (1H, s), 8.62 (1H, d, J = 7.0 Hz), 8.35 (1H, d, J = 9.5 Hz), 8.30 (1H, d, J = 9.0 Hz), 8.07 (1H, m), 8.01 (2H, m), 7.83 (1H, s), 7.09 (1H, s), 6.18 (2H, s), 4.98 (2H, t, J = 6.5 Hz), 4.05 (3H, s), 3.22 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 161.9, 150.4, 150.0, 147.7, 144.1, 138.3, 133.8, 133.1, 133.0, 132.9, 132.5, 130.9, 127.6, 127.5, 127.4, 126.0, 120.9, 120.8, 120.3, 108.6, 108.5, 108.4, 105.6, 102.2, 57.3, 55.5, 26.2; HRMS (ESI) calcd for C27H19ClF3NO5 [M–Cl]+ 494.12098, found 494.1216.
10-methoxy-9-((2-nitrobenzoyl)oxy)-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (19): Yellow powder (0.162 g, 32%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.92 (1H, s), 9.12 (1H, s), 8.46 (1H, m), 8.35 (1H, d, J = 9.0 Hz), 8.30 (1H, d, J = 9.0 Hz), 8.18 (1H, m), 8.03 (2H, m), 7.83 (1H, s), 7.09 (1H, s), 6.18 (2H, s), 4.95 (2H, t, J = 6.0 Hz), 4.09 (3H, s), 3.22 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 161.0, 150.4, 150.1, 149.1, 147.8, 144.0, 138.4, 134.9, 133.2, 133.1, 132.4, 131.6, 130.9, 127.6, 126.0, 124.3, 122.5, 121.2, 120.8, 120.3, 108.5, 105.6, 102.2, 57.5, 55.6, 26.2.
9-((4-cyanobenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (20): Brown powder (0.138 g, 28%); 1H NMR (500 MHz, DMSO-d6): δ ppm 10.02 (1H, s), 9.07 (1H, s), 8.42 (2H, d, J = 9.0 Hz), 8.34 (1H, d, J = 9.5 Hz), 8.29 (1H, d, J = 9.0 Hz), 8.18 (2H, d, J = 8.5 Hz), 7.82 (1H, s), 7.09 (1H, s), 6.17 (2H, s), 4.88 (2H, t, J = 6.0 Hz), 4.03 (3H, s), 3.20 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 162.4, 150.4, 150.2, 147.8, 144.8, 138.4, 133.2, 133.1, 133.0, 132.0, 131.1, 131.0, 127.4, 126.0, 121.1, 120.7, 120.4, 118.1, 116.6, 108.5, 105.7, 102.2, 57.5, 55.4, 26.2; HRMS (ESI) calcd for C27H19ClN2O5 [M–Cl]+ 451.12885, found 451.1288.
9-((2,6-dichlorobenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (21): Yellow powder (0.098 g, 18%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.64 (1H, s), 9.12 (1H, s), 8.40 (1H, d, J = 9.0 Hz), 8.32 (1H, d, J = 9.5 Hz), 7.83 (1H, s), 7.79 (1H, s), 7.78 (1H, d, J = 8.5 Hz), 7.71 (1H, dd, J = 8.5, 7.0 Hz), 7.11 (1H, s), 6.19 (2H, s), 4.91 (2H, t, J = 6.5 Hz), 4.12 (3H, s), 3.22 (2H, t, J = 7.0 Hz); 13C NMR (125 MHz, DMSO-d6): 161.3, 151.2, 150.8, 143.6, 139.1, 134.0, 133.9, 132.4, 131.9, 131.8, 131.5, 129.5, 129.4, 128.3, 128.2, 126.4, 124.0, 121.3, 120.8, 120.4, 108.8, 106.0, 102.8, 57.6, 57.2, 26.2; HRMS (ESI) calcd for C26H18Cl3NO5 [M–Cl]+ 494.05565, found 494.0565.
10-methoxy-9-((2-methylbenzoyl)oxy)-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (22): Yellow powder (0.014 g, 3%); 1H NMR (500 MHz, DMSO-d6): δ ppm 10.00 (1H, s), 9.09 (1H, s), 8.34 (1H, d, J = 9.5 Hz), 8.32 (1H, d, J = 7.0 Hz), 8.27 (1H, d, J = 9.0 Hz), 7.83 (1H, s), 7.68 (1H, t, J = 7.5 Hz), 7.51 (1H, d, J = 7.5 Hz), 7.50 (1H, t, J = 8.0 Hz), 7.09 (1H, s), 6.18 (2H, s), 4.92 (2H, t, J = 6.0 Hz), 4.04 (3H, s), 3.20 (2H, t, J = 6.0 Hz), 2.64 (3H, s); 13C NMR (125 MHz, DMSO-d6): 164.4, 150.9, 150.5, 148.2, 145.1, 141.6, 138.6, 134.3, 134.2, 133.5, 132.6, 132.2, 131.4, 127.7, 127.4, 126.9, 126.4, 121.8, 121.2, 120.9, 109.0, 106.1, 102.7, 57.8, 55.8, 26.7, 21.9; HRMS (ESI) calcd for C27H22ClNO5 [M–Cl]+ 440.14925, found 440.1498.
10-methoxy-9-((4-methylbenzoyl)oxy)-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (23): Yellow powder (0.2 g, 42%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.96 (1H, s), 9.09 (1H, s), 8.33 (1H, d, J = 9.0 Hz), 8.27 (1H, d, J = 9.0 Hz), 8.16 (2H, d, J = 8.0 Hz), 7.83 (1H, s), 7.50 (2H, d, J = 8.5 Hz), 7.08 (1H, s), 6.18 (2H, s), 4.91 (2H, t, J = 6.0 Hz), 4.02 (3H, s), 3.19 (2H, t, J = 6.5 Hz), 2.48 (3H, s); 13C NMR (125 MHz, DMSO-d6): 163.4, 150.5, 150.0, 147.7, 145.3, 144.5, 138.2, 133.7, 133.0, 130.9, 130.5, 129.7, 126.9, 125.9, 125.2, 121.3, 120.7, 120.4, 108.4, 105.6, 102.2, 57.3, 55.3, 26.2, 21.4.
9-((4-ethylbenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (24): Yellow powder (0.017 g, 3%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.96 (1H, s), 9.05 (1H, s), 8.33 (1H, d, J = 9.0 Hz), 8.26 (1H, d, J = 9.5 Hz), 8.17 (2H, d, J = 8.0 Hz), 7.82 (1H, s), 7.53 (2H, d, J = 8.0 Hz), 7.08 (1H, s), 6.18 (2H, s), 4.89 (2H, t, J = 6.5 Hz), 4.01 (3H, s), 3.18 (2H, t, J = 6.5 Hz), 2.77 (2H, q, J = 7.5 Hz), 1.25 (3H, t, J = 7.5 Hz); 13C NMR (125 MHz, DMSO-d6): 163.6, 151.6, 150.6, 150.2, 147.9, 144.6, 138.3, 133.8, 133.1, 131.0, 130.8, 128.7, 127.1, 126.0, 125.6, 121.4, 120.8, 120.5, 108.6, 105.7, 102.3, 57.4, 55.4, 28.6, 26.3, 15.6; HRMS (ESI) calcd for C28H24ClNO5 [M–Cl]+ 454.16490, found 454.1654.
9-((4-(tert-butyl)benzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (25): Yellow powder (0.174 g, 34%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.95 (1H, s), 9.04 (1H, s), 8.32 (1H, d, J = 9.0 Hz), 8.26 (1H, d, J = 9.0 Hz), 8.19 (2H, d, J = 8.0 Hz), 7.82 (1H, s), 7.71 (2H, d, J = 8.5 Hz), 7.08 (1H, s), 6.17 (2H, s), 4.89 (2H, t, J = 6.5 Hz), 4.01 (3H, s), 3.18 (2H, t, J = 6.0 Hz), 1.36 (9H, s); 13C NMR (125 MHz, DMSO-d6): 163.5, 158.1, 150.6, 150.1, 147.8, 144.6, 138.3, 133.7, 133.1, 131.0, 130.5, 127.1, 126.1, 126.0, 125.3, 121.4, 120.8, 120.5, 108.6, 105.7, 102.3, 57.4, 55.3, 35.2, 30.9, 26.2.
9-((3,5-di-tert-butylbenzoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (26): Yellow powder (0.205 g, 36%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.99 (1H, s), 9.07 (1H, s), 8.34 (1H, d, J = 9.5 Hz), 8.27 (1H, d, J = 9.5 Hz), 8.06 (2H, d, J = 2.0 Hz), 7.90 (1H, t, J = 2.0 Hz), 7.83 (1H, s), 7.09 (1H, s), 6.18 (2H, s), 4.90 (2H, t, J = 5.5 Hz), 4.03 (3H, s), 3.19 (2H, t, J = 6.0 Hz), 1.38 (18H, s); 13C NMR (125 MHz, DMSO-d6): 164.1, 158.4, 151.5, 150.1, 147.8, 144.6, 138.2, 133.8, 133.0, 130.9, 129.0, 127.3, 126.8, 125.6, 124.2, 121.4, 120.7, 120.4, 108.5, 105.6, 102.2, 57.3, 55.5, 34.8, 31.1, 26.2; HRMS (ESI) calcd for C34H36ClNO5 [M–Cl]+ 538.25880, found 538.2592.
9-((1-naphthoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (27): Yellow powder; (0.172 g, 34%); 1H NMR (500 MHz, DMSO-d6): δ ppm 10.07 (1H, s), 9.13 (1H, s), 8.91 (1H, d, J = 8.5 Hz), 8.74 (1H, d, J = 7.5 Hz), 8.41 (1H, d, J = 8.5 Hz), 8.37 (1H, d, J = 9.5 Hz), 8.31 (1H, d, J = 9.5 Hz), 8.16 (1H, d, J = 8.0 Hz), 7.84 (1H, s), 7.80 (1H, t, J = 7.5 Hz), 7.75 (1H, t, J = 7.0 Hz), 7.69 (1H, t, J = 8.0 Hz), 7.08 (1H, s), 6.18 (2H, s), 4.93 (2H, t, J = 6.5 Hz), 4.07 (3H, s), 3.20 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 163.8, 150.5, 150.0, 147.7, 144.6, 138.2, 135.1, 133.8, 133.6, 133.1, 132.3, 131.0, 130.9, 129.1, 128.6, 127.0, 126.8, 125.9, 125.1, 125.0, 124.1, 121.4, 120.7, 120.4, 108.4, 105.6, 102.2, 57.4, 55.3, 26.2.
9-((2-naphthoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (28): Yellow powder (0.155 g, 30%); 1H NMR (500 MHz, DMSO-d6): δ ppm 10.06 (1H, s), 9.13 (1H, s), 9.01 (1H, s), 8.36 (1H, d, J = 9.5 Hz), 8.30 (1H, d, J = 9.5 Hz), 8.26 (1H, d, J = 8.0 Hz), 8.23 (1H, d, J = 8.5 Hz), 8.20 (1H, d, J = 9.0 Hz), 8.12 (1H, d, J = 8.0 Hz), 7.84 (1H, s), 7.78 (1H, t, J = 7.0 Hz), 7.71 (1H, t, J = 7.0 Hz), 7.08 (1H, s), 6.18 (2H, s), 4.92 (2H, t, J = 7.0 Hz), 4.04 (3H, s), 3.20 (2H, t, J = 7.0 Hz); 13C NMR (125 MHz, DMSO-d6): 163.6, 150.5, 150.0, 147.7, 144.5, 138.2, 135.6, 133.7, 133.0, 132.4, 132.1, 130.9, 129.6, 129.4, 128.7, 127.9, 127.4, 127.0, 125.9, 125.4, 125.2, 121.3, 120.7, 120.4, 108.4, 105.6, 102.2, 57.3, 55.3, 26.2.
9-(cinnamoyloxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (29): Yellow powder (0.051 g, 10%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.94 (1H, s), 9.02 (1H, s), 8.29 (1H, d, J = 9.5 Hz), 8.24 (1H, d, J = 9.0 Hz), 7.99 (1H, d, J = 16.0 Hz), 7.86 (2H, m), 7.80 (1H, s), 7.51 (1H, s), 7.50 (2H, m), 7.07 (1H, s), 7.05 (1H, d, J = 16.0 Hz), 6.16 (2H, s), 4.92 (2H, t, J = 6.0 Hz), 4.03 (3H, s), 3.20 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 163.9, 150.7, 150.2, 147.9, 147.8, 144.5, 138.3, 133.9, 133.7, 133.1, 131.5, 131.0, 129.4, 129.0, 127.0, 126.0, 121.4, 120.8, 120.5, 116.4, 108.6, 105.7, 102.3, 57.4, 55.5, 26.4.
(E)-10-methoxy-9-((2-methyl-3-phenylacryloyl)oxy)-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (30): Yellow powder (0.025 g, 5%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.95 (1H, s), 9.08 (1H, s), 8.32 (1H, d, J = 9.5 Hz), 8.25 (1H, d, J = 9.5 Hz), 8.05 (1H, s), 7.83 (1H, s), 7.66 (2H, d, J = 7.5 Hz), 7.53 (2H, t, J = 7.5 Hz), 7.47 (1H, t, J = 7.5 Hz), 7.10 (1H, s), 6.18 (2H, s), 4.96 (2H, t, J = 6.5 Hz), 4.05 (3H, s), 3.21 (2H, t, J = 6.5 Hz), 2.32 (3H, d, J = 9.5 Hz); 13C NMR (125 MHz, DMSO-d6): 165.9, 151.0, 150.5, 148.2, 145.0, 142.0, 138.6, 135.3, 134.4, 133.5, 131.4, 130.7, 129.9, 129.3, 127.3, 126.7, 126.4, 121.8, 121.2, 120.9, 109.0, 106.1, 102.7, 57.8, 55.8, 26.7, 15.0; HRMS (ESI) calcd for C29H24ClNO5 [M–Cl]+ 466.16490, found 466.1654.
(E)-9-((3-(benzo[d][1,3]dioxol-5-yl)acryloyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (31): Brown powder (0.108 g, 20%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.93 (1H, s), 9.05 (1H, s), 8.31 (1H, d, J = 9.5 Hz), 8.23 (1H, d, J = 9.0 Hz), 7.90 (1H, d, J = 16.0 Hz), 7.82 (1H, s), 7.59 (1H, d, J = 2.0 Hz), 7.36 (1H, dd, J = 8.5, 2.0 Hz), 7.09 (1H, s), 7.04 (1H, d, J = 8.0 Hz), 6.92 (1H, d, J = 16.0 Hz), 6.18 (2H, s), 6.14 (2H, s), 4.93 (2H, t, J = 6.0 Hz), 4.04 (3H, s), 3.21 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 164.2, 150.8, 150.3, 150.2, 148.4, 147.9, 147.8, 144.6, 138.3, 133.8, 133.2, 131.1, 128.4, 127.0, 126.2, 126.0, 121.6, 120.8, 120.5, 114.0, 108.9, 108.7, 107.2, 105.8, 102.4, 102.1, 57.4, 55.6, 26.4.
(E)-9-((3-(2,6-dichlorophenyl)acryloyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (32): Brown powder (0.024 g, 4%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.96 (1H, s), 9.04 (1H, s), 8.32 (1H, d, J = 9.0 Hz), 8.26 (1H, d, J = 9.0 Hz), 8.03 (1H, d, J = 16.5 Hz), 7.81 (1H, s), 7.66 (2H, d, J = 8.5 Hz), 7.51 (1H, t, J = 8.5 Hz), 7.09 (1H, d, J = 16.5 Hz), 7.09 (1H, s), 6.17 (2H, s), 4.92 (2H, t, J = 6.5 Hz), 4.06 (3H, s), 3.21 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 163.0, 150.2, 147.9, 144.3, 140.5, 139.9, 138.4, 134.4, 133.3, 132.1, 130.5, 129.6, 128.0, 127.4, 127.0, 126.0, 124.7, 121.3, 120.9, 120.4, 108.6, 105.6, 102.2, 57.4, 55.5, 26.3; HRMS (ESI) calcd for C28H20Cl3NO5 [M–Cl]+ 520.07130, found 520.07867.
9-(((2E,4E)-5-(benzo[d][1,3]dioxol-5-yl)penta-2,4-dienoyl)oxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (33): Yellow powder (0.219 g, 39%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.85 (1H, s), 8.98 (1H, s), 8.27 (1H, d, J = 9.0 Hz), 8.22 (1H, d, J = 9.0 Hz), 7.78 (1H, s), 7.69 (1H, ddd, J = 15.0, 7.5, 3.0 Hz), 7.30 (1H, d, J = 2.0 Hz), 7.20 (1H, d, J = 15.5 Hz), 7.16 (1H, d, J = 15.5 Hz), 7.09 (1H, dd, J = 8.0, 1.5 Hz), 7.07 (1H, s), 6.96 (1H, d, J = 7.5 Hz), 6.41 (1H, d, J = 15.5 Hz), 6.15 (2H, s), 6.06 (2H, s), 4.90 (2H, t, J = 6.0 Hz), 4.02 (3H, s), 3.19 (2H, t, J = 6.5 Hz); 13C NMR (125 MHz, DMSO-d6): 164.0, 160.8, 156.6, 150.8, 150.2, 148.8, 148.3, 148.0, 144.7, 142.6, 138.4, 133.7, 133.2, 131.2, 130.6, 126.1, 124.8, 124.1, 121.6, 120.9, 120.6, 117.7, 108.9, 108.7, 106.2, 106.0, 102.6, 101.8, 57.4, 55.6, 26.4; HRMS (ESI) calcd for C31H24ClNO7 [M–Cl]+ 522.15473, found 522.1556.
9-(2-(2,6-dichlorophenyl)acetoxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (34): Yellow powder (0.018 g, 3%); 1H NMR (500 MHz, DMSO-d6): δ ppm 9.96 (1H, s), 9.01 (1H, s), 8.24 (2H, s), 7.78 (1H, s), 7.57 (2H, d, J = 8.0 Hz), 7.42 (1H, t, J = 8.0 Hz), 7.09 (1H, s), 6.16 (2H, s), 4.96 (2H, s), 4.59 (2H, s), 3.98 (3H, s), 3.24 (2H, s); 13C NMR (125 MHz, DMSO-d6): 167.0, 150.5, 150.1, 147.8, 144.2, 138.3, 135.7, 133.2, 133.0, 131.0, 130.5, 130.4, 128.6, 127.2, 126.0, 121.0, 120.8, 120.4, 108.6, 105.7, 102.3, 57.2, 55.6, 36.2, 26.3; HRMS (ESI) calcd for C27H20Cl3NO5 [M–Cl]+ 508.07130, found 508.0720.
9-(2,2-diphenylacetoxy)-10-methoxy-5,6-dihydro-[1,3]dioxolo[4,5-g]isoquinolino[3,2-a]isoquinolin-7-ium chloride (35): Yellow powder (0.063 g, 11%); 1H NMR (500 MHz, DMSO-d6): δ ppm 10.04 (1H, s), 9.04 (1H, s), 8.22 (2H, m), 7.79 (1H, s), 7.57 (4H, m), 7.42 (4H, m), 7.33 (2H, m), 7.08 (1H, s), 6.16 (2H, s), 6.09 (1H, s), 4.99 (2H, t, J = 5.5 Hz), 4.00 (3H, s), 3.22 (2H, t, J = 6.0 Hz); 13C NMR (125 MHz, DMSO-d6): 169.6, 150.4, 150.0, 147.8, 144.3, 138.6, 138.1, 133.3, 133.0, 132.6, 130.9, 129.0, 128.7, 126.1, 124.2, 121.0, 120.6, 120.3, 115.1, 108.4, 105.6, 102.2, 57.0, 55.4, 25.6; HRMS (ESI) calcd for C33H26ClNO5 [M–Cl]+ 516.18055, found 516.1815.
α-Glucosidase inhibitory assay
Berberrubine derivatives will be assayed for yeast α-glucosidase inhibitory activity. The protocol described by Ramadhan et al. will be used37. Briefly, yeast α-glucosidase (0.1 U/mL) and substrate (1 mM p-nitrophenyl-α-D-glucopyranoside) were dissolved in 0.1 M phosphate buffer (pH 6.9). A 10 μL test sample was pre-incubated with α-glucosidase (40 μL) at 37 °C for 10 min. A substrate solution (50 μL) was then added to the reaction mixture and incubated at 37 °C for an additional 20 min, and terminated by adding 1 M Na2CO3 solution (100 μL). Enzymatic activity was quantified by measuring the absorbance at 405 nm (ALLSHENG AMR-100 microplate reader). The percentage inhibition of activity was calculated as follows: % Inhibition = [(A0 − A1)/A0] × 100, where: A0 is the absorbance without the sample; A1 is the absorbance with the sample. The IC50 value was deduced from the plot of % inhibition versus the concentration of the test sample. Acarbose was used as standard control and the experiment was performed in triplicate.
Kinetic study of α-glucosidase inhibition
The mode of inhibition of α-glucosidase was determined from Lineweaver–Burk plots. The inhibition type was determined using various concentrations of p-NPG substrate in the absence or presence of compounds at different concentrations. The Ki value was determined from secondary plots of slope versus [I].
Cells culture
HEK-293 cells (ATCC® CRL-1573™) were maintained in growth medium consisting of DMEM (Gibco®, Langley, OK) supplemented with 10% fetal bovine serum (Gibco®, Langley, OK), 100 I.U./ml penicillin (Bio Basic Canada®, Ontario, CA), 100 μg/ml streptomycin (Bio Basic Canada®, Ontario, CA), and 10 mM HEPES (4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid) (Sigma Aldrich®, St. Louis, MO) at 37 °C under 5% CO2.
Cytotoxicity study
The cytotoxicity of the active compounds (9, 26, 28 and 33) was tested with HEK-293 cells. The cells were seeded at 1 × 104 cells per well into 96-well plates in growth medium and incubated overnight. The compounds were prepared at the indicated concentrations in dimethylsulfoxide (DMSO) and added to the cells. Cells were incubated for 48 h before analyzing the cell viability using CellTiter 96® AQueous One Solution Cell Proliferation Assay kit (Promega, USA) according to manufacturer’s protocol. The plate was read at the A490 by VICTORTM X3 microplate reader (PerkinElmer, USA). The 1%DMSO treated cells as a positive control (100% viability). The CC50 values were calculated from nonlinear regression analysis. Results were reported as means and standard error mean (SEM) from three biological independent experiments.
Computational details
The 3D structure of α-glucosidase MAL12 derived from the alphafold2 database (UniProt ID: P5334150) was used as a protein receptor. The protonation state of protein was assigned at pH 7.0 using PDB2PQR51 and then the structure was minimized by SANDER52. The 3D structures of potent compounds 9, 26, 28, and 33 were constructed by gaussview 6.053, and subsequently optimized with density function of theory (DFT) with B3LYP/ 6-31G* basis set using gaussian 1653. The antechamber and parmchk2 tools were adopted to parameterize the ligand force field using GAFF54, while their partial charges were assigned using the RESP charge fitting method. To predict the binding pose and affinity of potent compounds at the active site of α-glucosidase, a molecular docking was applied using AutoDock Vina 1.2.3 software55. Then, molecular dynamics (MD) simulation for these complexes was performed using the AMBER20 package program52. In brief, the ligand/protein complex was prepared by adding all missing hydrogen atoms, neutralizing by ions, and solvating in the TIP3P56 water model with 12 Å from the protein surface by the tLeap module52. Then, the complex was slowly minimized and structurally relaxed by harmonic potentials, as previously described in our studies57,58,59. Each system was heated to 300 K for 20 ps with the canonical ensemble under periodic boundary conditions and then simulated for 300 ns. The production phase extracted from the last 100 ns-MD trajectories was selected to calculate the binding free energy using MM/GBSA and QM-MM/GBSA60 with three different theories AM1, PM3, and PM6 treated on ligand molecule. The key residues that interacted with the most potent compound were evaluated based on MM/GBSA method. The 3D structures and 2D binding interactions were visualized by UCSF Chimera V1.1661 and BIOVIA Discovery Studio Visualizer62.
The distribution functions of solvent were produced by 3D-RISM theory. The 3D-RISM equation was solved with the Kovalenko-Hirata closure63. The system temperature and the density of solvent water were set at 300 K and 1.0 g/cm3. The Lennard–Jones parameters for solute molecules were taken from the GAFF64 parameter set assigned by antechamber. The TIP3P56 arranged for the geometrical and potential parameters for the solvent water was employed with modified hydrogen parameters (σ = 0.4 Å and ε = 0.046 kcal/mol). The 3D grid spacing with 0.5 Å was assigned for the number of grid points with 128. The calculation was performed using the RISMiCal program package65,66,67.
Data availability
All data generated or analyzed during this study are included in this published article and its Supplementary Information files.
References
Nguyen, D. V., Shaw, L. C. & Grant, M. B. Inflammation in the pathogenesis of microvascular complications in diabetes. Front. Endocrinol. 3, 170 (2012).
edition, I. D. A. T. Diabetes around the world in 2021, https://diabetesatlas.org/ (2021).
Organization, W. H. Diabetes, https://www.who.int/news-room/fact-sheets/detail/diabetes (2022).
Lebovitz, H. E. Alpha-glucosidase inhibitors. Endocrinol. Metab. Clin. North Am. 26, 539–551 (1997).
Chai, T.-T., Kwek, M.-T., Ong, H.-C. & Wong, F.-C. Water fraction of edible medicinal fern Stenochlaena palustris is a potent α-glucosidase inhibitor with concurrent antioxidant activity. Food Chem. 186, 26–31 (2015).
Ghani, U. Re-exploring promising α-glucosidase inhibitors for potential development into oral anti-diabetic drugs: Finding needle in the haystack. Eur. J. Med. Chem. 103, 133–162 (2015).
Ghani, U. et al. The 4-(dimethylaminoalkyl) piperazine inhibitors of α-glucosidase: allosteric enzyme inhibition and identification of interacting chemical groups. Turk. J. Chem. 46, 1484–1492 (2022).
Ghani, U. et al. Dithiocarbamate derivatives inhibit α-glucosidase through an apparent allosteric site on the enzyme. Chem. Biol. Drug Des. 98, 283–294 (2021).
Ghani, U. et al. Thiazole inhibitors of α-glucosidase: Positional isomerism modulates selectivity, enzyme binding and potency of inhibition. Comput. Biol. Chem. 98, 107647 (2022).
Battu, S. K. et al. Physicochemical characterization of berberine chloride: a perspective in the development of a solution dosage form for oral delivery. Aaps Pharmscitech. 11, 1466–1475 (2010).
Gao, W.-W. et al. Discovery of 2-aminothiazolyl berberine derivatives as effectively antibacterial agents toward clinically drug-resistant Gram-negative Acinetobacter baumanii. Eur. J. Med. Chem. 146, 15–37 (2018).
Sobolova, K. et al. Discovery of novel berberine derivatives with balanced cholinesterase and prolyl oligopeptidase inhibition profile. Eur. J. Med. Chem. 203, 112593 (2020).
Wang, L. et al. Synthesis of disaccharide modified berberine derivatives and their anti-diabetic investigation in zebrafish using a fluorescence-based technology. Org. Biomol. Chem. 18, 3563–3574 (2020).
Guo, Z., Sun, H., Zhang, H. & Zhang, Y. Anti-hypertensive and renoprotective effects of berberine in spontaneously hypertensive rats. Clin. Exp. Hypertens. 37, 332–339 (2015).
Wang, Y. X. et al. Synthesis and biological evaluation of new berberine derivatives as cancer immunotherapy agents through targeting IDO1. Eur. J. Med. Chem. 143, 1858–1868 (2018).
Wang, Y.-X. et al. Synthesis and identification of novel berberine derivatives as potent inhibitors against TNF-α-induced nf-κb activation. Molecules 22, 1257–1271 (2017).
Cicero, A. & Ertek, S. Metabolic and cardiovascular effects of berberine: from preclinical evidences to clinical trial results. Clin. Lipidol. 4, 553–563 (2009).
Zuo, F., Nakamura, N., Akao, T. & Hattori, M. Pharmacokinetics of berberine and its main metabolites in conventional and pseudo germ-free rats determined by liquid chromatography/ion trap mass spectrometry. Drug Metab. Dispos. 34, 2064–2072 (2006).
Yu, F. et al. Monodisperse microparticles loaded with the self-assembled berberine-phospholipid complex-based phytosomes for improving oral bioavailability and enhancing hypoglycemic efficiency. Eur. J. Pharm. Biopharm. 103, 136–148 (2016).
Chen, W. et al. Bioavailability study of berberine and the enhancing effects of TPGS on intestinal absorption in rats. Aaps Pharmscitech. 12, 705–711 (2011).
Pan, G.-Y. et al. The antihyperglycaemic activity of berberine arises from a decrease of glucose absorption. Planta Med. 69, 632–636 (2003).
Li, Z.-Q. et al. Berberine acutely inhibits the digestion of maltose in the intestine. J. Ethnopharmacol. 142, 474–480 (2012).
Pan, G. et al. Inhibitory action of berberine on glucose absorption. Acta Pharm. Sin. 38, 911–914 (2003).
Liu, L. et al. Berberine attenuates intestinal disaccharidases in streptozotocin-induced diabetic rats. Pharmazie 63, 384–388 (2008).
Liu, L. et al. Berberine suppresses intestinal disaccharidases with beneficial metabolic effects in diabetic states, evidences from in vivo and in vitro study. Naunyn Schmiedebergs Arch. Pharmacol. 381, 371–381 (2010).
Lo, C.-Y. et al. Synthesis and anticancer activity of a novel series of 9-O-substituted berberine derivatives: A lipophilic substitute role. Bioorg. Med. Chem. Lett. 23, 305–309 (2013).
Park, K. D. et al. Synthesis of 13-(substituted benzyl) berberine and berberrubine derivatives as antifungal agents. Bioorg. Med. Chem. Lett. 16, 3913–3916 (2006).
Fu, S. et al. Discovery of mitochondria-targeting berberine derivatives as the inhibitors of proliferation, invasion and migration against rat C6 and human U87 glioma cells. MedChemComm 6, 164–173 (2015).
Zhang, S. et al. Synthesis and hypoglycemic activity of 9-O-(lipophilic group substituted) berberine derivatives. Bioorg. Med. Chem. Lett. 26, 4799–4803 (2016).
Ding, S.-M. et al. Novel oxazolxanthone derivatives as a new type of α-glucosidase inhibitor: synthesis, activities, inhibitory modes and synergetic effect. Bioorg. Med. Chem. 26, 3370–3378 (2018).
Ye, G.-J. et al. Design and synthesis of novel xanthone-triazole derivatives as potential antidiabetic agents: α-Glucosidase inhibition and glucose uptake promotion. Eur. J. Med. Chem. 177, 362–373 (2019).
Li, G.-L. et al. Synthesis of 3-acyloxyxanthone derivatives as α-glucosidase inhibitors: A further insight into the 3-substituents’ effect. Bioorg. Med. Chem. 24, 1431–1438 (2016).
Kan, L. et al. Inhibition of α-glucosidases by tea polyphenols in rat intestinal extract and Caco-2 cells grown on Transwell. Food Chem. 361, 130047 (2021).
Kamiyama, O. et al. In vitro inhibition of α-glucosidases and glycogen phosphorylase by catechin gallates in green tea. Food Chem. 122, 1061–1066 (2010).
Chantarasriwong, O., Jang, D. O. & Chavasiri, W. A practical and efficient method for the preparation of sulfonamides utilizing Cl3CCN/PPh3. Tetrahedron Lett. 47, 7489–7492 (2006).
Kasemsuknimit, A., Satyender, A., Chavasiri, W. & Jang, D.-O. Efficient amidation and esterification of phosphoric acid using Cl3CCN/Ph3P. Bull. Korean Chem. Soc. 32, 3486–3488 (2011).
Ramadhan, R. & Phuwapraisirisan, P. New arylalkanones from Horsfieldia macrobotrys, effective antidiabetic agents concomitantly inhibiting α-glucosidase and free radicals. Bioorg. Med. Chem. Lett. 25, 4529–4533 (2015).
Liu, Y., Ma, L., Chen, W.-H., Wang, B. & Xu, Z.-L. Synthesis of xanthone derivatives with extended π-systems as α-glucosidase inhibitors: Insight into the probable binding mode. Bioorg. Med. Chem. 15, 2810–2814 (2007).
Liu, Y. et al. Synthesis and pharmacological activities of xanthone derivatives as α-glucosidase inhibitors. Bioorg. Med. Chem. 14, 5683–5690 (2006).
Cai, C.-Y. et al. Analogues of xanthones—Chalcones and bis-chalcones as α-glucosidase inhibitors and anti-diabetes candidates. Eur. J. Med. Chem. 130, 51–59 (2017).
Liu, Y. et al. Synthesis, inhibitory activities, and QSAR study of xanthone derivatives as α-glucosidase inhibitors. Bioorg. Med. Chem. 16, 7185–7192 (2008).
Li, G.-L. et al. Toward potent α-glucosidase inhibitors based on xanthones: A closer look into the structure–activity correlations. Eur. J. Med. Chem. 46, 4050–4055 (2011).
Proença, C. et al. Inhibitory activity of flavonoids against human sucrase-isomaltase (α-glucosidase) activity in a Caco-2/TC7 cellular model. Food Funct. 13, 1108–1118 (2022).
Liu, Y. et al. α-Glucosidase inhibitors from Chinese bayberry (Morella rubra Sieb. et Zucc.) fruit: molecular docking and interaction mechanism of flavonols with different B-ring hydroxylations. RSC Adv. 10, 29347–29361 (2020).
Son, H.-U. et al. Effects of synergistic inhibition on α-glucosidase by phytoalexins in soybeans. Biomolecules 9, 828 (2019).
Askarzadeh, M. et al. Design, synthesis, in vitro α-glucosidase inhibition, docking, and molecular dynamics of new phthalimide-benzenesulfonamide hybrids for targeting type 2 diabetes. Sci. Rep. 12, 10569 (2022).
He, C. et al. Interaction mechanism of flavonoids and α-glucosidase: Experimental and molecular modelling studies. Foods 8, 355 (2019).
Kovalenko, A. & Hirata, F. Three-dimensional density profiles of water in contact with a solute of arbitrary shape: A RISM approach. Chem. Phys. Lett. 290, 237–244 (1998).
Phanich, J. et al. A 3D-RISM/RISM study of the oseltamivir binding efficiency with the wild-type and resistance-associated mutant forms of the viral influenza B neuraminidase. Protein Sci. 25, 147–158 (2016).
Teste, M.-A., François, J. M. & Parrou, J.-L. Characterization of a new multigene family encoding isomaltases in the yeast Saccharomyces cerevisiae, the IMA family. J. Biol. Chem. 285, 26815–26824 (2010).
Dolinsky, T. J., Nielsen, J. E., McCammon, J. A. & Baker, N. A. PDB2PQR: an automated pipeline for the setup of Poisson–Boltzmann electrostatics calculations. Nucleic Acids Res. 32, W665–W667 (2004).
D.A. Case et al AMBER 2020 (2020).
Frisch, M. J. et al Gaussian 16 Rev. C.01 (2016).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Eberhardt, J., Santos-Martins, D., Tillack, A. F. & Forli, S. AutoDock Vina 1.2.0: New docking methods, expanded force field, and python bindings. J. Chem. Inf. Model. 61, 3891–3898 (2021).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Nutho, B. et al. Discovery of C-12 dithiocarbamate andrographolide analogues as inhibitors of SARS-CoV-2 main protease: In vitro and in silico studies. Comput. Struct. Biotechnol. J. 20, 2784–2797 (2022).
Hengphasatporn, K. et al. Halogenated baicalein as a promising antiviral agent toward SARS-CoV-2 main protease. J. Chem. Inf. Model. 62, 1498–1509 (2022).
Hengphasatporn, K. et al. Promising SARS-CoV-2 main protease inhibitor ligand-binding modes evaluated using LB-PaCS-MD/FMO. Sci. Rep. 12, 17984 (2022).
Mishra, S. K. & Koča, J. Assessing the performance of MM/PBSA, MM/GBSA, and QM–MM/GBSA approaches on protein/carbohydrate complexes: effect of implicit solvent models, QM methods, and entropic contributions. J. Phys. Chem. B 122, 8113–8121 (2018).
Pettersen, E. F. et al. UCSF Chimera—A visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Biovia, D. S. Discovery Studio Visualizer (Dassault Systèmes, 2021).
Kovalenko, A. & Hirata, F. A replica reference interaction site model theory for a polar molecular liquid sorbed in a disordered microporous material with polar chemical groups. J. Chem. Phys. 115, 8620–8633 (2001).
Wang, J., Wang, W., Kollman, P. A. & Case, D. A. Automatic atom type and bond type perception in molecular mechanical calculations. J. Mol. Graph. 25, 247–260 (2006).
Yoshida, N. in IOP Conference Series: Materials Science and Engineering. 012062 (IOP Publishing).
Maruyama, Y. & Hirata, F. Modified Anderson method for accelerating 3D-RISM calculations using graphics processing unit. J. Chem. Theory Comput. 8, 3015–3021 (2012).
Maruyama, Y. et al. Massively parallel implementation of 3D-RISM calculation with volumetric 3D-FFT. J. Comput. Chem. 35, 1347–1355 (2014).
Acknowledgements
This research project is supported by the Second Century Fund (C2F), Chulalongkorn University, the sponsorship of a high-performance computing infrastructure project (Grant Number hp200157), Kakehashi project of Tsukuba Innovation Arena (TIA) collaborative research program, CREST JST, Japan (Grant Number JP20338388), and partially by the AMED, Japan (Grant Number JP21ae0101047h0001) to Y.S.
Author information
Authors and Affiliations
Contributions
Duy Vu Nguyen: synthesis of target molecules, analysis the structures, alpha-glucosidase testing, analysis the results Ade Danova: synthesis of some target molecules and characterize by spectroscopy Kowit Hengphasatporn, Ryo Fujiki, Yasuteru Shigeta, Thanyada Rungrotmongkol: computational study Aphinya Suroengrit and Siwaporn Boonyasuppayakorn: biological study Warinthorn Chavasiri: design the target molcules and discussion the experiment results All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nguyen, D.V., Hengphasatporn, K., Danova, A. et al. Structure—yeast α-glucosidase inhibitory activity relationship of 9-O-berberrubine carboxylates. Sci Rep 13, 18865 (2023). https://doi.org/10.1038/s41598-023-45116-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-45116-0
This article is cited by
-
Synthesis and Biological Activity Evaluation of Two Derivatives of 7-Aminocephalosporanic Acid
Chemistry Africa (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.