Abstract
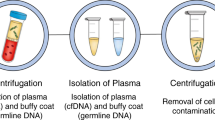
Liquid biopsy (LB) provides a unique minimally invasive tool to follow-up cancer patients over time, to detect minimal residual disease (MRD), to study metastasis-biology and mechanisms of therapy-resistance. Molecular characterization of CTCs offers additionally the potential to understand resistance to therapy and implement individualized targeted treatments which can be modified during the disease evolution and follow-up period of a patient. In this study, we present a long-term follow-up of operable breast cancer patients based on a comprehensive liquid biopsy analysis. We performed a comprehensive liquid biopsy analysis in peripheral blood of 13 patients with early-stage operable breast cancer at several time points for a period of ten years, consisting of: (a) CTC enumeration using the CellSearch system, (b) phenotypic analysis of CTCs using Immunofluorescence, (c) gene expression analysis, in EpCAM(+) CTCs for CK-19, CD24,CD44, ALDH1, and TWIST1, (d) analysis of PIK3CA and ESR1 mutations in EpCAM(+) CTCs and corresponding plasma ctDNA and (e) DNA methylation of ESR1 in CTCs. 10/13 (77%) patients were found negative for LB markers in PB during the whole follow-up period, and these patients did not relapse during the follow-up. However, 3/13(18%) patients that were positive for at least one LB marker relapsed within the follow-up period. The molecular characteristics of CTCs were highly different even for the same patient at different time points, and always increased before the clinical relapse. Our results indicate that liquid biopsy can reveal the presence of MRD at least 4 years before the appearance of clinically detectable metastatic disease demonstrating that a comprehensive liquid biopsy analysis provides highly important information for the therapeutic management of breast cancer patients.
Similar content being viewed by others
Introduction
Minimal residual disease (MRD) detection and monitoring remains a high challenge for the management of patients with solid tumours1,2. A considerable number of patients with breast cancer will develop metastasis within five years of primary tumor resection despite initially being free of detectable metastases depending on the tumor type and stage. In ER(+) breast cancer, after 5 years of adjuvant endocrine therapy, breast cancer recurrences continue to occur steadily throughout the study period from 5 to 20 years1. A significant proportion of these early-stage breast cancer patients that seemingly response to treatment have occult micrometastases or MRD that perseveres after initial therapy as a potential source of subsequent metastatic relapse at distant sites. The early identification of MRD in individual patients is highly challenging; towards this goal, real-time high-sensitivity liquid biopsy (LB) assays are highly promising and offer a great potential to address this2.
LB is a minimally invasive approach that is mainly based on the analysis of circulating tumor cells (CTCs) and circulating tumor DNA (ctDNA)3,4,5. LB is an important tool in the fight against cancer, since it provides the ability to monitor in serial samples a molecular portrait of the tumor in real time3,6,7. CTC analysis currently presents a powerful tool for the management of advanced- and early-stage cancer patients; especially CTC molecular characterization offers the unique potential to better understand the biology of metastasis and resistance mechanisms to specific treatments3,6. In the early stages of cancer, CTCs are usually detected at very low numbers and are characterized by a high heterogeneity7. It is more than ten years that CTCs enumeration performed in the CellSearch system has been FDA-cleared as a prognostic marker and has been associated with progression-free survival (PFS) and overall survival (OS) in early and metastatic breast, metastatic prostate and colorectal cancer3,8,9. The STIC CTC trial has shown that CTC count can be a reliable biomarker for choosing between chemotherapy and single-agent endocrine therapy as the first-line treatment in hormone receptor–positive HER2-negative metastatic breast cancer10. Müller et al.11 demonstrated that the presence of even one CTC with strong HER2 staining was associated with shorter OS, supporting a biological role of HER2 expression on CTCs.
During the last years CTC biology is under intense study12. Beyond CTC enumeration, molecular characterization of CTCs at the gene expression level has the potential to clarify the critical signalling pathways involved in metastasis biology and even improve patient management3,6. We have previously shown that the detection of cytokeratin-19 (CK-19) transcripts in CTCs by RT-qPCR in breast cancer patients is of prognostic significance before, during and after adjuvant chemotherapy13,14,15,16,17,18,19,20. It is also known that the expression of epithelial-mesenchymal transition (EMT) and stem cell markers like vimentin, N-cadherin, SNAI1 (also known as Snail), SNAI2 (also known as Slug), TWIST, zinc finger E-box binding homeobox 1 (ZEB1), and ZEB2 in CTCs is associated with an invasive phenotype21. Moreover, a subset of tumor cells with stem cell properties can contribute to tumor initiation and growth but can also play an important role to the development of metastasis and drug resistance3. Cancer stem cell (CSC) markers like ALDH1, CD24, CD44, and CD133 have been detected in CTCs22,23,24,25,26,27, and their clinical relevance has already been reported15,28.
In breast cancer, detection of PIK3CA mutations is highly important, since recently there are specific targeted therapies developed29. Detection of PIK3CA in CTCs could have important clinical applications for the follow-up of these patients. However, this is very challenging, since CTCs are heterogeneous and cells carrying these mutations consist often a minority in the CTC population. Our group has developed highly sensitive and specific assays for the detection of PIK3CA hotspot mutations (E545K, H1047R)30,31,32 and ESR1 mutations33 in EpCAM(+) CTC of early and metastatic breast cancer patients. There are also several studies in which the PIK3CA mutational status of CTCs has been investigated at the single cell level revealing the heterogeneity of CTC in the same patient34,35,36. ESR1 gene mutations consist one of the resistant mechanisms to endocrine therapies for ER-positive metastatic breast cancer patients37 and have been also detected in single CTCs exposing important information about their mutational heterogeneity and subsequently for the resistance to therapies38,39. Beyond mutations, epigenetic modifications, especially DNA methylation of tumor and suppressor genes’ promoters, play an important role in cancer development40. We have recently reported that methylation of the ESR1 promoter is an alternative and additional mechanism of ESR1 inactivation leading to resistance to hormone treatment in breast cancer41.
In this study, we present for the first time a comprehensive liquid biopsy analysis for 13 patients all diagnosed with early breast cancer, based on a long-term follow-up. Our analysis was based on CTC enumeration, CTC phenotypic characterization and molecular monitoring of CTCs at the gene expression, DNA mutation and DNA methylation level, and on corresponding plasma cell free DNA for DNA mutation and DNA methylation level. Our results indicate that a comprehensive liquid biopsy analysis provides valuable and important information for the therapeutic management of breast cancer patients since it is the only way to shed light into the black box that was sealed till now between the information taken from the primary tumor and from a distant metastatic site. Our results demonstrate that liquid biopsy analysis could reveal the presence of MRD years before the appearance of clinically detectable metastatic disease and progression of disease.
Results
CK-19 mRNA expression
The presence of CK-19 transcripts was assessed in all patient samples at different time points during the follow-up period (Fig. 1). Among the ten patients remained free from disease progression over the ten years of follow-up, CK-19 transcripts were not detected in eight (8/10, 80%) of them (Pt#1, Pt#2, Pt#4, Pt#5, Pt#7, Pt#8, Pt#9, Pt#10) at all time points. The other two patients (Pt#3, Pt#6) were found positive for CK-19 transcripts; more specifically, Pt#3 was found positive for CK-19 transcripts on month 53 after diagnosis and Pt#6 on months 28 and 39, while CK-19 transcripts were not detected at all previous and ensuing time points.
All breast cancer patients (Pt#11, Pt#12, Pt#13) that were positive for CK-19 transcripts early on, and at various time points later developed metastasis (Fig. 1). Pt#11 at the initial time point was found negative for CK-19 transcripts. However, after 78 and 81 months after initial diagnosis, high levels of CK-19 transcripts were detected in CTCs and two months later progression of disease (PD) was confirmed by detecting metastatic lesions on adrenal gland and liver. During the metastatic phase of the disease, all samples analyzed for CK-19 transcripts were found positive and PD was confirmed a few months later at each time point. This patient died three years after metastasis was documented clinically and by imaging studies and one month after a new progression. Pt#12 was found positive for CK-19 transcripts 61 months since diagnosis and one month later after confirmation of PD (Fig. 1). In Pt#13, no CK-19 transcripts were detected before the initiation of adjuvant chemotherapy. On September 2014, a significant increase in the number of CK-19 transcripts was observed. Eight months later a liver metastasis was confirmed by a liver biopsy. During the metastatic phase of the disease (Fig. 1) all samples analyzed for CK-19 transcripts were found positive.
Detection of ESR1 & PIK3CA mutations
We further analyzed all available plasma-cfDNA and CTC-derived gDNA samples from the three breast cancer patients (Pt#11, Pt#12, Pt#13) that developed metastasis at different time points for ESR1 and PIK3CA mutations (Fig. 2). We first used a drop-off ddPCR to detect ESR1 mutations in plasma-ctDNA samples at different time points41. Pt#12 was found negative for ESR1 mutations in plasma-ctDNA at all the time points tested; this patient experienced a clinical objective response to endocrine treatment and remained free from disease progression at the time of analysis for 61 months since diagnosis. Pt#13 was found positive for ESR1 mutations in plasma-ctDNA consistently at five different time points during the follow-up period (Fig. 2). This patient later developed resistance to endocrine therapy. In parallel, all available plasma-cfDNA and CTC-derived gDNA samples were analyzed for PIK3CA mutations. PIK3CA E545K mutation was detected in 2/10 gDNA samples from Pt#11 extracted from EpCAM(+) fractions (Fig. 2). Pt#12 was found positive for PIK3CA E545K mutation in 8/14 gDNA samples extracted from EpCAM(+) fractions and in 1/14 paired plasma-ctDNA samples (Fig. 2). Pt#13 was found positive for PIK3CA H1047R mutation in 3/10 gDNA samples extracted from EpCAM(+) cell fractions and in 2/10 paired plasma-ctDNA samples. PIK3CA E545K mutation was detected in 2/10 paired plasma-ctDNA samples.
Comprehensive liquid biopsy analysis for Pt#13
An overview presenting timeline of the disease course from diagnosis onward, therapeutic interventions and the results for CTC enumeration, and the phenotypic and molecular characterization of CTCs at different time points for Pt#13 is shown in Fig. 3A; therapeutic interventions for this patient at different time points are shown in Fig. 3B.
-
CTC enumeration and therapeutic interventions
CTC enumeration using the CellSearch system was performed at regular time points during the follow-up period (Fig. 4A). Before the initiation of adjuvant systemic chemotherapy, CTC enumeration revealed the presence of 10CTCs/23 mL PB (time point: 0 months, Fig. 4A). The patient received a dose-dense chemotherapy (AC every 2 weeks followed by 4 cycles docetaxel every 2 weeks followed by adjuvant radiotherapy42,43. At the time of radiotherapy completion (Fig. 3B), a new CTC enumeration test revealed the presence of 7CTCs/23 mL PB. During adjuvant hormone treatment (LH/LH analogs and tamoxifen) (time point: 15 months, Fig. 4A) a slight increase of the CTC number (10CTCs/23 mL PB) was observed which was further confirmed 3 months later. On January 2011, letrozol was added instead of tamoxifen (Fig. 3B) and a more detailed characterization of CTCs by double immunofluorescence demonstrated that practically all the detected cells were CK(+)/HER2(+) and CK(+)/Vimentin(+) (Fig. 3A). Based on these results the patient was enrolled in a phase II randomized clinical trial of secondary adjuvant Trastuzumab versus standard hormone treatment42 and was allowed to receive trastuzumab along with hormone treatment; treatment (April 2011–December 2011) resulted in a clear decrease of the absolute number of CTCs (time point: 15 months, Fig. 4A). On February 2012 a significant increase of CTCs was noted, (time point: 26 months, Fig. 4A) bearing the HER2(−), EGFR(+), Ki67(−), M30(−) phenotype. Trastuzumab was discontinued and a 3-month treatment with lapatinib was administered based on the EGFR + phenotype of the cells without success (time point: 26 months, Fig. 4A). On July 2012, the patient received afinitor/aromasin with a clear decrease of the CTCs’ number from 25/23 mL PB (time points: 26 months and 37 months, Fig. 4A) until September 2014 when 275 CTCs/23 mL PB reappeared in the blood. A parallel analysis based on IF in the same PB samples (Fig. 3A) has shown that these cells were CK(+)/HER2(−) (time point: 57 months, Fig. 4A). At this period tumor markers were moderately increased for the first time (CEA: 6 mg/mL and CA19.9: 51 IU/mL). On December 2014, whole bone scintigraphy demonstrated lesions suspicious for bone metastases which, however, could not be confirmed by MRIs. Until September 2015 the patient was without medical complains and the imaging studies with CT scans every 4 months were practically negative; the patient continued to receive afinitor/aromasin despite the clinical suspicion of disease progression. At this period the MRI of the abdomen as well as a PET scan demonstrated the presence of multiple hypodense lesions of the liver and a liver biopsy has confirmed the metastatic nature of the lesions (ER+, PR+, HER2-neg, Ki67 = 50%), compatible with the primary tumor. The patient received front line chemotherapy with weekly paclitaxel + bevacizumab which resulted in a significant decrease of the CTC number despite the stable disease which was documented as best clinical response 1254 CTCs/7.5 mL PB (time point: 74 months, Fig. 4A) and 108/7.5 mL PB, (time point: 82 months, Fig. 4A), respectively. Subsequently, because of a new increase of CTC count [CTCs: 411/7.5 mL PB (time point: 76 months, Fig. 4A)] a combination of capecitabine/bevacizumab was initiated with major objective response after 3 cycles, resulting to a long lasting clinical partial response with a decreased CTC count [CTCs:411/7.5 mL PB (time point: 76 months, Fig. 4A), to 108 CTCs/7,5 mL PB (time point: 82 months, Fig. 4A), to CTCs: 82 CTCs/7.5 mL PB (time point: 103 months, Fig. 4A)]; however, the CTC counts were increased again (478 CTCs/7.5 mL PB time point: 116 months, Fig. 4A) and were further increased despite under a combination of a CDK4/6 inhibitor/Faslodex [1423CTCs/7.5 mL PB (time point: 122 months, Fig. 4A)]. On March 2020, the number of CTCs was decreased (703 CTCs/7.5 mL PB, time point: 124 months, Fig. 4A) while on May 2020, at the time of carcinomatous meningitis diagnosis, an increase of CTC values was observed.
-
Molecular analysis of CTCs and corresponding ctDNA
P#13: (A) CTC enumeration values (CTCs/7.5 mL PB) based on CellSearch, during the timeline of the disease (red stars represent the time of progression disease). (B) Molecular analysis of EpCAM(+) CTC fractions and CTCs from CellSearch cartridges for gene expression, PIK3CA and ESR1 mutations and ESR1 methylation, and corresponding plasma ctDNA (purple: positive, green: negative, crossed circle: missing value). (C) Representative images of CTCs by immunofluorescence experiments and confocal laser scanning microscopy analysis.
Gene expression
CK-19 mRNA
Before the initiation of adjuvant chemotherapy, CK-19 transcripts were not detected in PB of Pt#13. On September 2014, CK-19 transcripts were observed: this has been confirmed by a parallel analysis in the CellSearch, using the same blood draw (275 CTCs/7.5 mL of blood; time point: 57 months, Fig. 4A). Eight months later (time point: 66 months, Fig. 4A) a liver metastasis, confirmed by a new biopsy, was detected. During the metastatic phase of the disease (February 2016 until May 2020; time points: 74–103 months, 116–126 months, Fig. 4B) all samples analyzed for CK-19 transcripts were found positive.
CD24, CD44, ALDH1, TWIST1 mRNA
The CD24−/low/CD44high profile was detected in all samples (time points: 74–103 months, 116–126 months, Fig. 4B) and the CD24−/low/ALDH1high in 5/10 (50%) samples (time points: 74, 82, 100, 122, 124 months, Fig. 4B). TWIST1 overexpression was detected in 3/10 (30%) samples (time points: 74, 100, 118 months, Fig. 4B). It should be mentioned that TWIST1 overexpression was detected in samples with CD24−/low/CD44very high profile.
DNA mutations
PIK3CA mutations
PIK3CA hotspot mutations were detected in 25 gDNA samples extracted from CellSearch cartridges (n = 15) and EpCAM(+) CTC fractions (n = 10) (Fig. 4B). In particular, PIK3CA hotspot mutations were detected in 5/15(36%) gDNA samples extracted from CTCs isolated from CellSearch cartridges. E545K mutation was detected in 4/15 (27%) samples (time points: 59, 63, 69, 74 months, Fig. 4B) and H1047R mutation in another 1/15 (7%) sample (time point: 61 months, Fig. 4B). All samples found positive for PIK3CA mutations were collected one year after the patient started to receive Afinitor/Aromasin therapy. In addition, PIK3CA H1047R hotspot mutation was detected in 3/10 (30%) gDNA samples extracted from EpCAM(+) CTC fractions (time points: 76, 100, 103 months, Fig. 4B). It is important to mention that PIK3CA mutations were not detected in gDNA from the primary tumor (FFPEs) when the same highly sensitive methodology was used. There was no concordance in the findings for the detection of PIK3CA mutations between EpCAM(+)fractions and CellSearch cartridges. On March 2020 the patient received Alpelisib based on the detection of PIK3CA mutations in ctDNA and CTCs for a period of 2 months. But the drug was discontinued after two months due to serious adverse events. In paired plasma-ctDNA samples, PIK3CA H1047R hotspot mutation was detected in 2/10 (20%) (time points: 76, 118 months, Fig. 4B) and PIK3CA E545K in 2/10 (20%) (time points: 124, 126 months, Fig. 4B). It is noteworthy that the same mutations were also detected in CTCs obtained from both CellSearch cartridges and EpCAM(+) fractions.
ESR1 mutations
ESR1 hotspot mutations, were detected in 20 gDNA samples extracted from CellSearch cartridges (n = 12) and EpCAM(+) CTC fractions (n = 8) (Fig. 4B). In particular, ESR1 hotspot mutations were detected in 6/12(50%) gDNA samples extracted from CTCs isolated from CellSearch cartridges. More specifically, ESR1 D538G hotspot mutation was detected in 6/12(50%) DNA samples (time points: 59, 61, 63, 69, 70, 74 months, Fig. 4B), the Y537S in 1/12(8%) samples (time point: 70 months, Fig. 4B), Y537C in 5/12 (42%) samples (time points: 59, 61, 63, 70, 74 months, Fig. 4B) and the Y537N in 1/12(8%) samples (time point: 61 months, Fig. 4B). All samples found positive for ESR1 mutations were collected one year after the patient started Afinitor/Aromasin therapy. At time point #28, 11 months after the first detection of ESR1 mutations in CellSearch cartridges the patient relapsed while she was under Afinitor/Aromasin therapy. ESR1 Y537N hotspot mutation was detected in 1/8(13%) gDNA samples extracted from EpCAM(+) CTC fractions (time point: 116 months, Fig. 4B). In paired plasma-ctDNA samples, ESR1 D538G hotspot mutation was detected in 7/9(78%) of tested cases (time points: 74, 82, 116, 118, 122, 124, 126 months, Fig. 4B).
ESR1 methylation
The presence of ESR1 methylation was investigated in 22 gDNA samples extracted from CellSearch cartridges (n = 12) and EpCAM(+) cell fractions (n = 10) during the follow-up period (Fig. 4B). ESR1 methylation was detected in 8/10 (80%) of the SB-treated gDNA samples extracted from EpCAM(+) cell fractions (time points: 74, 76, 100, 103, 116, 118, 122, 126 months, Fig. 4Β). In all paired plasma-ctDNA samples (time points: 74, 76, 82, 100, 103, 116, 118, 122, 124, 126 months, Fig. 4B) ESR1 methylation was detected with a significant agreement with the corresponding EpCAM(+) cell fraction in 8/10 (80%) cases.
-
Phenotypic characterization of CTCs by IF
The phenotypic characterization of CTCs using double IF staining revealed that a substantial number of CTCs were HER2+ even if the primary tumor was negative for HER2 amplification. In particular, we have noticed changes of HER2+ expression in CTCs during the follow-up period. In 2012, the patient was enrolled in a prospective randomized trial of secondary adjuvant Herceptin trastuzumab versus standard treatment in patients with HER2(+) CTCs after completion of adjuvant chemotherapy43. At the time point of 26 months follow-up, 25CTCs/23 mL of blood were enumerated using the CellSearch (Fig. 4Α). In a paired PB sample, using the same blood draw, the phenotypic characterization of CTCs has revealed that: 1 CTC was CK(+)/HER2(−), 13 CTCs were CK(+)/EGFR(+), 14 CTCs were CK(+)/EGFR(−) and 112 CTCs were CK(+)/M30(−). No CTCs were detected for the following phenotypes: CK(+)/HER2(+), CK(+)/M30(+), CK(+)/ki67(+), CK(+)/ki67(−), CK(+)/VEGF(+), CK(+)/VEGFR2(+), CK(+)/VEGFR2(−) (Fig. 3A). At the time point of 29 months (Fig. 4A), when 17CTCs/23 mL of blood were enumerated using the CellSearch, the phenotypic characterization of CTCs demonstrated that the cells were HER2(−) (HER2(−), EGFR(+), Ki67(−), M30(−) and VEGFR2(−)) (Fig. 3A). In September 2014, after 57 months of follow-up (Fig. 4A), when 275 CTCs/7.5 mL were enumerated in peripheral blood our IF analysis has confirmed that these cells were CK(+)/HER2(−) (Fig. 3A). Representative images of CTCs by immunofluorescence experiments and confocal laser scanning microscopy analysis are shown in Fig. 4C.
Discussion
We present a comprehensive liquid biopsy analysis of 13 ER(+) breast cancer patients initially diagnosed with operable disease, based on the analysis of serial peripheral blood samples at different time points in a period of ten years, and correlate our findings with the clinical outcome.
In all ten patients that remained free from disease progression during the follow-up period, CK-19 transcripts were detected only at 3/135 (2.2%) time points tested. Until now, all these patients are alive and remained free from progression disease, except Pt#8 that was lost to follow-up. On the contrary, an important increase in the copy number of CK-19 transcripts was observed for two patients (Pt#11, Pt#13) a few years after initial diagnosis and before the confirmation of PD and diagnosis of metastasis. Our results are in agreement with previous clinical studies who have shown that the detection of CK-19(+) mRNA cells in the peripheral blood of patients with operable breast cancer before, during and after adjuvant treatment is an independent prognostic factor associated with an increased risk of disease relapse and shorter survival16,17,18,44,45. It has also been shown that the presence of CK-19(+) CTCs after the completion of chemotherapy is associated with increased risk of late relapse and poor survival in metastatic breast cancer46,47,48. Matikas et al. have recently shown that breast cancer patients with CK-19(+) CTCs at baseline and at post-therapy had worse DFS and OS compared with patients with CK-19(−) CTCs at both time-points48.
Our findings indicate that in 8/10 (80%) of early breast cancer patients that did not relapse no CK-19 transcripts were detected in the EpCAM(+) CTC fraction. However, there were two patients P#3 and P#6 that did not relapse during the time of the follow-up. In these two patients we detected CK-19 transcripts in EpCAM(+) CTC fractions, but in only one time point out of 14 (1/14, 7%) time points tested for P#3, and in only two timepoints out of 16 (1/16, 6%) for P#6. It is well known that most of CTCs are destroyed during circulation by immune cells, and this very low detection rate of CTC in peripheral blood could possibly be an explanation for the lack of disease recurrence in these two patients.
Three out of these thirteen patients (Pt#11, Pt#12, Pt#13) developed distant metastasis during the follow-up period. In these patients we further analyzed DNA samples extracted from EpCAM(+) cell fractions and paired plasma for the detection of PIK3CA and ESR1 mutations. ESR1 mutations were not detected in any sample for Pt#11 and Pt#12. ESR1 mutations were detected in the plasma ctDNA of Pt#13 consistently at five different time points during the follow-up period. This patient received everolimus and exemestane for a long time, and after disease progression on this therapy, ESR1 mutations were detected in serial samples of plasma ctDNA. Our results are in accordance with those reported in the EROS1 study, where ESR1 mutations were identified in patients previously treated with tamoxifen or aromatase inhibitors, revealing a possible correlation between long term aromatase inhibitor therapy and the existence of ESR1 mutations49. Additional studies have shown that acquired ESR1 mutations are a major mechanism of resistance to aromatase inhibitors50,51,52,53. Our results reflect the reports indicating that ESR1 mutations (especially D538G, Y537S) are associated with more aggressive disease38,53,54. In the phase III PADA-1 trial presented at 2021 San Antonio Breast Cancer Symposium, it was observed that switching from an aromatase inhibitor plus palbociclib to fulvestrant and palbociclib upon early identification of the ESR1 mutation in plasma-before disease progression- the median PFS was doubled. This trial has also shown that ESR1 mutations are rarely detected in plasma-cfDNA of ER+ HER2− MBC patients with no overt resistance to aromatase inhibitor and that the detection of ESR1 mutations was associated with a significantly shorter PFS, suggesting that the existence of ESR1 mutation at baseline could accelerate the outset of resistance to AI-palbociclib55. The first results of this trial have recently been published showing that the early therapeutic targeting of ESR1 mutations in blood results in significant clinical benefit56. It is an important study as the original design explored in PADA-1 might help with tackling acquired resistance with new drugs in future trials56. In our study, PIK3CA mutations were detected in all patients that later developed metastasis. More specifically in Pt#11, E545K mutation was detected in 1/10 (10%) EpCAM(+) CTC fractions but not in paired plasma-ctDNA samples. In Pt#12, E545K mutation was detected in 8/14 (57%) EpCAM(+) CTC fractions and in 1/14 (7%) paired plasma-ctDNA samples checked. In Pt#13, E545K mutation was detected in 2/10 plasma-ctDNA samples but not in paired EpCAM(+) CTC fractions. In the same patient, H1047R mutation was detected in 3/10 (30%) EpCAM(+) CTC fractions and in 2/10 (20%) paired plasma-ctDNA samples. We have already shown before in a direct comparison study using the same blood draw, and the same detection methodology that there is no a 100% concordance in the detection of PIK3CA mutations in CTCs and corresponding ctDNA31. In addition, this discordance may be explained as a result of potential intratumor heterogeneity and as an effect of therapeutic pressure on different cancerous subclones.
We further focused our analysis on Pt#13 that was diagnosed with operable BC in 2010 and died from metastatic breast cancer in 2020. CTC enumeration revealed a bad prognosis for this patient, even before the initiation of systemic chemotherapy, that was based on the relatively high number of CTCs. During the first five years (2010–2015) of follow-up, even before metastasis in the liver was histologically confirmed in September 2015 (time point: 69 months), the patient was constantly positive for the presence of CTCs, CTC counts were constantly more than 5CTCs /23 mL PB and a dramatic rise in CTC counts in September 2014 (timepoint: 57 months) has clearly suggested disease progression at least four years before imaging documentation of metastasis.
Rapid increases in CTC numbers at months 74 and 122, were associated with metastatic disease documented by biopsy 6 months earlier. In this patient, verification of recurrence and administration of systemic therapy was verified by tissue biopsy of the metastatic lesions in the liver and was not based on CTC detection. The increase of the CTC number was associated with clinically and radiologically documented disease progression and the different administered treatments were given according to the national guidelines using approved drugs and combinations.
Molecular characterization of CTCs isolated from this patient by double immunofluorescence demonstrated that practically all the detected cells were CK(+)/HER2(+) and CK(+)/Vimentin(+), while the primary tumor was HER2 negative. Based on our initial data demonstrating that trastuzumab can eliminate HER2+ CTCs57, the patient was enrolled in an open-label randomized Phase II trial of secondary adjuvant trastuzumab in early-stage breast cancer patients harboring HER2+ CTCs after the completion of adjuvant chemotherapy and radiotherapy42 and trastuzumab administration resulted in a clear decrease of the CTCs numbers lasting for 8 months. However, at the time of relapse the significant increase of CTCs was characterized by the presence of exclusively HER2- but EGFR+ CTCs. It is to note that lapatinib also resulted in decreased number of CTCs but for a short period of time. Retrospective molecular analysis of DNA derived from CellSearch cartridges, (P#13, time point: 59 months, Nov 2014) has revealed the presence of the E545K PIK3CA mutation in CTCs that is known to confer resistance to trastuzumab as well as ESR1 mutations (D538G, Y537C), that are now known to confer resistance to the everolimus/exemestane combination. In September 2015 (time point: 69 months), CTC number increased again while imaging studies revealed multiple liver metastatic lesions which were histologically confirmed. CTC enumeration has indicated metastasis at least one year earlier (September 2014, time point: 57 months) than radiologically revealed and histologically confirmed (September 2015, time point: 69 months).
It is remarkable that even one year before clinical manifestation of metastasis, and till 2020, PIK3CA hotspot mutations (E545K and H1047R) and ESR1 mutations (D538G, Y537C, Y537S, Y537N), were continuously detected, not only in CTC-derived DNAs from CellSearch cartridges, but also in plasma-cfDNA, and EpCAM(+) CTC fractions as well. At the same time period (Feb 2016–May 2020), ESR1 methylation, that was also shown to be associated with lack of response to hormonotherapy41, was detected in plasma-cfDNA and corresponding EpCAM(+) CTC fractions, but not in CTC-derived DNAs from CellSearch cartridges. Based on these results, it is evident that while the rise in CTC counts indicated very early the metastatic spread, CTC molecular characterization at the DNA level clearly indicated that the tumor had already developed resistance to the treatment. During Feb 2016–May 2020 gene expression analysis in EpCAM(+) CTCs fractions, has revealed the EMT nature of these cells, since the epithelial marker CK-19 that was expressed in all samples tested, was co-expressed with stem cell markers and EMT markers. CD24-/low/CD44high profile was detected in all samples and the CD24−/low/ALDH1high in 50% of samples analyzed during this period of time. Our results on CK-19, TWIST1, CD24 and ALDH1 expression in CTCs are in accordance to those presented in a previous study58.
Molecular characterization of CTC at the DNA level revealed one year before the clinical manifestation of metastasis the presence of PIK3CA and ESR1 mutations in CTCs that are now known to be highly important for therapy resistance. Our retrospective analysis indicates that these findings were an early indication that disease would not respond to the therapy given to this patient (Trastuzumab, lapatinib, and Afinitor/Aromacin), but at that time this was not known, and moreover Alpelisib was not FDA-approved. Alpelisib was administered to P#13 for 2 months, but severe adverse events (diabetes and septicemia necessitating patient’s hospitalization for 18 days) resulted to early treatment discontinuation. The patient died from disease progression 6 months later.
To the best of our knowledge, this is the first time that early detection of Minimal Residual Disease in breast cancer based on a comprehensive liquid biopsy analysis and a ten year follow-up of operable breast cancer patients is reported. According to our results, CTC analysis provided highly important information for the management of these patients. CTC enumeration as expected indicated at the very beginning whether these patients were at a very high risk for metastatic spread. Our results indicate that a comprehensive liquid biopsy analysis provides highly important information for the therapeutic management of breast cancer patients, and could guide oncologists to select molecular targeted treatments, based on precision medicine. Up to now, the use of CTCs to guide systemic therapy remains controversial, and there are no definitive data to support its clinical utility. However, a number of clinical trials based on liquid biopsy are ongoing and are expected to demonstrate the clinical utility of CTCs59. Our results indicate that comprehensive liquid biopsy analysis could reveal the presence of MRD four years before the appearance of clinically detectable metastatic disease. We strongly believe that interventional clinical trials based on a comprehensive liquid biopsy analysis are highly needed to change clinical practice for the benefit of breast cancer patients.
Methods
Patients
All patients were diagnosed with early and operable ER(+) breast cancer. Peripheral blood (PB) (20 mL) was obtained from all patients at different time points during a long-term follow-up period since diagnosis, two weeks after the removal of the primary tumor. Ten of these patients did not relapse and remained metastasis free during the follow-up period, while three of them relapsed within 5–7 years. In all PB samples EpCAM(+) cell fractions were enriched using immunomagnetic capture beads, and analyzed for CK-19 mRNA expression, and in parallel, ctDNA, isolated from paired plasma using the same blood draw (2 mL), was analyzed for all these patients. All samples were analyzed for ESR1 and PIK3CA mutations in genomic DNA (gDNA), extracted from EpCAM(+) fractions and in paired plasma ctDNA at different time points. CTC enumeration using the CellSearch system (Menarini, Silicon Biosystems) was also performed. For one patient that later developed metastasis, CellSearch analysis was performed at regular intervals within a year for 10 consecutive years and phenotypic characterization of CTCs was also performed through double immunofluorescence. gDNA was extracted from the isolated CTCs from CellSearch cartridges to detect ESR1 and PIK3CA mutations for this patient and all CTC-derived gDNA samples were also analyzed for ESR1 methylation status. All patients have signed informed consent for MRD screening, and the study was conducted in accordance with the Declaration of Helsinki and has been approved the Medical Ethical Committee of the General University Hospital of Heraklion, Crete, Greece (Ethical Allowance: 8756/23-6-2014).
Molecular analysis of CTCs
Gene expression
RNA isolation: cDNA synthesis
All steps including, the isolation of EpCAM(+) fractions, total RNA extraction using Trizol-LS reagent (Invitrogen, USA) were performed as previously described15,60,61. cDNA synthesis was carried out with the High-Capacity RNA-to-cDNA Kit (Applied Biosystems, USA) according to the manufacturer’s protocol in 20μL of total volume reaction. All cDNA samples were kept at − 20 °C until use.
RT-qPCR
RT-qPCR was performed for the following genes: (a) CK-19, (b) EMT-associated marker TWIST-1, and (c) stem-cell markers CD24, CD44, ALDH-1. B2M (beta-2-microglobulin) was used as a reference gene61. RT-qPCR assays for the quantification of CK-19, TWIST-1, CD24, CD44 and ALDH-1 transcripts were performed as previously described14,15,62. The cut-off for CD24, CD44 and ALDH-1 was estimated in respect to HPRT expression as previously reported61 in a group of 10 healthy individuals whose peripheral blood has been analyzed in exactly the same way as patient’s.
DNA analysis
DNA isolation
Genomic DNA (gDNA) was isolated from CellSearch cartridges, and EpCAM(+) CTC fractions at regular timer intervals from initial diagnosis during the follow-up period as follows: (a) CellSearch cartridges: The pre-enriched sample (pre-stained CTCs and WBCs) was aspirated from the corresponding CellSearch cartridges, gDNA was extracted using the QIAamp DNA Micro Kit (Qiagen, Germany) in accordance with manufacturer’s instructions42. (b) EpCAM(+) fractions: Similarly, gDNA from all available EpCAM(+) fractions was extracted from CTCs in Trizol LS (Invitrogen, USA) as previously described30,63. In parallel to the isolation of CTC-derived gDNA, plasma cell-free DNA was isolated as previously described31. The QIAamp Circulating Nucleic Acid Kit (Qiagen) was used to isolate ctDNA from 2.0 mL of plasma according to the manufacturer’s instructions. DNA concentration was measured in all samples, using the Nanodrop ND-1000 spectrophotometer (Thermo Scientific, USA) and DNA integrity was assessed prior to the analysis by amplifying a wild-type region in exon 20 of PIK3CA gene64.
PIK3CA and ESR1 mutations
All available DNA samples were analyzed for PIK3CA hotspot mutations (c.1633G > A: E545K exon 9 and c.3140A > G: H1047R exon 20) as well as for ESR1 hotspot mutations [Y537S (c.1610A > C), Y537C (c.1610A > G), Y537N (c.1609T > A) and D538G (c.1613A > G)] as previously described30,33. For plasma ctDNA analysis, a drop-off ddPCR for screening the ESR1 mutations in exon 8 (Y537S, Y537C, Y537N, D538G and L536R) was performed as previously reported65 in order to detect the presence of the ESR1 mutations before identifying individual mutations using the ESR1-NAPA assay33.
ESR1 methylation
All available DNA samples were also processed to sodium bisulfite (SB) treatment using the EZ DNA Methylation Gold Kit (ZYMO Research Corp., USA) according to manufacturer’s instructions. Only samples that were positive for exon 20 PIK3CA amplification were further processed to SB-treatment. SB-treated DNA was stored at − 80 °C until further use. After the SB-treatment, SB-converted DNA integrity was assessed by a real-time methylation-specific PCR (MSP) for b-actin (ACTB) as previously described41,60. Subsequently, all SB-treated samples were analyzed for ESR1 promoter methylation using real-Time MSP, based on our previously developed and validated protocols41,66.
CTC enumeration
CTC enumeration was performed using the FDA-cleared CellSearch system (Menarini, Silicon Biosystems) in PB samples collected during all these years at regular time intervals for most patients. For Pt#13 CellSearch analysis was performed for 10 consecutive years in peripheral blood draws in sequential patient samples During the first 5 years of follow-up, before the development of clinically confirmed metastatic disease, 23 ml of peripheral blood collected in CellSave preservative tubes was analyzed. Conversely, during the metastatic phase of Pt#13 disease 7.5 mL of peripheral blood were used for CTC enumeration every 3–6 months.
Double immunofluorescence (IF)
Phenotypic characterization of CTCs was performed only for Pt#13. CTCs isolated from 20 ml of peripheral blood were used for their phenotypic characterization during the follow-up. The first 5 mL of blood were discarded to avoid contamination with epithelial cells from the skin. Peripheral blood mononuclear cells (PBMCs) were isolated with Ficoll-Hypaque density gradient (d = 1.077 g/mL) centrifugation at 1800 rpm (600 g) for 30 min. Cytospins of 5 105 PBMCs were prepared and stored at − 80 °C until their use for double staining experiments. The presence of CK-positive cells in PBMCs’ cytospins was investigated using the A45-B/B3 mouse antibody (anti-CK-8, CK-18, CK-19, Micromet Munich) and is further referred as CK+ in the text. At some time points the presence of CTCs in PBMCs’ cytospins was investigated using monoclonal antibodies against ki67 (proliferation marker, Abcam) or/and M30-FITC conjugated (apoptotic marker, Roche Diagnostics, Basel). Double immunofluorescence staining was performed as previously described67,68,69,70,71,72.
Data availability
The data presented in this study are available on request from the corresponding author. The data are not publicly available due to ethical restrictions.
References
Pan, H. et al. 20-Year risks of breast-cancer recurrence after stopping endocrine therapy at 5 years. N. Engl. J. Med. 377, 1836–1846. https://doi.org/10.1056/NEJMoa1701830 (2017).
Pantel, K. & Alix-Panabières, C. Liquid biopsy and minimal residual disease—Latest advances and implications for cure. Nat. Rev. Clin. Oncol. 16, 409–424. https://doi.org/10.1038/s41571-019-0187-3 (2019).
Lianidou, E. & Hoon, D. Circulating tumor cells and circulating tumor DNA, in Tietz Textbook of Clinical Chemistry and Molecular Diagnostics 1111–1144 (2017).
Ma, N. & Jeffrey, S. S. Deciphering cancer clues from blood. Science 1979(367), 1424–1425. https://doi.org/10.1126/science.abb0736 (2020).
Alix-Panabières, C. The future of liquid biopsy. Nature 579, S9. https://doi.org/10.1038/d41586-020-00844-5 (2020).
Lianidou, E. & Pantel, K. Liquid biopsies. Genes Chromosom. Cancer 58, 219–232. https://doi.org/10.1002/gcc.22695 (2019).
Cortés-Hernández, L. E., Eslami, Z. S., Pantel, K. & Alix-Panabières, C. Molecular and functional characterization of circulating tumor cells: From discovery to clinical application. Clin. Chem. 66, 97–104. https://doi.org/10.1373/clinchem.2019.303586 (2020).
Rack, B. et al. Circulating tumor cells predict survival in early average-to-high risk breast cancer patients. J. Natl. Cancer Inst. 106(5), dju066. https://doi.org/10.1093/jnci/dju066 (2014).
Janni, W. J. et al. Pooled analysis of the prognostic relevance of circulating tumor cells in primary breast cancer. Clin. Cancer Res. 22, 2583–2593. https://doi.org/10.1158/1078-0432.CCR-15-1603 (2016).
Bidard, F. C. et al. Efficacy of circulating tumor cell count-driven vs clinician-driven first-line therapy choice in hormone receptor-positive, ERBB2-negative metastatic breast cancer: The STIC CTC randomized clinical trial. JAMA Oncol 7, 34–41. https://doi.org/10.1001/jamaoncol.2020.5660 (2021).
Müller, V. et al. Prognostic relevance of the HER2 status of circulating tumor cells in metastatic breast cancer patients screened for participation in the DETECT study program. ESMO Open https://doi.org/10.1016/j.esmoop.2021.100299 (2021).
Alix-Panabières, C. & Pantel, K. Liquid biopsy: From discovery to clinical application. Cancer Discov. 11, 858–873. https://doi.org/10.1158/2159-8290.CD-20-1311 (2021).
Stathopoulou, A. et al. Real-time quantification of CK-19 mRNA-positive cells in peripheral blood of breast cancer patients using the LightCycler system. Clin. Cancer Res. 9, 5145–5151 (2003).
Stathopoulou, A. et al. A highly specific real-time RT-PCR method for the quantitative determination of CK-19 mRNA positive cells in peripheral blood of patients with operable breast cancer. Int. J. Cancer 119, 1654–1659. https://doi.org/10.1002/ijc.22017 (2006).
Strati, A. et al. Gene expression profile of circulating tumor cells in breast cancer by RT-qPCR. BMC Cancer https://doi.org/10.1186/1471-2407-11-422 (2011).
Xenidis, N. et al. Cytokeratin-19 mRNA-positive circulating tumor cells after adjuvant chemotherapy in patients with early breast cancer. J. Clin. Oncol. 27, 2177–2184. https://doi.org/10.1200/JCO.2008.18.0497 (2009).
Xenidis, N. et al. Differential effect of adjuvant taxane-based and taxane-free chemotherapy regimens on the CK-19 mRNA-positive circulating tumour cells in patients with early breast cancer. Br. J. Cancer 108, 549–556. https://doi.org/10.1038/bjc.2012.597 (2013).
Xenidis, N. et al. Clinical relevance of circulating CK-19 mRNA-positive cells detected during the adjuvant tamoxifen treatment in patients with early breast cancer. Ann. Oncol. 18, 1623–1631. https://doi.org/10.1093/annonc/mdm208 (2007).
Xenidis, N. et al. Predictive and prognostic value of peripheral blood cytokeratin-19 mRNA-positive cells detected by real-time polymerase chain reaction in node-negative breast cancer patients. J. Clin. Oncol. 24, 3756–3762. https://doi.org/10.1200/JCO.2005.04.5948 (2006).
Xenidis, N. et al. Peripheral blood circulating cytokeratin-19 mRNA-positive cells after the completion of adjuvant chemotherapy in patients with operable breast cancer. Ann. Oncol. 14, 849–855. https://doi.org/10.1093/annonc/mdg259 (2003).
Alix-Panabières, C., Mader, S. & Pantel, K. Epithelial-mesenchymal plasticity in circulating tumor cells. J. Mol. Med. (Berl.) 95(2), 133–142. https://doi.org/10.1007/s00109-016-1500-6 (2017).
Li, W. et al. Unraveling the roles of CD44/CD24 and ALDH1 as cancer stem cell markers in tumorigenesis and metastasis. Sci. Rep. https://doi.org/10.1038/s41598-017-14364-2 (2017).
Aktas, B. et al. Stem cell and epithelial-mesenchymal transition markers are frequently overexpressed in circulating tumor cells of metastatic breast cancer patients. Breast Cancer Res https://doi.org/10.1186/bcr2333 (2009).
Kasimir-Bauer, S., Hoffmann, O., Wallwiener, D., Kimmig, R. & Fehm, T. Expression of stem cell and epithelial-mesenchymal transition markers in primary breast cancer patients with circulating tumor cells. Breast Cancer Res. https://doi.org/10.1186/bcr3099 (2012).
Giordano, A. et al. Epithelial-mesenchymal transition and stem cell markers in patients with HER2-positive metastatic breast cancer. Mol. Cancer Ther. 11, 2526–2534. https://doi.org/10.1158/1535-7163.MCT-12-0460 (2012).
Papadaki, M. A. et al. Circulating tumor cells with stemness and epithelial-to-mesenchymal transition features are chemoresistant and predictive of poor outcome in metastatic breast cancer. Mol. Cancer Ther. 18, 437–447. https://doi.org/10.1158/1535-7163.MCT-18-0584 (2019).
Pore, M. et al. Cancer stem cells, epithelial to mesenchymal markers, and circulating tumor cells in small cell lung cancer. Clin. Lung Cancer 17, 535–542. https://doi.org/10.1016/j.cllc.2016.05.015 (2016).
Markou, A. et al. Multiplex gene expression profiling of in vivo isolated circulating tumor cells in high-risk prostate cancer patients. Clin. Chem. 64, 297–306. https://doi.org/10.1373/clinchem.2017.275503 (2018).
André, F. et al. Alpelisib for PIK3CA-mutated, hormone receptor-positive advanced breast cancer. N. Engl. J. Med. 380, 1929–1940. https://doi.org/10.1056/NEJMoa1813904 (2019).
Markou, A. et al. PIK3CA mutational status in circulating tumor cells can change during disease recurrence or progression in patients with breast cancer. Clin. Cancer Res. 20, 5823–5834. https://doi.org/10.1158/1078-0432.CCR-14-0149 (2014).
Tzanikou, E. et al. PIK3CA hotspot mutations in circulating tumor cells and paired circulating tumor DNA in breast cancer: A direct comparison study. Mol. Oncol. 13(12), 2515–2530. https://doi.org/10.1002/1878-0261.12540 (2019).
Markou, A., Tzanikou, E., Ladas, I., Makrigiorgos, G. M. & Lianidou, E. Nuclease-assisted minor allele enrichment using overlapping probes-assisted amplification-refractory mutation system: An approach for the improvement of amplification-refractory mutation system-polymerase chain reaction specificity in liquid biopsies. Anal. Chem. 91, 13105–13111. https://doi.org/10.1021/acs.analchem.9b03325 (2019).
Stergiopoulou, D. et al. ESR1 NAPA assay: Development and analytical validation of a highly sensitive and specific blood-based assay for the detection of ESR1 mutations in liquid biopsies. Cancers (Basel) 13, 1–18. https://doi.org/10.3390/cancers13030556 (2021).
Lampignano, R. et al. A novel workflow to enrich and isolate patient-matched EpCAMhigh and EpCAMlow/negative CTCs enables the comparative characterization of the PIK3CA status in metastatic breast cancer. Int. J. Mol. Sci. https://doi.org/10.3390/ijms18091885 (2017).
Deng, G. et al. Single cell mutational analysis of PIK3CA in circulating tumor cells and metastases in breast cancer reveals heterogeneity, discordance, and mutation persistence in cultured disseminated tumor cells from bone marrow. BMC Cancer https://doi.org/10.1186/1471-2407-14-456 (2014).
Pestrin, M. et al. Heterogeneity of PIK3CA mutational status at the single cell level in circulating tumor cells from metastatic breast cancer patients. Mol. Oncol. 9, 749–757. https://doi.org/10.1016/j.molonc.2014.12.001 (2015).
Carausu, M. et al. ESR1 mutations: A new biomarker in breast cancer. Expert Rev. Mol. Diagn. 19, 599–611. https://doi.org/10.1080/14737159.2019.1631799 (2019).
Paolillo, C. et al. Detection of activating estrogen receptor gene (ESR1) mutations in single circulating tumor cells. Clin. Cancer Res. 23, 6086–6093. https://doi.org/10.1158/1078-0432.CCR-17-1173 (2017).
Franken, A. et al. Detection of ESR1 mutations in single circulating tumor cells on estrogen deprivation therapy but not in primary tumors from metastatic luminal breast cancer patients. J. Mol. Diagn. 22, 111–121. https://doi.org/10.1016/j.jmoldx.2019.09.004 (2020).
Hanahan, D. Hallmarks of cancer: New dimensions. Cancer Discov. 12, 31–46. https://doi.org/10.1158/2159-8290.CD-21-1059 (2022).
Mastoraki, S. et al. ESR1 methylation: A liquid biopsy-based epigenetic assay for the follow-up of patients with metastatic breast cancer receiving endocrine treatment. Clin. Cancer Res. 24, 1500–1510. https://doi.org/10.1158/1078-0432.CCR-17-1181 (2018).
Georgoulias, V. et al. Trastuzumab decreases the incidence of clinical relapses in patients with early breast cancer presenting chemotherapy-resistant CK-19mRNA-positive circulating tumor cells: Results of a randomized phase II study. Ann. Oncol. 23, 1744–1750. https://doi.org/10.1093/annonc/mds020 (2012).
Mavroudis, D. et al. Six versus 12 months of adjuvant trastuzumab in combination with dose-dense chemotherapy for women with HER2-positive breast cancer: A multicenter randomized study by the Hellenic Oncology Research Group (HORG). Ann. Oncol. 26, 1333–1340. https://doi.org/10.1093/annonc/mdv213 (2015).
Stathopoulou, A. et al. Molecular detection of cytokeratin-19-positive cells in the peripheral blood of patients with operable breast cancer: Evaluation of their prognostic significance. J. Clin. Oncol. 20, 3404–3412. https://doi.org/10.1200/JCO.2002.08.135 (2002).
Ignatiadis, M. et al. Prognostic value of the molecular detection of circulating tumor cells using a multimarker reverse transcription-PCR assay for cytokeratin 19, mammaglobin A, and HER2 in early breast cancer. Clin. Cancer Res. 14, 2593–2600. https://doi.org/10.1158/1078-0432.CCR-07-4758 (2008).
Saloustros, E. et al. Cytokeratin-19 mRNA-positive circulating tumor cells during follow-up of patients with operable breast cancer: Prognostic relevance for late relapse. Breast Cancer Res. 13, R60. https://doi.org/10.1186/bcr2897 (2011).
Georgoulias, V. et al. Effect of front-line chemotherapy on circulating CK-19 mRNA-positive cells in patients with metastatic breast cancer. Cancer Chemother. Pharmacol. 74, 1217–1225. https://doi.org/10.1007/s00280-014-2598-2 (2014).
Matikas, A. et al. Detection of circulating tumour cells before and following adjuvant chemotherapy and long-term prognosis of early breast cancer. Br. J. Cancer 126(11), 1563–1569. https://doi.org/10.1038/S41416-022-01699-5 (2022).
Najim, O. et al. The prevalence of estrogen receptor-1 mutation in advanced breast cancer: The estrogen receptor one study (EROS1). Cancer Treat. Res. Commun. https://doi.org/10.1016/j.ctarc.2019.100123 (2019).
Schiavon, G. et al. Analysis of ESR1 mutation in circulating tumor DNA demonstrates evolution during therapy for metastatic breast cancer. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.aac7551 (2015).
Jeselsohn, R. et al. Emergence of constitutively active estrogen receptor-α mutations in pretreated advanced estrogen receptor-positive breast cancer. Clin. Cancer Res. 20, 1757–1767. https://doi.org/10.1158/1078-0432.CCR-13-2332 (2014).
Toy, W. et al. Activating ESR1 mutations differentially affect the efficacy of ER antagonists. Cancer Discov. 7, 277–287. https://doi.org/10.1158/2159-8290.CD-15-1523 (2017).
Chandarlapaty, S. et al. Prevalence of ESR1 mutations in cell-free DNA and outcomes in metastatic breast cancer: A secondary analysis of the BOLERO-2 clinical trial. JAMA Oncol. 2, 1310–1315. https://doi.org/10.1001/jamaoncol.2016.1279 (2016).
Paoletti, C. et al. Comprehensive mutation and copy number profiling in archived circulating breast cancer tumor cells documents heterogeneous resistance mechanisms. Can. Res. 78, 1110–1122. https://doi.org/10.1158/0008-5472.CAN-17-2686 (2018).
Bidard, F. C. et al. Prognostic impact of ESR1 mutations in ER+ HER2− MBC patients prior treated with first line AI and palbociclib: An exploratory analysis of the PADA-1 trial. 38, 1010–1010. https://doi.org/10.1200/JCO.2020.38.15_suppl.1010 (2020).
Bidard, F. C. et al. Switch to fulvestrant and palbociclib versus no switch in advanced breast cancer with rising ESR1 mutation during aromatase inhibitor and palbociclib therapy (PADA-1): A randomised, open-label, multicentre, phase 3 trial. Lancet Oncol. https://doi.org/10.1016/S1470-2045(22)00555-1 (2022).
Bozionellou, V. et al. Trastuzumab administration can effectively target chemotherapy-resistant cytokeratin-19 messenger RNA-positive tumor cells in the peripheral blood and bone marrow of patients with breast cancer. Clin. Cancer Res. 10, 8185–8194. https://doi.org/10.1158/1078-0432.CCR-03-0094 (2004).
Bredemeier, M. et al. Gene expression signatures in circulating tumor cells correlate with response to therapy in metastatic breast cancer. Clin. Chem. 63, 1585–1593. https://doi.org/10.1373/clinchem.2016.269605 (2017).
Vasseur, A., Kiavue, N., Bidard, F. C., Pierga, J. Y. & Cabel, L. Clinical utility of circulating tumor cells: An update. Mol. Oncol. 15, 1647–1666. https://doi.org/10.1002/1878-0261.12869 (2021).
Strati, A. et al. Prognostic significance of PD-L1 expression on circulating tumor cells in patients with head and neck squamous cell carcinoma. Ann. Oncol. 28, 1923–1933 (2017).
Zavridou, M. et al. Evaluation of preanalytical conditions and implementation of quality control steps for reliable gene expression and DNA methylation analyses in liquid biopsies. Clin. Chem. 64, 1522–1533. https://doi.org/10.1373/clinchem.2018.292318 (2018).
Strati, A., Nikolaou, M., Georgoulias, V. & Lianidou, E. Prognostic significance of TWIST1, CD24, CD44, and ALDH1 transcript quantification in EpCAM-positive circulating tumor cells from early stage breast cancer patients. Cells 8, 652. https://doi.org/10.3390/cells8070652 (2019).
Chimonidou, M. et al. DNA methylation of tumor suppressor and metastasis suppressor genes in circulating tumor cells. Clin. Chem. 57, 1169–1177. https://doi.org/10.1373/clinchem.2011.165902 (2011).
Vorkas, P. A. et al. PIK3CA hotspot mutation scanning by a novel and highly sensitive high-resolution small amplicon melting analysis method. J. Mol. Diagn. 12, 697–704. https://doi.org/10.2353/jmoldx.2010.100008 (2010).
Jeannot, E. et al. A single droplet digital PCR for ESR1 activating mutations detection in plasma. Oncogene https://doi.org/10.1038/s41388-020-1174-y (2020).
Chimonidou, M. et al. Direct comparison study of DNA methylation markers in EpCAM-positive circulating tumour cells, corresponding circulating tumour DNA, and paired primary tumours in breast cancer. Oncotarget 8, 72054–72068. https://doi.org/10.18632/oncotarget.18679 (2017).
Kallergi, G. et al. Apoptotic circulating tumor cells in early and metastatic breast cancer patients. Mol. Cancer Ther. 12, 1886–1895. https://doi.org/10.1158/1535-7163.MCT-12-1167 (2013).
Kallergi, G., Mavroudis, D., Georgoulias, V. & Stournaras, C. Phosphorylation of FAK, PI-3K, and impaired actin organization in CK-positive micrometastatic breast cancer cells. Mol. Med. 13, 79–88. https://doi.org/10.2119/2006-00083.Kallergi (2007).
Kallergi, G. et al. Phosphorylated EGFR and PI3K/Akt signaling kinases are expressed in circulating tumor cells of breast cancer patients. Breast Cancer Res. https://doi.org/10.1186/bcr2149 (2008).
Kallergi, G. et al. Hypoxia-inducible factor-1α and vascular endothelial growth factor expression in circulating tumor cells of breast cancer patients. Breast Cancer Res. https://doi.org/10.1186/bcr2452 (2009).
Spiliotaki, M. et al. Evaluation of proliferation and apoptosis markers in circulating tumor cells of women with early breast cancer who are candidates for tumor dormancy. Breast Cancer Res. https://doi.org/10.1186/s13058-014-0485-8 (2014).
Kallergi, G. et al. Expression of truncated human epidermal growth factor receptor 2 on circulating tumor cells of breast cancer patients. Breast Cancer Res. https://doi.org/10.1186/s13058-015-0624-x (2015).
Acknowledgements
The current research was partly supported by Stavros Niarchos Foundation within the framework of a Grant to the National and Kapodistrian University of Athens, Grant No. 16785. We acknowledge all the patients’ contribution who participated in this study. Especially, we would like to express our deep thanks to our late patient #13 who was so positive in giving her blood samples for ten years throughout this study, since she believed that Liquid Biopsy research will help many patients to come.
Author information
Authors and Affiliations
Contributions
Conceptualization, E.L. and V.G.; Methodology: D.S., A.M., A.S., M.Z., E.T., S.M., G.K.; Validation, D.S, G.K., E.L.; Formal Analysis, D.S., A.M., A.S., M.Z., E.T., S.M., G.K, E.L.; Investigation: D.S., A.M., A.S., M.Z., E.T., S.M., G.K.; Resources, E.L. V.G.; Data Curation, D.S., A.M., A.S., M.Z., G.K., V.G., E.L.; Writing—Original Draft Preparation, D.S., V.G., E.L.; Writing—Review and Editing, V.G., E.L.; Supervision, E.L.; Project Administration, E.L.; Funding Acquisition, E.L.
Corresponding author
Ethics declarations
Competing interests
Prof Evi Lianidou and Dr. Athina Markou are inventors in the following patent: Method of determining PIK3CA mutational status in a sample. Inventors: Lianidou E, Markou A. (http://www.freepatentsonline.com/WO2016020710A1.html). Authors state no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Stergiopoulou, D., Markou, A., Strati, A. et al. Comprehensive liquid biopsy analysis as a tool for the early detection of minimal residual disease in breast cancer. Sci Rep 13, 1258 (2023). https://doi.org/10.1038/s41598-022-25400-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-25400-1
This article is cited by
-
Nanobiotechnology augmented cancer stem cell guided management of cancer: liquid-biopsy, imaging, and treatment
Journal of Nanobiotechnology (2024)
-
Liquid biopsy: from concept to clinical application
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.