Abstract
Although recent studies have revealed the association between the human microbiome especially gut microbiota and longevity, their causality remains unclear. Here, we assess the causal relationships between the human microbiome (gut and oral microbiota) and longevity, by leveraging bidirectional two-sample Mendelian randomization (MR) analyses based on genome-wide association studies (GWAS) summary statistics of the gut and oral microbiome from the 4D-SZ cohort and longevity from the CLHLS cohort. We found that some disease-protected gut microbiota such as Coriobacteriaceae and Oxalobacter as well as the probiotic Lactobacillus amylovorus were related to increased odds of longevity, whereas the other gut microbiota such as colorectal cancer pathogen Fusobacterium nucleatum, Coprococcus, Streptococcus, Lactobacillus, and Neisseria were negatively associated with longevity. The reverse MR analysis further revealed genetically longevous individuals tended to have higher abundances of Prevotella and Paraprevotella but lower abundances of Bacteroides and Fusobacterium species. Few overlaps of gut microbiota-longevity interactions were identified across different populations. We also identified abundant links between the oral microbiome and longevity. The additional analysis suggested that centenarians genetically had a lower gut microbial diversity, but no difference in oral microbiota. Our findings strongly implicate these bacteria to play a role in human longevity and underscore the relocation of commensal microbes among different body sites that would need to be monitored for long and healthy life.
Similar content being viewed by others
Introduction
Population aging is accelerating, and it is of great significance to identify the biological factors that affect aging and how to extend life to achieve a healthy and long life. The heritability of a human’s lifespan is reported to be only 25%1,2, which means the human lifespan is not only determined by the genome but also affected by both the human external environment and internal environment (microbiome) as well as their interactions3,4. Increasing evidence indicates that there is a link between gut microbes and longevity5,6,7,8,9. A study of Italian centenarians (99–104 years old) and ultracentenarians (105–109 years old) showed that there is a different core flora that decreases with aging dominated by Ruminococcaceae, Lachnospiraceae, and Bacteroides6. Another independent study of Chinese centenarians showed that aging is negatively related to abundances of Bacteroides vulgatus, Ruminococcus sp.5139BFAA, and Clostridium sp. AT5 in the gut7. These studies have reported that some specific intestinal microbes are significantly associated with longevity, however, the causal relationships between them remain unexplored. In addition, the oral microbiome influenced not only oral but also systemic health and disease10,11,12. Few studies have focused on the associations between the oral microbiome and longevity. Therefore, it is particularly important to explore in detail which specific intestinal or oral microbes link to longevity, and to explore the functions of these intestinal microbes, to achieve aging intervention and prolong lifespan.
Mendelian randomization (MR)13 has been developed to investigate the potential causal relationship between exposure and outcome by using genetic variants as instrumental variables (IVs) in genetic epidemiology research. Several MR analyses have investigated the biomarkers (genes/proteins/metabolites/diseases) contributing to longevity or lifespan14,15,16,17,18. The microbiota influencing longevity remains unknown. Twin-based heritability estimation19,20 and increasing microbiome GWAS analyses21,22,23,24,25 consistently indicated the genetic influence on gut microbiota. Our recent microbial GWAS analyses have identified genetic determinants for both gut21 and oral26 microbial features, which enable us to investigate potential causations between the human microbiome and longevity.
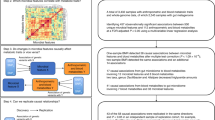
Here, we performed bidirectional two-sample MR analyses to dissect the potential causal links between the human microbiome and longevity. Specifically, the forward MR was performed using gut-, salivary-, tongue dorsum- microbiome, and microbial α-diversity from the 4D-SZ cohort (N = 2984) as exposures and longevity from the CLHLS phase I dataset (N = 4477) as the outcome, to identify the microbial features that may affect longevity. The reverse MR was performed using the longevity phase I dataset (N = 4477) as discovery sample exposure and the longevity phase II dataset (N = 6548) as replication sample exposure, together with gut-, salivary-, tongue dorsum- microbiome, and microbial α-diversity as outcomes, to identify the causal effects of genetically predicted longevity phenotype on the microbiota. An overview of the study design and results was presented in Supplementary Fig. 1. Additionally, we performed the main MR analyses in Chinese populations and replicated the MR results with that of in European populations. In this large-scale MR study, we discovered a large number of potential causal correlations between the gut, oral microbiota and longevity, suggesting the human microbiome is a promising alternative for microbiome-based management of human healthy aging.
Results
Causal relationships between the gut microbiome and longevity
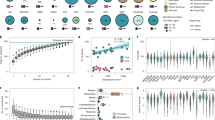
We first investigate causal relationships between the gut microbiome and longevity by performing a bidirectional two-sample MR analysis using GWAS summary data of the gut microbiome (500 independent microbial features; N = 1539) obtained from the 4D-SZ cohort21, together with longevity GWAS from the CLHLS cohort27 (N = 4477) (Supplementary Fig. 1, “Methods” section). Descriptions of samples and cohorts were presented in Supplementary Table 1. In the forward MR, 23 gut microbial features were found to influence longevity (p < 0.05; Fig. 1; Supplementary Table 2). Family Coriobacteriaceae (β = 0.21, p = 0.008), phylum Tenericutes (β = 0.089, p = 0.008), genus Oxalobacter (β = 0.09, p = 0.0096), microbial function/module MF0039:methionine degradation II (β = 0.32, p = 0.014) and species Lactobacillus amylovorus (a probiotic28 altering body adiposity; β = 0.09, p = 0.014) were the top five strongest positively associated with longevity. However, some features like Methylobacillus flagellates (β = − 0.094, p = 0.003), Neisseria genus, Streptococcus genus and its species (the dental caries pathogen S. mutans29 and unclassified Streptococcus sp. Oral taxon 071), and Fusobacterium nucleatum (colorectal cancer (CRC) pathogen30; β = − 0.118, p = 0.036) were negatively associated with longevity. These causal relationships were robust with p < 0.05 in at least one method when the other five two-sample MR tests were performed (“Methods” section), and there was no evidence of horizontal pleiotropy (both PMR-PRESSO.Global.test > 0.05 and PMR_egger_intercept > 0.05; Supplementary Table 2).
Bidirectional two-sample MR analyses identified causal relationships between the gut microbiome and longevity. (a) The barplot represented the causal effects of 23 specific gut microbial features on longevity (p < 0.05), as estimated using the GCTA-GSMR approach in the forward MR analysis. The MR effect size estimate (bxy) was plotted. (b) The barplot represented the causal effects of longevity on 78 gut microbial features in the reverse MR analysis (p < 0.05). Discovery used the longevity phase I dataset (N = 4477) as exposure. Replication used the longevity phase II dataset (N = 6548) as exposure. Both used the gut microbiome dataset (N = 1539) as the outcome. *p < 0.05, **p < 0.01, and ***p < 0.001. s_ denotes species, g_ denotes genus, f_ denotes family, p_ denotes phylum and MF denotes microbial function/module.
In the reverse MR analysis to use longevity as exposure (N = 4477 with 45 independent IVs; Supplementary Fig. 1), we found longevity was associated with relative abundances of 78 microbial features (p < 0.05). Notably, genetically predicted longevous individuals tended to have increased abundances of some features like the genus Prevotella and its seven species such as P. copri and P. multiformis, Paraprevotella Mitsuokella, and MF0047:glutamine degradation II, but decreased abundances of other features such as Bacteroides and its nine species such as B. thetaiotaomicron and B. uniformis, phylum Fusobacteria and its species (F. varium and F. ulcerans), genus Clostridium and its species (C. bolteae, C. ramosum, unclassified Clostridium sp. HGF2, and C. leptum). In general, Bacteroides and Escherichia were more prevalent than Prevotella after 50 years old20,31. Bacteroides and Fusobacteria were also major taxa that metabolize pectin, implicated in body fat and hormone levels32.
70 of the 78 causal relationships were replicated in the same direction of effect, of which 17 were well validated with p < 0.05 while applying the reverse MR analyses to another independent longevity dataset33 as exposure (N = 6548 with 30 independent IVs; Supplementary Table 3), suggesting these causalities were robust. We confirmed the correlation of longevity with increased abundances of Prevotella and its species (Fig. 2) but decreased abundances of Bacteroides and its species (Supplementary Fig. 3), in line with previous observational studies34. Additionally, longevity was related to increased abundances of microbial pathways such as MF0047: glutamine degradation II, MF0098: hydrogen metabolism, and MF0051: arginine degradation I, but the decreased abundances of pathways like MF0092: lactate production, MF0040: proline degradation, F0070: pentose phosphate pathway (oxidative_phase) and MF0095: propionate production III, which were all replicated in the MR analyses when using two independent longevity cohorts as exposure datasets.
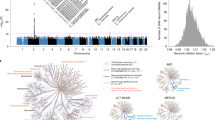
Comparisons of causalities between Chinese and European populations. (a–c) represented the causalities from gut microbiome to longevity by comparing the effect sizes of all available causalities between Chinese and MiBioGen (mainly European, (a)), Finnish (b), and Dutch (c), respectively. (d–f) represented the causalities from longevity to gut microbiome by comparing the effect sizes of all available causalities between Chinese and MiBioGen (mainly European, (d)), Finnish (e), and Dutch (f), respectively. Spearman test was used with correlation coefficients and p-value marked. The red, blue, purple, and green dots indicated the causalities were significant in Chinese, MiBioGen, Dutch, and Finnish, respectively (p < 0.05). The black dots indicated the causalities were not significant in either of the two compared populations. The red dots with a green rectangular box in the Dutch panel represented the shared MR by the Chinese and Dutch.
Few overlaps of causalities between the Chinese and European populations
To confirm if the causalities identified in the Chinese were replicated in other populations, we systematically compared the correlations of the gut microbiome and longevity with those of the European cohorts. We performed bidirectional two-sample MR analyses by leveraging GWAS summary statistics of the gut microbiome from MiBioGen (mainly European populations)22, Dutch35, and Finnish36 cohorts, together with the largest meta-analysis of longevity GWAS of European ancestry37. Overall, MR results from the MiBioGen, Dutch, and Finnish cohorts (Supplementary Tables 4–6) showed clear differences from those in the Chinese cohort. Beta effects of the causalities available between Chinese and the three European cohorts consistently showed negative correlations, except the only one positive correlation of causality from longevity to microbiome between Chinese and the MiBioGen cohort (Fig. 2; Supplementary Figs. 4, 5).
Despite the overall negative correlations of the causalities between the Chinese and European populations (Fig. 2; Supplementary Tables 4–6), we found two shared causalities: longevity was causally linked to an increased abundance of Lactococcus lactis, but the decreased abundance of Ruminococcus in both Chinese and Finnish (Fig. 2f). Oral L. lactis enriched in healthy people in comparison with RA10, could inhibit bacteria such as Streptococcus38, and could produce GABA39. We also observed several overlapped causalities among the three European cohorts. Bacteroidia was positively associated with longevity in the Dutch cohort and so with two Bacteroides species (B. sp003545565 and B. ovatus) in the Finnish cohort. Sutterellaceae in the Dutch cohort and Sutterella in the Finnish cohort were negatively linked to longevity. We called for further validation of these shared microbiome-longevity interactions and the discovery of more overlapped causalities across populations in the future, given mainly 16s rRNA sequencing classifying into only microbial genera but not species was performed in the European populations.
Causalities between the salivary microbiome and longevity
We observed 88 significant associations of the salivary microbiome with longevity (p < 0.05; Supplementary Table 7), among which 23 correlations with p < 0.01 were presented in Fig. 3a. Several microbes such as Streptococcus mitis AZ (β = 0.623, p = 0.0012), Pauljensenia odontolyticus A (β = 0.399, p = 0.0013), Treponema C lecithinolyticum (β = 0.458, p = 0.0035), and Streptococcus uSGB 1603 (β = 0.625, p = 0.0045) linked to increased longevity odds. Streptococcus uSGB 1603 reduced risks of uterine fibroids and gastric cancer in the diseases’ MR analysis by integrating with biobank Japan (BBJ) diseases data (Supplementary Table 8). Oribacterium sinus which has been reported depleted in the COVID-19 patients40 and potentially negatively correlated with congestive heart failure (Supplementary Table 8), exhibited a positive association with longevity. Reversely, some oral microbiota such as Neisseria flavescens (β = − 0.565, p = 0.0015), Lachnoanaerobaculum sp000287675 (β = − 0.523, p = 0.0017), Solobacterium uSGB 2890 (β = − 0.780, p = 0.0017), Lancefieldella uSGB 3043 (β = − 0.477, p = 0.0032) decreased longevity odds. Lachnoanaerobaculum sp000287675 increased the risk of chronic obstructive pulmonary disease, biliary tract cancer, and osteoporosis in the disease MR analysis. Solobacterium uSGB 2890 increased the risks of lung cancer and asthma (Supplementary Table 8). These findings suggest that the microbiota may influence longevity by acting on disease risks.
Bidirectional two-sample MR analyses identified causal relationships between the salivary microbiome and longevity. (a) The barplot represented the causal effects of 23 specific salivary microbial features on longevity (p < 0.01), as estimated using the GCTA-GSMR approach in the forward MR analysis. The MR effect size estimate (bxy) was plotted. (b) The barplot represented the causal effects of longevity on 16 salivary microbial features in the reverse MR analysis (p < 0.01). Discovery used the longevity phase I dataset (N = 4477) as exposure. Replication used the longevity phase II dataset (N = 6548) as exposure. Both used the gut microbiome dataset (N = 1539) as the outcome. **p < 0.01. Unclassified species-level cluster denotes uSGB, known species-level clusters denote kSGB and MF denotes microbial function/module.
The reverse MR analysis (testing the causal effect of longevity on saliva microbiome) identified 122 suggestive links with p < 0.05 and 16 of them with p < 0.01 (Fig. 3b; Supplementary Table 7). For example, longevity was correlated with higher abundances of Veillonellaceae F0422 uSGB 3578 (β = 0.024, p = 0.0095) whereas lower abundances of genus Aggregatibacter (β = − 0.023, p = 0.013) and a handful of its species such as A. segnis and A. sp000466335, family Porphyromonadaceae and its several species such as P. sp000467855 and P. endodontalis. Veillonellaceae might be better adapted to an environment with lower saliva flow10, and could produce carbon dioxide from lactate, which might be an important source when elderly people do not eat as much41. Longevous individuals exhibited a lower abundance of phylum Fusobacteria (β = − 0.023, p = 0.018) but higher Firmicutes (β = 0.019, p = 0.039), consistent with the above gut MR findings. Additionally, longevity was found to be negatively associated with the microbial function/module MF0025:tryptophan degradation for both saliva and gut, in agreement with a previous report that aging and the aging-associated decline in protein homeostasis could be delayed by inhibition of tryptophan degradation42.
Causalities between the tongue microbiome with longevity
We also examined the causal relationships between the tongue microbiome and longevity. 73 tongue microbiome features linked to longevity (Supplementary Table 9), among which 12 links with p < 0.01 were presented in Fig. 4a. For example, Streptococcus mitis AC (β = 0.531, p = 0.0005), Streptococcus uSGB 1472 (β = 0.464, p = 0.005), Prevotella nanceiensis (β = 0.481, p = 0.0048), and Catonella uSGB 1617 (β = 0.346, p = 0.0064) were found to be positively correlated with longevity. However, Streptococcus uSGB 1204 (β = − 0.513, p = 0.00096), Prevotella enoeca (β = − 0.629, p = 0.0019) and Campylobacter A uSGB 2909 (β = − 0.517, p = 0.006) were negatively correlated with longevity. Moreover, tongue dorsum microbial pathway MF0047: glutamine degradation II promoted longevity (β = 0.551, p = 0.00098), consistent with the above gut MR result that longevity was positively linked to fecal MF0047: glutamine degradation II. Aggregatibacter (β = − 0.383, p = 0.017) and A. segnis showed negative links with longevity, in line with the longevity influences on Aggregatibacter spp. as identified in the saliva MR result.
Bidirectional two-sample MR analyses identified causal relationships between the tongue dorsum microbiome and longevity. (a) The barplot represented the top 20 causal effects of specific tongue dorsum microbial features on longevity (p < 0.05), as estimated using the GCTA-GSMR approach in the forward MR analysis. The MR effect size estimate (bxy) was plotted. (b) The barplot represented the top 20 causal effects of longevity on tongue dorsum microbial features in the reverse MR analysis (p < 0.01). Discovery used the longevity phase I dataset (N = 4477) as exposure. Replication used the longevity phase II dataset (N = 6548) as exposure. Both used the gut microbiome dataset (N = 1539) as the outcome. *p < 0.05, **p < 0.01, and ***p < 0.001. Unclassified species-level cluster denotes uSGB, known species-level clusters denote kSGB and MF denotes microbial function/module.
Further, we also found 29 causal effects of longevity on the tongue dorsum microbiome (p < 0.05). Longevity was genetically linked to decreased abundances of Prevotella intermedia that was associated with increased risks of CRC43 and severe asthma44, HOT-345 uSGB 3129, F0040 uSGB 94, but increased abundances of Oribacterium uSGB 1411, Saccharimonadaceae UBA2866 uSGB 2333, Rothia uSGB 551, Streptococcus uSGB 2448, Bacteroidaceae uSGB 1496 (Fig. 4b, Supplementary Table 9).
Longevity associated with microbial α-diversity
We further investigate whether there is a potential correlation between microbial α-diversity and longevity. We identified 7, 21, and 11 independent host genetic variants associated with microbial α-diversity (Shannon diversity index) for the gut, saliva, and tongue dorsum at p < 10–5, respectively (Supplementary Table 10). Using these genetic variants as instrumental variables (IVs), we performed two-sample BMR analyses and observed no significant influences of microbial diversity across all three body niches on longevity (Supplementary Table 11). However, the reverse MR inference showed that genetically longevous individuals exhibited a lower Shannon diversity in the gut (Fig. 5). This causal relationship was robust when using two different longevity datasets as exposures (Fig. 5). Unexpectedly, this finding was in contrast to the previous papers45.
Bidirectional two-sample MR inference for the microbial α-diversity and longevity. (a,b). Forest plots represented the MR estimates and 95% CI values of the causal effects of longevity on microbial alpha diversity, as estimated in the discovery ((a), blue) and replication ((b), red) stages, respectively. (c,d) plotted the instruments of longevity phase I (discovery, (c)) and phase II (replication, (d)) datasets, respectively, showing the causal effect of longevity on lower gut microbial α-diversity. The x-axes in (c,d) show the SNP-exposure effect and the y-axes show the SNP-outcome effect. Error bars in (c,d) represent the standard errors.
Discussion
This is the first study using the MR approach to reveal causal relationships between the human microbiome (gut and oral) and longevity. We provided a good repository of resources containing a large number of potential correlations between the gut-, salivary-, tongue dorsum- microbiome, and longevity using bidirectional two-sample MR analyses. Those specific microbes that were identified significantly associated with longevity could be potential causal targets for study on healthy aging8.
Some of the identified longevity-associated microbes were found to correlate with diseases, indicating these microbes may affect longevity by mediating the risks of diseases. Coriobacteriaceae was significantly reduced in heart failure patients with reduced ejection fraction and cardiomyopathy46, suggesting this family may contribute to longevity by reducing the risk of cardiovascular diseases. Phylum Tenericutes was more abundant in the healthy group than in the MetS group in a Korean study47 and decreased the risk of lung cancer in our previous MR analysis21. Oxalobacter could potentially metabolize oxalate which hurts kidney functions48,49, was related to pro-longevity maybe also through reducing the risk of rheumatoid arthritis (RA) as reported in an MR study22. The probiotic Lactobacillus amylovorus altering body adiposity28 positively correlated with longevity, whereas the CRC pathogen Fusobacterium nucleatum30 negatively correlated with longevity. Consistently, our diseases’ two-sample MR analysis also supported a “microbiota—disease—longevity” axis (Supplementary Table 8). For example, the gut Tenericutes that decreased the risk of lung cancer in our previous MR analysis21 promoted longevity. The salivary Oribacterium sinus and Pauljensenia uSGB 2326 that were negatively associated with congestive heart failure promoted longevity. Some other pro-longevity features such as Streptococcus infantis I and Lancefieldella uSGB 1105 were also negatively associated with peripheral artery disease that caused a crude five-year death rate of 33.2%50. These findings further help us to understand the interactions among microbiota, disease, and longevity.
We not only excavated the specific microbiota that may affect longevity but also investigated which microbes the genetically predicted longevous individuals tended to colonize. Here, longevity was genetically related to a higher abundance of Prevotella but a lower abundance of Bacteroides. Prevotella was associated with plant-rich diets (high levels of complex carbohydrates and fruit and vegetable intake), whereas Bacteroides was dominated by the high-fat, high-sodium “westernized” diet51,52, indicating that long-lived individuals may genetically tend to prefer healthier diets with higher intakes of vegetables and lower intakes of chloride, sodium, and saturated fat food that caused higher Prevotella and lower Bacteroides abundances,respectively53,54.
There are two major limitations in this study. First, the identified causalities didn’t reach a multiple-testing corrected significance given that we performed MR tests on thousands of microbial features. Despite this, we were still able to replicate 17 of the 78 identified causalities from longevity to the gut microbiome by using an independent dataset as validation, supporting the robustness of these MR results. Our data-driven MR results also supported and confirmed some previously reported observational correlations or animal experimental results. For example, observational studies showed that longevity was associated with a lower abundance of Bacteroides6,34 but higher abundances of Firmicutes and Tenericutes34, Corprococcus6,55. The positive correlation between longevity and microbial pathway MF0047: glutamine degradation II in both tongue dorsum (β = 0.551, p = 0.00098) and gut (β = 0.021, p = 0.00022) agreed with previous findings that glutamine-supplemented mice also showed remarkably a full one-third extension of lifespan56. Longevity was positively correlated with the gut microbial pathway hydrogen metabolism, which extends the lifespan of C. elegans by reducing reactive oxygen species57. Second, the lack of individual-level data prevented us from directly comparing the contributions of the microbiome and other host factors (economic/behavioral/environmental) to longevity. Our previous study has reported that the most influential factors related to longevity included sex, education level, career status, smoking, drinking, and diseases such as hypertension, type 2 diabetes mellitus, cardiovascular disease, dyslipidemia, gastroenteric ulcer, arthritis or cholelithiasis33. These factors may have effects on longevity independent of microbiome profiles. We hope to collect more individual-level data including the microbiome, genome, socioeconomic, behavioral, and environmental factors in the longevity cohort to comprehensively explore the impact of these factors on longevity in the future.
There were two important points to notice in this and future related research. One is that causalities of the gut microbiome and longevity generally exhibited negative correlations among Chinese and European populations, which may be partly attributed to geography or ethnicity-specific variations in the microbiome compositions that could be affected by diets, lifestyles, and genetic backgrounds58,59. For example, Verrucomicrobia and its genus Akkermansia negatively correlated with longevity in the Netherlands but positively associated with longevity in the Chinese. Akkermansia muciniphila is an emerging probiotic that has been reported to promote longevity and slow down the aging process60,61. Lactobacillus and Streptococcus negatively contributed to longevity in the Chinese but showed no associations in the Europeans. The genetically predicted longevous individuals tend to exhibit higher abundances of Prevotella and Paraprevotella in the Chinese but not in the Europeans which was consistent with the observational study25, suggesting the higher prevalence of Prevotella in Asian countries might be determined by ancestral genomes in addition to the diet. Future studies in cohorts of different ethnicities with larger sample sizes were called for to further substantiate these findings.
Another is to distinguish the associations of microbiota across different body sites with longevity. Longevous individuals causally exhibited a lower abundance of phylum Fusobacteria and its species (F. nucleatum30 and F. varium21) in both gut and saliva. However, some bacteria that decreased longevity in the gut appeared to increase longevity in the mouth, where they normally belong. For example, Streptococcus was negatively associated with longevity in the gut, whereas, Streptococcus mitis, Streptococcus uSGB 1603, uSGB 1349, uSGB 2667, and Streptococcus infantis G were positively associated with longevity in the saliva. These findings also implied the similarities and differences in the gut and oral microbial compositions and diversities10,26,62. The transmission and colonization of commensal microbes among different body sites would need to be monitored for a long and healthy life.
In summary, our MR analysis illustrates the value of human genetic information to help prioritize human microbiome features for aging-related mechanistic studies and clinical trials.
Methods
Microbiome cohort
The microbiome data in this study were originally from the 4D-SZ cohort21,25,26,32,63, which is a multi-omics cohort, and all individuals in which were enlisted for the whole genome sequencing (WGS) and whole metagenomic sequencing (WMS).
We obtained GWAS summary statistics of 500 gut microbial features (401 taxa and 99 pathways) in 1539 individuals as previously detailly described21. Briefly, the gut microbial taxonomy profiles were generated by aligning high-quality sequencing reads to the integrated gene catalog (IGC)64 of gut using SOAP2 with the criterion of identity ≥ 95% and were next determined based on the aligned gene abundances. Gut metabolic modules or pathways profiles were built through an extensive review of the literature and metabolic databases, inclusive of MetaCyc65 and KEGG66, followed by expert curation and delineation of modules and alternative pathways.
For oral microbiome, we obtained 1665 tongue dorsum microbial features (1583 taxa and 82 pathways) in 2017 individuals and 1770 salivary microbial features (1685 taxa and 85 pathways) in 1915 individuals with available GWAS summary statistics as detailly described here26. Briefly, the oral microbial taxonomy profiles were constructed by aligning reads to a previously well-constructed high-quality oral genome catalog63 and were next determined based on the aligned contig abundances. Oral metabolic modules or pathways profiles were built through an extensive review of the literature and metabolic databases, inclusive of MetaCyc65 and KEGG66, followed by expert curation and delineation of modules and alternative pathways. All those GWAS summary statistics included ~ 10 M common variants (MAF > 0.005) in this 4D-SZ cohort.
Longevity cohort
The longevity GWAS data in this study comes from the Chinese Longitudinal Healthy Longevity Surveys (CLHLS), which were conducted in 1998, 2000, 2002, 2005, 2008, 2011, and 2014 in a randomly selected half of the counties and cities in 22 out of 31 provinces in China27,33,67,68. Two longevity datasets were included in the MR analysis. The longevity phase I dataset27 was the main longevity dataset that included 4477 samples (2178 centenarians and 2299 middle-aged controls) with 5.6 M imputed genotypes. The longevity phase II dataset33 contained 287 K longevity candidate variants in 6548 individuals that were independent and had no overlapping samples with the phase I dataset. Due to the longevity phase II dataset only containing longevity candidate variants, we took it as the replication exposure dataset to increase power when calculating the MR from longevity to microbiome direction.
Two-sample MR design
We applied a bidirectional two-sample MR strategy: (1) In the forward direction, the GWAS summary statistics of gut and oral microbiota obtained from 4D-SZ cohort (500 gut microbial features for 1539 individuals, 1665 tongue dorsum microbial features for 2017 individuals and 1770 salivary microbial features for 1915 individuals; all individuals with high-depth whole genome and metagenome sequencing) were used as sample exposure, and the GWAS summary statistics from longevity phase I dataset (N = 4477 for 5.6 M genome-wide imputed variants) were used as sample outcome, to identify the causal effects of the microbiota on longevity; (2) In the reverse direction, the GWAS summary statistics from longevity phase I project were used as discovery sample exposure, and significant candidate longevity genes from longevity phase II dataset (N = 6548 independent and no overlapping samples with phase I project for 287 K candidate variants) were used as replication sample exposure, and the gut and oral microbiota obtained from 4D-SZ cohort were used as sample outcome, to identify the causal effects of longevity on the microbiota.
Instrumental variable selection
For each genome-wide association result of microbial features and host longevity phenotype, we selected genetic variants that showed association at p < 1 × 10−5 and then performed the linkage disequilibrium (LD) estimation with a threshold of LD r2 < 0.1 for clumping analysis to get independent genetic variants as IVs. As previously described21, the p-value threshold of 1 × 10−5 was used for the selection of genetic IVs associated with microbial features by maximizing the strength of genetic instruments and the amount of the average genetic variance explained by the genetic predictors in an independent sample. Therefore, we used a more liberal threshold of p < 1 × 10–5 to select the instruments for microbial features, and the instrumental mean F statistics reached 51.4 for the gut microbiome, 22 for the tongue dorsum microbiome, and 18 for the salivary microbiome (all greater than 10) that indicates strong instruments, consistent with our previous study. We assessed the IV assumptions69 by directly testing the association of the selected IVs with a range of potential confounders. In this study, we selected potential confounders comprised of 220 phenotypes in BBJ70, including diseases such as type 2 diabetes, and colorectal cancer and metabolic traits such as body mass index (BMI), high-density lipoprotein (HDL), low-density lipoprotein (LDL), and triglycerides that were commonly reported linked to the microbiome. As shown in Supplementary Table 12, we identified an average of 45 IVs for the 500 gut microbial features, 63.6% (= 318/500) microbial features exhibited no IVs associated with any of the 220 potential confounders (p > 10−5) and 28.6% microbial features exhibited only one genetic variant that was associated with one of the 220 potential confounders (p < 10−5 according to the p-value cut-off definition of significant IVs). Altogether, 92.2% microbial features had no more than 1 potential confounder-associated IV given a mean of 45 IVs for each microbial feature. Similarly, for the 1770 salivary microbial features (an average of 39 IVs for each feature), 69.72% (= 1234/1770) microbial features exhibited no IVs associated with any of the 220 potential confounders and 94.29% microbial features had no more than 1 potential confounder-associated IV. These results indicated the MR analyses in this study basically met the instrumental variable assumptions.
For consistency, we used the same threshold and procedure for selecting genetic IVs of longevity traits in both the longevity discovery and the replication cohort. We retained a total of 45 IVs (F statistics = 59.2) from the longevity exposure cohort and 30 IVs (F statistics = 46.7) from the longevity replication cohort for the MR analysis. A list of genetic instruments could be checked in Supplementary Fig. 2.
Two-sample MR analysis
The bidirectional two-sample MR analysis was performed by applying the GCTA-GSMR (Generalised Summary-data-based Mendelian Randomization) method, which excludes SNPs that show evidence of pleiotropic effects by the heterogeneity in dependent instruments outlier analysis (HEIDI-outlier test, 0.01). We therefore estimated a causal association between the exposure and the outcomes using the GSMR method71 as our principal MR analytical approach.
We subsequently performed a sensitivity and validation analysis for each MR using five other methods (inverse-variance weighting (IVW)72,73, MR-Egger regression74, weighted median75, mode-based estimate (MBE)76 including Simple mode and Weighted mode) implemented in the “TwoSampleMR” R package. A consistent effect across the four methods is less likely to be a false positive. We used the MR–Egger intercept and MR-PRESSO global test to test for the presence of directional pleiotropy. MR tests were considered statistically significant when p < 0.05.
Two-sample MR analysis for European populations
For comparison, we also performed a two-sample MR analysis to investigate causal effects between microbial features and longevity in European populations. GWAS summary statistics of the gut microbiome were obtained from three published studies involving MiBioGen (mainly European populations)22, Dutch35, and Finnish36 cohorts. The MiBioGen performed a meta-analysis of the gut microbiota GWAS in 18,340 participants with 13,266 European ancestry22. The Dutch microbial GWAS used metagenomic sequencing data on 7738 individuals from the northern Netherlands35. The Finnish microbial GWAS was performed on 5959 genotyped individuals with matched gut microbial shotgun metagenomes in the Finnish cohort36. GWAS summary statistics of longevity was from the largest meta-analysis of human longevity GWAS of European ancestry37. We used the 99th percentile longevity data whose individuals lived beyond the 99th (N = 3484) percentile, which was comparable to the Chinese longevity study with centenarians (N = 2187) as cases. For consistency, genetic variants with p < 1 × 10−5 and LD r2 < 0.1 were selected as instrumental variables and the GCTA-GSMR approach was used for MR analysis. The 1kgenome Phase 3 dataset with only European population ancestry is used as the reference panel for LD clumping.
Two-sample MR analysis for diseases in Japan Biobank
To investigate the causal effect of microbial features on diseases, we performed a bidirectional two-sample MR analysis using summary statistics of oral microbiota from our cohort and disease information from Biobank Japan. We downloaded both summary statistics data for 42 diseases from Biobank Japan77 (http://jenger.riken.jp/en/result). The bidirectional two-sample MR analysis was performed by applying the GCTA-GSMR71 using the HEIDI-outlier analysis to remove horizontal pleiotropic SNPs. For consistency, genetic variants with p < 1 × 10−5 and LD r2 < 0.1 were selected as instrumental variables for oral microbiota in our cohort. For disease exposures, SNP instruments were selected at a genome-wide significant threshold (p < 5 × 10–8) in the Japan Biobank study.
Data availability
All statistical results were available in supplementary tables. The summary statistics including the associations between host genetics and the gut microbiomes, host genetics and metabolites are publicly available from https://ftp.cngb.org/pub/CNSA/data2/CNP0000794/. The summary statistics including the associations between host genetics and tongue dorsum microbiome, host genetics and saliva microbiome are publicly available from https://db.cngb.org/search/project/CNP0001664. The release of these summary statistics data was approved by the Ministry of Science and Technology of China (Project ID: 2020BAT1137; 2021BAT1539). The summary statistics of microbial features from the MiBioGen cohort are available at www.mibiogen.org. The summary statistics of microbial features from the Finnish cohort are available via the NHGRI-EBI GWAS catalog https://www.ebi.ac.uk/gwas/studies/GCST90032172. The summary statistics of microbial features from the Dutch cohort are available at https://molgenis26.gcc.rug.nl/downloads/dutchmicrobiomeproject/summarystats/taxa. The full meta-analyses summary statistics of longevity are available for download at www.longevitygenomics.org/downloads.
References
Hjelmborg, J. V. B. et al. Genetic influence on human lifespan and longevity. Hum. Genet. 119, 312–321. https://doi.org/10.1007/s00439-006-0144-y (2006).
van den Berg, N. et al. Longevity defined as top 10% survivors and beyond is transmitted as a quantitative genetic trait. Nat. Commun. 10, 35. https://doi.org/10.1038/s41467-018-07925-0 (2019).
Badal, V. D. et al. The gut microbiome, aging, and longevity: A systematic review. Nutrients 12, 123759. https://doi.org/10.3390/nu12123759 (2020).
Kong, F., Deng, F., Li, Y. & Zhao, J. Identification of gut microbiome signatures associated with longevity provides a promising modulation target for healthy aging. Gut Microbes 10, 210–215. https://doi.org/10.1080/19490976.2018.1494102 (2019).
Han, B. et al. Microbial genetic composition tunes host longevity. Cell 173, 1058. https://doi.org/10.1016/j.cell.2018.04.026 (2018).
Biagi, E. et al. Gut microbiota and extreme longevity. Curr. Biol. 26, 1480–1485. https://doi.org/10.1016/j.cub.2016.04.016 (2016).
Wang, N. et al. Enriched taxa were found among the gut microbiota of centenarians in East China. PLoS ONE 14, e0222763. https://doi.org/10.1371/journal.pone.0222763 (2019).
Ghosh, T. S., Shanahan, F. & O’Toole, P. W. The gut microbiome as a modulator of healthy ageing. Nat. Rev. Gastroenterol. Hepatol. 19, 565–584. https://doi.org/10.1038/s41575-022-00605-x (2022).
Wilmanski, T. et al. Gut microbiome pattern reflects healthy ageing and predicts survival in humans. Nat. Metab. 3, 274–286. https://doi.org/10.1038/s42255-021-00348-0 (2021).
Zhang, X. et al. The oral and gut microbiomes are perturbed in rheumatoid arthritis and partly normalized after treatment. Nat. Med. 21, 895–905. https://doi.org/10.1038/nm.3914 (2015).
Dzidic, M. et al. Oral microbiome development during childhood: An ecological succession influenced by postnatal factors and associated with tooth decay. ISME J. 12, 2292–2306. https://doi.org/10.1038/s41396-018-0204-z (2018).
Willis, J. R. & Gabaldón, T. The human oral microbiome in health and disease: From sequences to ecosystems. Microorganisms 8, 020308. https://doi.org/10.3390/microorganisms8020308 (2020).
Burgess, S., Timpson, N. J., Ebrahim, S. & Davey Smith, G. Mendelian randomization: Where are we now and where are we going? Int. J. Epidemiol. 44, 379–388. https://doi.org/10.1093/ije/dyv108 (2015).
Perrot, N. et al. A trans-omic Mendelian randomization study of parental lifespan uncovers novel aging biology and therapeutic candidates for chronic diseases. Aging Cell 20, e13497. https://doi.org/10.1111/acel.13497 (2021).
Timmers, P. R. H. J. et al. Mendelian randomization of genetically independent aging phenotypes identifies LPA and VCAM1 as biological targets for human aging. Nat. Aging 2, 19–30. https://doi.org/10.1038/s43587-021-00159-8 (2022).
Huang, S.-Y. et al. Investigating causal relationships between exposome and human longevity: A Mendelian randomization analysis. BMC Med. 19, 150. https://doi.org/10.1186/s12916-021-02030-4 (2021).
Liu, Z. et al. Associations of triglyceride levels with longevity and frailty: A Mendelian randomization analysis. Sci. Rep. 7, 41579. https://doi.org/10.1038/srep41579 (2017).
Yoon, S. Y. & Narayan, V. P. Genetically predicted glucosamine and longevity: A Mendelian randomization study. Clin. Nutr. ESPEN 49, 556–559. https://doi.org/10.1016/j.clnesp.2022.03.029 (2022).
Goodrich, J. K. et al. Genetic determinants of the gut microbiome in UK twins. Cell Host Microbe 19, 731–743. https://doi.org/10.1016/j.chom.2016.04.017 (2016).
Xie, H. et al. Shotgun metagenomics of 250 adult twins reveals genetic and environmental impacts on the gut microbiome. Cell Syst. 3, 572–584. https://doi.org/10.1016/j.cels.2016.10.004 (2016).
Liu, X. et al. Mendelian randomization analyses support causal relationships between blood metabolites and the gut microbiome. Nat. Genet. 54, 52–61. https://doi.org/10.1038/s41588-021-00968-y (2022).
Kurilshikov, A. et al. Large-scale association analyses identify host factors influencing human gut microbiome composition. Nat. Genet. 53, 156–165. https://doi.org/10.1038/s41588-020-00763-1 (2021).
Qin, Y. et al. Combined effects of host genetics and diet on human gut microbiota and incident disease in a single population cohort. Nat. Genet. 54, 134–142. https://doi.org/10.1038/s41588-021-00991-z (2022).
Lopera-Maya, E. A. et al. Effect of host genetics on the gut microbiome in 7,738 participants of the Dutch Microbiome Project. Nat. Genet. 54, 143–151. https://doi.org/10.1038/s41588-021-00992-y (2022).
Liu, X. et al. A genome-wide association study for gut metagenome in Chinese adults illuminates complex diseases. Cell Discov. 7, 9. https://doi.org/10.1038/s41421-020-00239-w (2021).
Liu, X. et al. Metagenome-genome-wide association studies reveal human genetic impact on the oral microbiome. Cell Discov. 7, 117. https://doi.org/10.1038/s41421-021-00356-0 (2021).
Zeng, Y. et al. Novel loci and pathways significantly associated with longevity. Sci. Rep. 6, 21243. https://doi.org/10.1038/srep21243 (2016).
Omar, J. M., Chan, Y.-M., Jones, M. L., Prakash, S. & Jones, P. J. H. Lactobacillus fermentum and Lactobacillus amylovorus as probiotics alter body adiposity and gut microflora in healthy persons. J. Funct. Foods 5, 116–123. https://doi.org/10.1016/j.jff.2012.09.001 (2013).
Nomura, R. et al. Contribution of severe dental caries induced by Streptococcus mutans to the pathogenicity of infective endocarditis. Infect. Immunity 88, e00897. https://doi.org/10.1128/IAI.00897-19 (2020).
Kostic, A. D. et al. Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe 14, 207–215. https://doi.org/10.1016/j.chom.2013.07.007 (2013).
Zhang, X. et al. Sex- and age-related trajectories of the adult human gut microbiota shared across populations of different ethnicities. Nat. Aging 1, 87–100. https://doi.org/10.1038/s43587-020-00014-2 (2021).
Jie, Z. et al. A transomic cohort as a reference point for promoting a healthy human gut microbiome. Med. Microecol. 8, 100039. https://doi.org/10.1016/j.medmic.2021.100039 (2021).
Liu, X. et al. Integrated genetic analyses revealed novel human longevity loci and reduced risks of multiple diseases in a cohort study of 15,651 Chinese individuals. Aging Cell 20, e13323. https://doi.org/10.1111/acel.13323 (2021).
Kim, B. S. et al. Comparison of the gut microbiota of centenarians in longevity villages of South Korea with those of other age groups. J. Microbiol. Biotechnol. 29, 429–440. https://doi.org/10.4014/jmb.1811.11023 (2019).
Lopera-Maya, E. A. et al. Effect of host genetics on the gut microbiome in 7,738 participants of the Dutch Microbiome Project. BioRxiv. https://doi.org/10.1101/2020.12.09.417642 (2021).
Qin, Y. et al. Genome-wide association and Mendelian randomization analysis prioritizes bioactive metabolites with putative causal effects on common diseases. Nat. Genet. https://doi.org/10.1038/s41588-021-00991-z (2020).
Deelen, J. et al. A meta-analysis of genome-wide association studies identifies multiple longevity genes. Nat. Commun. 10, 3669. https://doi.org/10.1038/s41467-019-11558-2 (2019).
Tong, Z. et al. An in vitro investigation of Lactococcus lactis antagonizing cariogenic bacterium Streptococcus mutans. Arch. Oral Biol. 57, 376–382. https://doi.org/10.1016/j.archoralbio.2011.10.003 (2012).
Valenzuela, J. A., Flórez, A. B., Vázquez, L., Vasek, O. M. & Mayo, B. Production of γ-aminobutyric acid (GABA) by lactic acid bacteria strains isolated from traditional, starter-free dairy products made of raw milk. Benef. Microbes 10, 579–587. https://doi.org/10.3920/bm2018.0176 (2019).
Ma, S. et al. Metagenomic analysis reveals oropharyngeal microbiota alterations in patients with COVID-19. Signal Transduct. Target. Ther. 6, 191. https://doi.org/10.1038/s41392-021-00614-3 (2021).
Adamovich, Y. et al. Oxygen and carbon dioxide rhythms are circadian clock controlled and differentially directed by behavioral signals. Cell Metab. 29, 1092–1103. https://doi.org/10.1016/j.cmet.2019.01.007 (2019).
van der Goot, A. T. et al. Delaying aging and the aging-associated decline in protein homeostasis by inhibition of tryptophan degradation. Proc. Natl. Acad. Sci. 109, 14912–14917. https://doi.org/10.1073/pnas.1203083109 (2012).
Yu, L. et al. A systematic review of microbial markers for risk prediction of colorectal neoplasia. Br. J. Cancer 126, 1318–1328. https://doi.org/10.1038/s41416-022-01740-7 (2022).
Lopes, M. P. et al. Prevotella intermedia and periodontitis are associated with severe asthma. J. Periodontol. 91, 46–54. https://doi.org/10.1002/jper.19-0065 (2020).
Sato, Y. et al. Novel bile acid biosynthetic pathways are enriched in the microbiome of centenarians. Nature. https://doi.org/10.1038/s41586-021-03832-5 (2021).
Luedde, M. et al. Heart failure is associated with depletion of core intestinal microbiota. ESC Heart Fail. 4, 282–290. https://doi.org/10.1002/ehf2.12155 (2017).
Lim, M. Y. et al. The effect of heritability and host genetics on the gut microbiota and metabolic syndrome. Gut 66, 1031–1038. https://doi.org/10.1136/gutjnl-2015-311326 (2017).
Abratt, V. R. & Reid, S. J. Oxalate-degrading bacteria of the human gut as probiotics in the management of kidney stone disease. Adv. Appl. Microbiol. 72, 63–87. https://doi.org/10.1016/s0065-2164(10)72003-7 (2010).
Ticinesi, A. et al. Understanding the gut-kidney axis in nephrolithiasis: An analysis of the gut microbiota composition and functionality of stone formers. Gut 67, 2097–2106. https://doi.org/10.1136/gutjnl-2017-315734 (2018).
Caro, J., Migliaccio-Walle, K., Ishak, K. J. & Proskorovsky, I. The morbidity and mortality following a diagnosis of peripheral arterial disease: Long-term follow-up of a large database. BMC Cardiovasc. Disord. 5, 14. https://doi.org/10.1186/1471-2261-5-14 (2005).
Wu, G. D. et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 334, 105–108. https://doi.org/10.1126/science.1208344 (2011).
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563. https://doi.org/10.1038/nature12820 (2014).
Levy, R. et al. Longitudinal analysis reveals transition barriers between dominant ecological states in the gut microbiome. Proc. Natl. Acad. Sci. U.S.A. 117, 13839–13845. https://doi.org/10.1073/pnas.1922498117 (2020).
Singh, R. K. et al. Influence of diet on the gut microbiome and implications for human health. J. Transl. Med. 15, 73. https://doi.org/10.1186/s12967-017-1175-y (2017).
Wu, L. et al. A cross-sectional study of compositional and functional profiles of gut microbiota in Sardinian centenarians. Msystems 4, e00325 (2019).
Chen, J. et al. The impact of glutamine supplementation on the symptoms of ataxia-telangiectasia: A preclinical assessment. Mol. Neurodegener. 11, 60. https://doi.org/10.1186/s13024-016-0127-y (2016).
Zhang, M. et al. Hydrogen extends Caenorhabditis elegans longevity by reducing reactive oxygen species. PLoS ONE 15, e0231972. https://doi.org/10.1371/journal.pone.0231972 (2020).
Gupta, V. K., Paul, S. & Dutta, C. Geography, ethnicity or subsistence-specific variations in human microbiome composition and diversity. Front. Microbiol. 8, 1162–1162. https://doi.org/10.3389/fmicb.2017.01162 (2017).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227. https://doi.org/10.1038/nature11053 (2012).
Bárcena, C. et al. Healthspan and lifespan extension by fecal microbiota transplantation into progeroid mice. Nat. Med. 25, 1234–1242. https://doi.org/10.1038/s41591-019-0504-5 (2019).
Cerro, E.D.-D. et al. Daily ingestion of Akkermansia mucciniphila for one month promotes healthy aging and increases lifespan in old female mice. Biogerontology. https://doi.org/10.1007/s10522-021-09943-w (2021).
Maki, K. A., Kazmi, N., Barb, J. J. & Ames, N. The oral and gut bacterial microbiomes: Similarities, differences, and connections. Biol. Res. Nurs. 23, 7–20. https://doi.org/10.1177/1099800420941606 (2021).
Zhu, J. et al. Over 50,000 metagenomically assembled draft genomes for the human oral microbiome reveal new taxa. Genom. Proteom. Bioinform. https://doi.org/10.1016/j.gpb.2021.05.001 (2021).
Li, J. et al. An integrated catalog of reference genes in the human gut microbiome. Nat. Biotechnol. 32, 834–841. https://doi.org/10.1038/nbt.2942 (2014).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 44, D471–D480. https://doi.org/10.1093/nar/gkv1164 (2016).
Kanehisa, M. et al. Data, information, knowledge and principle: Back to metabolism in KEGG. Nucleic Acids Res. 42, D199–D205. https://doi.org/10.1093/nar/gkt1076 (2014).
Zeng, Y. et al. Sex differences in genetic associations with longevity. JAMA Netw. Open 1, e181670. https://doi.org/10.1001/jamanetworkopen.2018.1670 (2018).
Zhao, X. et al. The correlation of copy number variations with longevity in a genome-wide association study of Han Chinese. Aging (Albany NY) 10, 1206–1222. https://doi.org/10.18632/aging.101461 (2018).
Burgess, S., Foley, C. N. & Zuber, V. Inferring causal relationships between risk factors and outcomes from genome-wide association study data. Annu. Rev. Genom. Hum. Genet. 19, 303–327. https://doi.org/10.1146/annurev-genom-083117-021731 (2018).
Sakaue, S. et al. A cross-population atlas of genetic associations for 220 human phenotypes. Nat. Genet. 53, 1415–1424. https://doi.org/10.1038/s41588-021-00931-x (2021).
Zhu, Z. et al. Causal associations between risk factors and common diseases inferred from GWAS summary data. Nat. Commun. 9, 224. https://doi.org/10.1038/s41467-017-02317-2 (2018).
Burgess, S., Butterworth, A. & Thompson, S. G. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet. Epidemiol. 37, 658–665. https://doi.org/10.1002/gepi.21758 (2013).
Bowden, J. et al. A framework for the investigation of pleiotropy in two-sample summary data Mendelian randomization. Stat. Med. 36, 1783–1802. https://doi.org/10.1002/sim.7221 (2017).
Bowden, J. et al. Assessing the suitability of summary data for two-sample Mendelian randomization analyses using MR-Egger regression: The role of the I2 statistic. Int. J. Epidemiol. 45, 1961–1974. https://doi.org/10.1093/ije/dyw220 (2016).
Bowden, J., Davey Smith, G., Haycock, P. C. & Burgess, S. Consistent estimation in mendelian randomization with some invalid instruments using a weighted median estimator. Genet. Epidemiol. 40, 304–314. https://doi.org/10.1002/gepi.21965 (2016).
Hartwig, F. P., Davey Smith, G. & Bowden, J. Robust inference in summary data Mendelian randomization via the zero modal pleiotropy assumption. Int. J. Epidemiol. 46, 1985–1998. https://doi.org/10.1093/ije/dyx102 (2017).
Ishigaki, K. et al. Large-scale genome-wide association study in a Japanese population identifies novel susceptibility loci across different diseases. Nat. Genet. 52, 669–679. https://doi.org/10.1038/s41588-020-0640-3 (2020).
Acknowledgements
The authors sincerely thank the support provided by colleagues of BGI-Shenzhen, supervisors and families. Collections of the Chinese Longitudinal Healthy Longevity Surveys (CLHLS) datasets and DNA samples analyzed in this paper were jointly supported by the National Key R&D Program of China (2018YFC2000400), National Natural Sciences Foundation of China (72061137004), the U.S. National Institute of Aging/ National Institute of Health (P01AG031719) and Duke/Duke-NUS/RECA(Pilot)/2019/0051. They thank for the support from the National Natural Science Foundation of China (No. 32200548).
Author information
Authors and Affiliations
Contributions
H.J. and Y.H. conceived and organized this study. X.X., H.Y., J.W., X.J., L.X., and Y.H. contributed to the organization of the microbiome cohort. Y.Z., J.M., C.N. contributed to the organization of the longevity cohort. X.L. and L.Z. performed Mendelian randomization analyses. H.J., T.Z., and Y.Q. participated in the discussion. X.L. wrote the raw manuscript. All authors contributed to the data and texts in this manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, X., Zou, L., Nie, C. et al. Mendelian randomization analyses reveal causal relationships between the human microbiome and longevity. Sci Rep 13, 5127 (2023). https://doi.org/10.1038/s41598-023-31115-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-31115-8
This article is cited by
-
Causal effects of denture wearing on epigenetic age acceleration and the mediating pathways: a mendelian randomization study
BMC Oral Health (2024)
-
Investigating the causal relationship between thyroid dysfunction diseases and osteoporosis: a two-sample Mendelian randomization analysis
Scientific Reports (2024)
-
Genetic insights into the association of statin and newer nonstatin drug target genes with human longevity: a Mendelian randomization analysis
Lipids in Health and Disease (2023)
-
Causal effect between gut microbiota and pancreatic cancer: a two-sample Mendelian randomization study
BMC Cancer (2023)
-
Causal effect of gut microbiota on DNA methylation phenotypic age acceleration: a two-sample Mendelian randomization study
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.