Abstract
Tenofovir disoproxil fumarate (TDF) is one of the nucleotide analogs capable of inhibiting the reverse transcriptase (RT) activity of HIV and hepatitis B virus (HBV). There is no known HBV resistance to TDF. However, detectable variation in duration of HBV persistence in patients on TDF therapy suggests the existence of genetic mechanisms of on-drug persistence that reduce TDF efficacy for some HBV strains without affording actual resistance. Here, the whole genome of intra-host HBV variants (N = 1,288) was sequenced from patients with rapid (RR, N = 5) and slow response (SR, N = 5) to TDF. Association of HBV genomic and protein polymorphic sites to RR and SR was assessed using phylogenetic analysis and Bayesian network methods. We show that, in difference to resistance to nucleotide analogs, which is mainly associated with few specific mutations in RT, the HBV on-TDF persistence is defined by genetic variations across the entire HBV genome. Analysis of the inferred 3D-structures indicates no difference in affinity of TDF binding by RT encoded by intra-host HBV variants that rapidly decline or persist in presence of TDF. This finding suggests that effectiveness of TDF recognition and binding does not contribute significantly to on-drug persistence. Differences in patterns of genetic associations to TDF response between HBV genotypes B and C and lack of a single pattern of mutations among intra-host variants sensitive to TDF indicate a complex genetic encoding of the trait. We hypothesize that there are many genetic mechanisms of on-drug persistence, which are differentially available to HBV strains. These pervasive mechanisms are insufficient to prevent viral inhibition completely but may contribute significantly to robustness of actual resistance. On-drug persistence may reduce the overall effectiveness of therapy and should be considered for development of more potent drugs.
Similar content being viewed by others
Introduction
Resistance of viral strains to drugs is an important problem for patient management and public health. Viral drug resistance is usually associated with simple patterns of mutations involving only a few genomic sites1,2,3. One of the most studied and effective drugs are nucleotide and nucleoside analogs4. In hepatitis B virus (HBV), these analogs inhibit reverse transcriptase (RT) activity5,6. Development of HBV drug resistance is caused by specific viral mutations directly affecting recognition and binding of the analogs7 or excision of chain terminators by RT8, and may be accompanied by complementary mutations that correct fitness reduction usually associated with the primary mutation9. These patterns of mutations are generally referred to as a genetic barrier to resistance. Genetic patterns of greater complexity engender a greater genetic barrier to development of resistance10.
Although drug resistant mutations have a strong phenotypic effect, they are not independent from other genomic sites and genetic composition of the intra-host viral population. Estimates of the rates of mutation and viral replication indicate that all possible single and double mutations, and a large fraction of possible triple mutations are generated during each day of viral replication in infected hosts11,12, making many simple mutation patterns associated with drug resistance readily available to essentially any intra-host viral population. Nevertheless, despite such a wide presentation of drug-resistance mutations, not all viral strains develop resistance, indicating that phenotypic effects of these mutations are dependent on the genetic background to which they occur, emphasizing a significant role of epistasis and coevolution among viral genomic sites in development of resistance13,14. Therefore, strength of the genetic barrier is associated not only with complexity and availability of genetic changes required for resistance to viral population but with the overall fitness effects of these mutations in the background of the strain genetic composition13.
Worldwide, an estimated 248 million people had chronic HBV infection in 201515. HBV is a small DNA virus, the replication of which involves an RT step16. Thus, drugs developed to control the human immunodeficiency virus (HIV) RT activity are also effective against HBV RT. Many of these drugs have a low HBV genetic barrier to resistance, resulting in frequent development of resistance1,2. Tenofovir disoproxil fumarate (TDF) is one of the nucleotide analogs effectively inhibiting HIV and HBV replication. Unlike many drugs, however, TDF is not associated with development of HBV drug resistance17,18,19,20. Nevertheless, TDF treatment results in a variable duration of HBV replication in different patients. Some patients experience a rapid HBV clearance, while others have detectable HBV after almost 2 years of TDF treatment21,22. While treatment response depends on a variety of host and viral factors, such differences could also be a result of genetic mechanisms that reduce TDF efficacy for some HBV strains without affording actual resistance.
Here, we show that differential responses to TDF are associated with genetic composition of the entire HBV genome, with RT playing a non-dominant role. The findings suggest the existence of various HBV genetic mechanisms contributing in different combinations to the rate of the HBV on-TDF persistence. Without causing resistance, the on-drug persistence mechanisms may mitigate effects of many HBV RT-inhibiting drugs, thus should be considered for the development of more potent therapies.
Results
Changes in intra-host HBV population
1,288 whole genome HBV sequences (954 unique sequences) were obtained from ten patients (P1–P10), with 114–171 (44–122 unique) intra-host sequence variants obtained from each patient. HBV sequences were obtained at three time-points (0, 4 and 40 weeks after initiation of TDF therapy) from all slow response (SR) patients. However, no sequences were obtained from specimens collected at week 40 from rapid response (RR) patients P5, P6 and P7, and only five and six sequences were obtained from RR patients P4 and P9, correspondingly.
Among the RR cases, two (P6 and P7) were infected with a heterogeneous HBV population composed of many low-frequency variants at the baseline, which remained similarly diverse at week 4. However, three cases (P4, P5 and P9) were infected with HBV populations that had dominant or high-frequency variants. Although the dominant variants varied in frequency between time-points, they were present at week 4 (Fig. 1) but were replaced later with different variants in two cases, P4 and P9, who remained HBV PCR positive at week 40. Thus, in all RR cases, viral population largely preserved its structure at baseline and week 4, and experienced delayed shift, losing dominant variants, at week 40 in two cases, P4 and P9, who were still PCR-positive at the third time-point.
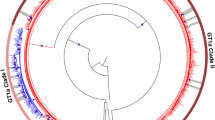
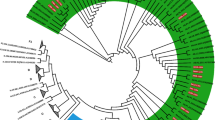
Heterogeneity and phylogeny of HBV quasispecies strains of genotype (GT) C, B and E. Shown are the median joining networks (MJN) of the full HBV genomic quasispecies sequences sampled from (A) five SR patients (P1-P3, P8 and P10) and from (B) five RR patients (P4-P7 and P9), and (C) phylogenetic tree. Sequences were sampled at three time points: baseline and week 4 and 40 during TDF therapy. GT C sequences organized into two clades or clusters (C1 and C2). Nodes in MJN and in phylogram tree represent HBV variants. Nodes colored based on time point of sampling (as denoted in color legend).
Among SR cases, two (P2 and P3) were infected with diverse HBV populations, containing only low-frequency variants at baseline, which became less heterogeneous at week 4 and 40. Both cases contained a high-frequency variant at week 4, which persisted throughout the observation to week 40 in P2 but was detectable only at week 4 in P3. The other three SR cases (P1, P8 and P10) had a high-frequency variant at baseline. In P1, a single variant was continuously dominant at all three time-points. In P8, the initial major variant was replaced with a different one at week 4, which remained detectable at week 40. In P10, the initial dominant variant declined in frequency by week 4 and turned undetectable at week 40 (Fig. 1).
Thus, while no substantial changes in the intra-host population occurred between weeks 0 and 4 among all RR cases, the intra-host HBV population experienced a detectable shift in three SR cases (P2, P3 and P8) between these two time-points. The rapid shifts in intra-host population of P2, P3 and P8, accompanied by increase in frequency of certain intra-host variants in P2 and P3 or by replacement of the dominant variant in P8, within 1 month after initiation of therapy indicate a capacity of these SR-HBV strains to adapt to TDF within a short period of time. This results in a slow HBV decline in patients on treatment, while none of the RR strains could produce intra-host variants of a similar replicative strength on TDF. Persistence of the major variant in P1 indicates that this SR strain was less sensitive to TDF initially, which, together with the observation of significant population shifts in the other three SR cases, suggests differential sensitivity of intra-host HBV variants to TDF, being especially detectable among SR strains.
SR/RR-associated mutations
Considering significant effects of TDF on the structure of intra-host HBV populations, it is conceivable that HBV variants from the SR group may carry specific mutations affording a greater protection against TDF than mutations in variants from the RR group. Inspection of the nucleotide (nt) sequence alignment of all intra-host variants did not reveal any mutations completely specific to RR or SR. However, application of the correlation-based feature selection (CFS) algorithm23 allowed for identification of 16 nt sites associated with RR or SR classes (Table 1). These sites were found to be scattered across all HBV genes (C, X, P and S), with nine mutations affecting genomic regions encoding all four domains (terminal protein (TP), spacer (Sp), RT and RNAse H) of the P protein. Among the 16 sites, 5 are 3rd positions of codons of the P (N = 4) and C (N = 1) open reading frames (ORF) that are in genomic regions outside of the ORF overlap (Table 1). The association of the 16 nt sites with RR/SR was confirmed by the targeted analysis using naïve Bayesian Network (BN) (Fig. 2). BN analysis showed a significant association of polymorphism at site 2573 with RR/SR classes (Kullback–Leibler divergence (KL) = 0.78; P < 0.001). All intra-host HBV variants (N = 458) sampled from RR cases (P4, P5, P6, P7 and P9) had cytosine at this site, while intra-host variants (N = 393) from 4 SR cases (P1, P2, P8 and P10) had thymine at this site. Only SR P3 had intra-host HBV variants (N = 103) containing cytosine at site 2573. Moreover, the naïve BN was found to have accuracy of 99.7% (95% CI 99.4–100) in leave-one-out cross-validation tests, while achieving the expected accuracy (~ 50.0%) in randomly labeled data (Tables S1 and Tables S2 in SI).
Relevant nt sites associated to the RR/SR. BN generated using full HBV genomic quasispecies sequences (N = 954) of GT C, B and E sampled from ten patients at three time points: baseline, and week 4 and 40 during TDF therapy. Round nodes in the graph represent 16 polymorphic nt sites (Table 1) and the square node represents the response (“target”) variable. Coloring of round nodes based on genomic region (see legend Fig. 4). Dependencies (relationships) between the response and nt sites are displayed as blue arcs and inter-dependencies between the sites as black arcs. The average strength of the relationship between a node and the target was small but significant (KL = 0.19, P < 0.05). However, four relationships in the network—arcs between the target and nodes representing genome positions (p): 866, 1946, 2075 and 2441—could not be statistically supported (P > 0.05). Nonetheless, this BN was found useful for prediction of RR/SR association (Tables S1 and Tables S2, in SI).
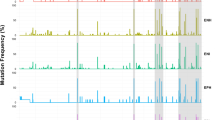
Additionally, a self-organizing artificial neural network (ANN) model24,25 constructed using the 16 nt sites shown in Table 1 (see “SI Methods”) showed a clear partitioning of HBV variants into two clusters concordant with RR/SR (Fig. 3). Among HBV genotype C (HBV/C) and HBV genotype B (HBV/B), the model accurately identified 97.2% of the RR-associated sequences. Physicochemical profiles of the 16 nt sites from only 13 sequence variants of the RR HBV/B strain infecting P9 (six from baseline; six from week 4, and one from week 40) were similar to variants (N = 496) obtained from SR patients infected with HBV/B, HBV/C and HBV/E. Thus, although genetic analysis did not allow for identifying a single mutation clearly distinguishing RR and SR, a combination of several mutations scattered across the entire HBV genome were found to be strongly associated with RR/SR as a group, suggesting that on-drug persistence is complexly encoded in the HBV genome.
Physicochemical clustering of HBV sequences. Shown is the SOTA-based24,25 grouping, by (A) response and by (B) genotype, of physicochemical profiles representing full HBV quasispecies sequences (N = 954) collected from ten TDF-treated patients at three time points. Group 1 (neuron 3, in red) was mostly (97.4%) comprised of HBV GT C (clusters C1 and C2—see Fig. 1C), B and E strains sampled from SR patients. Group 2 (neuron 2, in cyan) was comprised of strains derived solely from RR HBV/C- & B-infected patients. Only thirteen out of the 47 unique variants sampled from one HBV/B-infected RR patient were found to be members of Group 1. Red arrow denotes boundary between the two groups. The 16 nt-based physicochemical profile representation of HBV sequences was generated using a scale of five physicochemical properties of DNA nt’s26.
The RR/SR association among HBV genotype C strains
Considering a significant genetic diversity among HBV genotypes27,28, it is conceivable that molecular mechanisms responsible for the on-TDF persistence may be specific for each genotype. Thus, complex genetic encoding of the persistence suggested here may result from a set of simple genotype-specific genetic associations. Indeed, among the ten HBV strains used in this study, only six belonged to a single genotype C, while the other four belonged to genotypes B (N = 3) and E (N = 1). Since HBV/C strains were most represented, HBV/C sequences (N = 799) were used to generate BN (details in “SI methods”) to explore genotype-specific genetic associations with RR/SR. Among 1,020 HBV/C polymorphic genomic sites (~ 32% of all sites), 77.0% did not form significantly strong associations (SC ≥ 1; p ≤ 0.05) to each other (see Fig. S1 in SI). Meanwhile, ~ 21.0% of the sites were organized into a major BN component that included the RR/SR variable (Fig. 4), suggesting a certain genetic association of the involved sites with the on-TDF persistence. The BN was evaluated by Bayesian testing using Bayes factors (Bf)29,30 to measure the statistical significance of the influence of each state of every polymorphic site on the RR/SR state (see “SI methods”). Our analysis showed that nt states of polymorphic sites disconnected from the major component (N = 30) were non-informative with respect to the RR/SR state (exhibited neutral Bf; Bf = 0). However, Bf showed a strong association of 39.0% of nt states for 203 polymorphic sites composing the major BN component with the RR/SR as well as with phylogenetic clustering (see “SI methods”).
Genome-wide dependencies among polymorphic sites. BN generated using 799 whole-genome quasispecies from 6 HBV/C-infected patients treated with TDF (see “BN Section” in SI). Round nodes in the graph represent polymorphic nt sites and arcs represent significant (SC = 1.0, P < 0.001) dependency relationships. The BN comprises a major 215-varaible component (212 nt sites, and the response, time-point and phylogenetic cluster variables—square nodes in yellow) and 11 minor components representing 30 nt sites. Node coloring based on nine regions (genome positions in parenthesis): overlapping S–P genes (1–837), RT domain (838–1163), RNAse H domain (1164–1375), overlapping RNAse H–X (1376–1625), X gene (1626–1815), overlapping X–C genes (1816–1840), C gene (1841–2308), overlapping C–P genes (2309–2454) and Terminal protein domain (2455–3215). Position numbering based on reference sequence: GenBank accession number AY233278.
To sort out genetic associations with RR/SR vs. associations with phylogenetic clusters and time-points (Cluster and TP nodes in Fig. 4), we conducted target analysis (see “SI methods”). Among the 203 sites, 23 were found to be more significantly (KL ≥ 0.68; P < 0.001) associated with RR/SR than with phylogenetic cluster or time point (Fig. S2 in SI). The majority of these sites (N = 16) were located in three domains of the P protein: TP (N = 5), RT (N = 5) and RNAse H (N = 6). It is important that 4 of the 6 RNAse H sites (genome positions 1223, 1231, 1322 and 1501) were most strongly associated to RR/SR (Fig. S2B in SI). When the state for these sites is known, the state of the RR/SR variable becomes independent of the other variables in the BN (Table 2). Target analysis conducted without the 4 sites from RNAse H showed that the state of RR/SR variable can be accurately estimated from 72 polymorphic sites of the major BN component (Fig. 4). These sites had stronger associations (KL values ranging from 0.10 to 0.48) to RR/SR than to PC and T (Fig. S3 in SI). Considered together with Bf for each site in the BN (Fig. 4), these results suggest abundance of genetic pathways affecting RR or SR phenotypes, indicating involvement of many more genetic mechanisms in the on-drug persistence than usually associated with development of resistance. Although RNAse H seemingly plays a particularly important role in defining the rate of response to TDF for HBV/C strains, the identified 4 RNAse H sites had no association with RR/SR for HBV/B strains (data not shown), indicating differences in genetic mechanisms of on-drug persistence between these two genotypes.
The RR/SR association of the HBV/C P protein
HBV drug resistance to nucleotide analogs is usually afforded by amino acid (aa) changes in RT of the P protein1,2,3. Although our analysis showed that nucleotide changes associated with RR/SR are distributed across the entire HBV genome, aa substitutions in P should be expected to play an important role in defining the rate of TDF response. Taking into consideration the aforementioned genotype specificity of the on-drug persistence, analysis was performed using P protein aa sequences from HBV/C only. It was found that 48 of 265 aa polymorphic sites from P form a major BN component (SC = 0.95; P < 0.001), with seven of these sites (positions in P: 307, 321, 624, 713, 743, 803 and 828) having a strong association (KL ≥ 0.41) with the RR/SR variable as determined by target analysis (Fig. S4 in SI). Importantly, the identified aa sites matching the nt sites showing a strong association with RR/SR (Table1 and Fig. S4 in SI).
Generally, nucleotide analogs inhibit the HBV RT activity. Our findings support an important role of RT in defining the HBV RR/SR phenotypes. To examine a potential contribution of the RT nt and aa variability to the rate of response to TDF among HBV/C strains, we conducted additional analyses. Prior to constructing a BN, all baseline sequences were initially mined and analyzed using the CFS algorithm (details in “SI methods”) to select a minimal subset of nt or aa polymorphic sites that maximize the conditional (posterior) probability of observing RR or SR. A minimum subset of 10 nt polymorphic sites (genome positions: 253, 280, 376, 458, 708, 828, 836, 926, 995 and 1006) as potential predictors (Merit = 0.527) of RR/SR was identified (Fig. S5 in SI). The nt polymorphic sites at position 836, 926 and 995 in RT, which were observed to be strongly associated (KL > 0.21; P < 0.001) with RR/SR in the BN (Fig. S3 in SI), were also selected by CFS. Although association of sites at position 253, 458 and 1006 to RR/SR was not statistically supported (P > 0.05), robust classification (CA = 100%) into RR or SR by a model using all these 10 selected nt sites (Fig. S5 in SI) suggests that the RT nt sequence contains information pertinent to the rate of response to TDF among HBV/C strains.
In addition, CFS analysis applied to the RT protein sequences (N = 247) sampled at baseline from the 6 HBV/C-infected patients identified a minimum subset of 22 polymorphic aa sites (Table S3 in SI), which were strongly associated (Merit = 0.511) as a group with RR/SR. BN constructed using these sites was shown to detect RR or SR with high accuracy (CA = 98.8%), while achieving the expected accuracy (CA = 53.4%) on random-labeled data. It is interesting that among the selected 22 sites, 6 located at the RT positions 82, 139, 153, 191, 223 and 233 have been reported as related to drug resistance31,32,33. Thus, the findings of groups rather than individual nt or aa sites strongly associated with RR/SR indicate that the differential sensitivity of HBV/C to inhibition by TDF is not defined by a single mutation and most probably involves either a single compound function or several simple functions of RT.
Protein 3D-structure mapping
The 22-selected polymorphic aa sites were mapped onto the 3D-structures of RT to identify structural effects of mutations at these sites that can potentially explain RR/SR phenotypes. Using the predicted HBV-RT/DNA-RNA/TFV-DP protein–ligand complexes34, the 3D-models were constructed for major HBV GT/C variants from 2 SR cases (P1 and P2) and an RR case (P5) (Fig. S6 in SI). Two aa sites, L147 and K239, were found to be in vicinity to the nt binding pocket (Fig. 5). Among all polymorphic aa sites in HBV/C RT, 2 other sites were identified that can potentially affect binding nt and DNA directly: V191, located in the alpha helix structure forming the nt binding pocket interface, and Q288, located in the alpha helix structure forming the DNA binding interface in the RT thumb domain. The states of these four sites were, however, identical among the three studied here HBV/C RT variants, indicating that genetic variation at these sites do not have a clear effect on the on-TDF persistence at least for the HBV variants from the three cases.
Mapping of HBV/C RT polymorphic aa sites relevant to SR/RR association. Shown is the predicted 3D structure of the HBV-RT/DNA-RNA/TFV-DP complex representative of the 345aa-long RT protein of HBV GT C strains34. The 22 aa sites represented in the RT-BNC (Table S3 in SI) are denoted as sticks (in purple). Potential effectors (N = 4) of the ligand–protein interaction are marked with corresponding RT positions. RT 3D-structure coloring scheme: fingers, in cyan and gold; palm, in green, and thumb, in red. TDF and DNA/RNA ligands are depicted with ball-and-stick (cpk colors) and cartoon (grey color) representations, respectively. Rendering was done using the VMD software35. Position numbering based on reference sequence: GenBank accession number AF458665.1.
Binding patterns of TDF to RT variants
To further investigate potential roles of the 22 aa polymorphic sites, analysis was conducted to characterize interaction between the diphosphorylated tenofovir (TFV-DP) and HBV RT. Analysis was performed using five predicted HBV-RT/DNA-RNA/TFV-DP protein–ligand complexes34 for the three aforementioned major HBV/C variants (from P1, P2 and P5) and two additional major variants from GT/B strains, one from RR patient (P9) and another from SR patient (P8) (Fig. S6 in SI). Analysis indicates that the TFV-DP binds near the YMDD motif (HBV RT active site; RT positions 203–206), with M204 and D205 contributing hydrophobic and negative-charge contacts, respectively (Fig. S7 in SI). The triphosphate end of TFV-DP is stably anchored to the binding pocket by hydrogen bonds (H-bonds) with Y148, T150, R110 and K149, and is strongly coupled by an Mg2+ chelation network. The detected residue-specific interactions and metal coordination coupling persist throughout MD simulations in all the predicted protein complexes of HBV/C and HBV/B (Table S4 in SI). The base-ring end of the TFV-DP is anchored by two persisting H-bonds with U-bases from the template RNA. Analysis also indicates that sites L147 and K239, mapped in vicinity of the binding pocket (Fig. 5), contribute to the TFV-DP binding interaction, providing, respectively, hydrophobic and positive-charge contacts (Fig. S7 in SI). Thus, the data, summarized in Table S4 (in SI), show no substantial differences in TFV-DP-binding patterns among the studied HBV RT variants, suggesting that the TDF binding to RT is not strongly associated with RR/SR.
Discussion
Genetic analyses conducted here indicate that the differential response to TDF among HBV strains is not associated with a specific mutation, a unique mutation pattern or a single HBV protein. Rather, capacity to the on-TDF persistence is a complex genetic trait, which is intricately encoded across the entire HBV genome. Identification of 16 sites from different HBV genomic regions (Table 1), which are strongly associated as a group with the TDF response, suggests the existence of a few compound or many simple genetic pathways contributing to the TDF response. These sites are distributed across all HBV genes, with nine sites located in genomic regions encoding all four structural domains of the P protein. Only three sites, one of which is synonymous, were found to be in the RT domain, indicating that RT does not play a major role in defining HBV persistence on TDF. Thus, there is an important difference between molecular mechanisms responsible for resistance to nucleotide and nucleoside analogs and for on-TDF persistence. While functional dominance of RT mutations in development of resistance is well established31,32,33, contribution of RT to controlling the level of HBV replication during TDF treatment, although essential, is seemingly limited. This conclusion is supported by the lack of difference in TDF binding by RT from persistent and rapidly declining HBV variants in the 3D-models implemented here. Thus, a potential involvement of all HBV proteins and limited contribution of RT to the protracted HBV replication on TDF suggest that genetic mechanisms of on-drug persistence and drug resistance are essentially different.
Molecular mechanisms of resistance to nucleotide and nucleoside analogs generally lead to a significantly reduced drug recognition and/or binding by the RT active center7 or excision of chain terminators8, which result in indefinite survival of the resistant HBV strain in presence of drug. Usually, simple mutation patterns are associated with resistance1,2,3. Thus, small genetic changes result in a very strong phenotypic effect, making specific drug resistance readily selectable for many viral strains. In contrast, protection afforded by on-TDF persistence is incomplete. It only slows the decline of HBV population during treatment. A lesser phenotypic effect associated with complex mutation patterns seemingly makes on-drug persistence less selectable during TDF treatment. However, phylogenetic analysis showed rapid genetic changes in intra-host HBV populations of 3 SR strains (P2, P3 and P8) and continuous presentation of a dominant HBV variant in P1 during treatment, indicating a degree of adaptation of HBV population to TDF and certain resilience of some variants on treatment. The structure of intra-host HBV populations of RR strains, though, does not change as rapidly, showing inability to adapt to TDF. These observations indicate that not only do HBV strains differ in their ability to persist, there is also a substantial difference among intra-host HBV variants in their capacity to replicate on TDF. Thus, even small genetic changes generally observed among closely related intra-host HBV variants of a single strain seem to contribute to variation in sensitivity to TDF. Although the genetic mechanisms of on-TDF persistence can be disabled or enabled by few mutations in SR strains, these mechanisms cannot become fully functional in RR strains despite experiencing large numbers of mutations as in patients P6 and P7 infected with highly diverse HBV populations, which indicates a non-uniform distribution of the trait in the HBV genetic space.
A complex genetic encoding among SR strains coupled with small adaptive changes among intra-host HBV variants specific for each SR strain suggests the existence of many simple genetic mechanisms, various combinations of which set a specific path to persistence in each SR strain. In addition, it argues against the existence of a single or a dominant mechanism across all HBV strains, as generally observed for the actual drug resistance associated with simple and specific mutation patterns for all resistant strains. This observation suggests that the exact genetic mechanisms responsible for on-drug persistence may vary among HBV strains. The identification of different mutation spectra associated with persistence between HBV genotypes B and C, and lack of a single mutation pattern among persistent HBV strains studied here lends support to this supposition.
Owing to a high mutation rate and a large intra-host population size, it is estimated that HBV experiences all possible single and double mutations every day of infection in each infected individual13. However, despite the continuous occurrence of drug-resistance mutations, not all HBV strains develop resistance, indicating a fundamental role of HBV genetic background in phenotypic presentation of these mutations13. Epistatic connectivity among HBV sites is dense and can be organized into a network36. Genetic analyses show that this network defines HBV predisposition to drug resistance, making resistance mutations functionally acceptable in some HBV strains and, thus, selectable during treatment14. Like the resistance mutations, functional presentation of the TDF adaptive mutations in SR strains is epistatic or depends on the genetic background to which these mutations occur. Differences in genetic predisposition to persistence between SR and RR strains may explain adaptation of SR strains and lack of adaptation of RR strains to TDF treatment. Drug resistance and persistence are either convergent or independent of ancestry14, but persistence is highly genetically abundant or controlled by many genetic mechanisms, which alone are not as robust as resistance in controlling response to drugs and just protract HBV replication.
Cross-resistance of HBV strains to different nucleotide and nucleoside analogs is common1. Response to TDF, however, does not involve the development of actual resistance. Nevertheless, HBV infected patients preliminary treated with lamivudine or adefovir may have delayed or attenuated responses to TDF18,37, 38, suggesting a cross-selection for on-TDF persistence resulted from existence of genetic pathways for persistence shared by these three drugs. Genetic mechanisms of on-drug persistence may operate along with mechanisms of resistance. However, their effect is likely masked by the phenotypically dominant resistance. Identification of an HBV genotype A/G recombinant strain surviving during lamivudine treatment without development of the well-known lamivudine-resistance mutations14 suggests the existence of molecular mechanisms of on-lamivudine persistence, which are different from the actual lamivudine resistance.
The mechanisms of on-drug persistence are likely genotype specific. Indeed, a delayed response to TDF was observed for HBV genotype G38. Variation in susceptibility to TDF was reported for HBV genotype A vs. genotype C39,40. Here, mutation patterns associated with RR/SR were found to be different between HBV strains from genotypes B and C, additionally supporting the existence of genotype specific genetic pathways contributing to the TDF response. The nature of these mechanisms cannot be identified from the patterns alone. However, a strong association of the RNAse H sites with RR/SR in genotype C suggests a role of the enzymatic activity in on-TDF persistence. Although not yet observed for HBV, in HIV, mutations affecting the RNAse H conformation facilitate resistance to RT inhibitors likely by slowing degradation of the RNA genome during viral replication and, thus, providing more time for dissociation of the drug from the inhibited RT41. It is important to note that this mechanism is not specific to a certain drug. Many mechanisms associated with T- and B-cell responses42,43,44, as well as with functional states of the basal core promoter and pre-core regions of the HBV genome45 somewhat nonspecifically contribute to susceptibility to drugs and, thus, may serve to promote HBV persistence in absence of actual resistance.
In conclusion, capacity of HBV strains to persist on TDF is a complex trait genetically associated with mutations at many sites of the HBV genome. However, small genetic variations distinguishing persisting from non-persisting intra-host HBV variants indicate a potentially simple genetic nature of on-TDF persistence in each SR strain; while inconsistent presentation of these mutations among SR strains indicates a specific nature of these simple genetic mechanisms operating in each case. In contrast to drug resistance which is encoded by a dominant genetic mechanism across HBV strains, on-TDF persistence is likely controlled by many genetic mechanisms, each of which differentially operates in every persistent HBV strain. Although incapable to offset completely inhibition by drugs, on-drug persistence may contribute to or modify overall resistance. With drugs becoming ever more efficient, it is conceivable that complete resistance may become uncommon, and clinical management will rather face diminished responses to drugs, making mitigation of on-drug persistence essential for improving further quality of patients’ care by reducing duration of treatment as well as its cost. Understanding of genetic mechanisms of on-drug persistence should help in devising more potent drug therapies.
Methods
Patients
Whole-genome HBV quasispecies from ten immune tolerant patients (identified from Study GS-US-203-010146) were used for this study. All patients provided written informed consent. All methods were carried out in accordance with relevant guidelines and regulations. The study was approved by the Institutional Review Boards of each participating institution (Centers for Disease Control and Prevention’s Institutional Review Boards). All patients had received TDF monotherapy, and were matched by HBV titer, ALT and HBeAg at baseline. Patients were evaluated at base line, week 4 and week 40. Five patients had a slow response (SR), never achieving HBV DNA < 400 copies/ml by week 96, while five had a rapid response (RR), achieving HBV DNA < 400 copies/ml by week 96. Demographics and clinical features of patients are presented in the SI, Table S5.
Nucleic acid extraction and HBV whole genome quasispecies sequencing
Total nucleic acid was isolated from serum samples using the robotic Roche MagNA Pure LC system (software version 3.0.11) and the MagNA Pure LC Total Nucleic acid isolation kit (Roche Diagnostics GmbH, Mannheim, Germany), and eluted in 50 μl of lysis buffer according to the manufacturer’s instructions. Nearly full-length genomes of HBV genotype B (GT/B) and genotype C (GT/C) strains were amplified using two rounds of PCR as previously described47. Further details can be found in “SI Methods”.
Median joining network (MJN)
Intra-host heterogeneity of HBV strains at baseline, week 4 and week 40 was evaluated by MJN, constructed using the Phylogenetic Network software (NETWORK, version 4.112, Fluxus Technology Ltd, Suffolk, England48 (https://www.fluxus-engineering.com/sharenet.htm).
Bayesian networks (BN)
Genome-wide site-specific dependencies among nt/aa polymorphic sites in HBV genomes was assessed by BN modeling49. BN of polymorphic sites were estimated from alignments of HBV sequences using the SopLEQ method50,51 and a structural coefficient (SC) influence = 1 (significance threshold). The sequence alignment was done using the CLUSTALW program embedded in MEGA52 (version 6.06 https://www.megasoftware.net/). Prior to BN analysis, each sequence in the alignment was associated with respective metadata corresponding to response rate, genotype/subtype and time point of sampling. BN analyses were performed in three steps: learning step (SopLEQ method), analysis of associations and inference in order to characterize associations with response rates to TDF (target node). The Kullback–Leibler (KL) divergence53 was used to measure the strength of a direct relationship (link) between two variables (nodes), and the Bayes Factor (BF) metric29,30 to measure the impact of any given state of a node on the observed state of the target node. BN analyses were done with the BayesiaLab software (BayesiaLab, version 5.0.2 PE, Bayesia SAS, Laval, France, (https://www.bayesialab.com). Additional details are in “SI Methods”.
Feature selection (FS)
Statistical54 or machine-learning55 FS methods generally provide efficient means for identifying the most useful attributes for classification or regression tasks and are commonly employed to reduce the dimensionality of the data (i.e., number of attributes) without negatively affecting the accuracy of the prediction. FS was applied to HBV full or partial genome and protein sequence data, which comprised unique quasispecies variants within a sampled host. Samples were obtained from HBV-infected patients: six infected by HBV genotype C (HBV/C), three by genotype B (HBV/B) and one by genotype E (HBV/E). As in the case of BN analyses, FS analysis was performed on the genomic and/or protein sequence CSV formatted data, where each sequence variant was respectively annotated with corresponding RR or SR class-labels.
The correlation-based feature selection (CFS) algorithm was used for FS analysis of the sequence data, which is an FS technique based on the Merit heuristic56. Merit-based heuristics are founded on the idea that good feature subsets contain features highly correlated with the class label, yet poorly inter-correlated with each other. The basic strategy of this merit-based heuristics is to find the best minimal subset of features associated to a class label by accounting for the class-feature correlation and feature-feature correlations. In other words, CFS takes into account the usefulness of a feature subset for predicting the class label, while accounting for the level of inter-correlation between the features within the subset. Here, polymorphic sites comprised in a feature subset were evaluated by CFS to measure their joint (combined) correlation with respect to the class-label and inter-correlation among themselves. Merit measures returned by the CFS evaluation were then used to select the best feature subset for prediction of the HBV variants RR or SR association. We used the feature subset evaluation function implemented in WEKA (version 3.17, https://www.cs.waikato.ac.nz/ml/weka/)57, which is formalized as follows,
where, \(\overline{{r_{{cf}} }}\) is the average (avg.) class-feature correlation, and \(\overline{{r_{{ff}} }}\) is the avg. feature-feature intercorrelation in the feature subset S containing k features.
CFS identified among many feature subsets the most useful RR/SR predictive feature subset in HBV quasispecies genomes and proteomes. Polymorphic sites comprised in the best feature subset were then used to build the classification models presented herein. We note that other feature subsets may also have strong RR/SR predictive usefulness. However, due to statistical58 and computational limitations59,60 associated with FS methods, including the limitation in our patients cohort size (N = 10), it is not possible to determine the ground truth about which features are causal (or relevant) factors for the RR/SR characteristic. Nevertheless, we used the classification theory approach to examine the degree of reliability by which nucleotide or amino acid variations at the polymorphic sites identified by CFS help to associate HBV variants to the host’s RR/SR characteristics, and to establish the unlikelihood that such association could be attributed to genotype- and/or patient-related biases (refer to SI: Tables S1, and Table S2 & Fig. S5, respectively) or to random statistical correlations (details in “SI Methods”). Although the CFS-selected features identified herein as biomarkers for association of HBV strains to RR/SR are not definitive, and genomics experimentation is needed, the classification theory approach strongly supported the importance of CFS-selected features as biological factors that contributed to the specific and accurate identification of the HBV RR/SR predisposition to TDF treatment.
Classification models
Classifiers to identify/detect RR- and SR-strains were constructed using supervised and unsupervised machine learning61. Classification accuracy (CA) performances of supervised BN was evaluated using WEKA’s Experimenter module (version 3.6.157, https://www.cs.waikato.ac.nz/ml/weka/). An unsupervised self-organizing ANN was built using the self-organizing tree algorithm (SOTA)24,25 embedded in the KNIME software (KNIME Analytics Platform for Linux, version 2.8.2, Knime AG, Zurich, Switzerland, 201362 https://www.knime.com/). Further details can be found in “SI Methods”.
RT 3D-structure modeling
Structural modeling of five representative SR/RR-associated RT protein variants and MD simulation experiments was performed as previously described34. Ligand–protein interaction graphs were generated using the Maestro software (Schrödinger Release 2014-1: Maestro, version 9.7, Schrödinger, LLC, New York, NY, 2014). The 3-dimesional (3D) rendering of protein structures was done using VMD software35 (version 1.9.2; https://www.ks.uiuc.edu/Research/vmd/). Further details can be found in “SI Methods”.
Data availability
The sequence data that support the findings of this study were submitted to GenBank (accession numbers: MT426248–MT427201) and will be available to the public once they are processed. HBV data and datasets are available from the corresponding author upon reasonable request.
References
Guo, X. et al. Trends in hepatitis B virus resistance to nucleoside/nucleotide analogs in North China from 2009 to 2016: A retrospective study. Int. J. Antimicrob. Agents 52, 201–209 (2018).
Svicher, V. et al. Role of hepatitis B virus genetic barrier in drug-resistance and immune-escape development. Dig. Liver Dis. 43, 975–983 (2011).
Zoulim, F. Mechanism of viral persistence and resistance to nucleoside and nucleotide analogs in chronic hepatitis B virus infection. Antiviral Res. 64, 1–15 (2004).
Quan, D. J. & Peters, M. G. Antiviral therapy: Nucleotide and nucleoside analogs. Clin. Liver Dis. 8, 371–385 (2004).
Seigneres, B. et al. Inhibitory activity of dioxolane purine analogs on wild-type and lamivudine-resistant mutants of hepadnaviruses. Hepatology 36, 710–722 (2002).
Zoulim, F. et al. 2’,3’-Dideoxy-beta-l-5-fluorocytidine inhibits duck hepatitis B virus reverse transcription and suppresses viral DNA synthesis in hepatocytes, both in vitro and in vivo. Antimicrob. Agents Chemother. 40, 448–453 (1996).
Menéndez-Arias, L. Mechanisms of resistance to nucleoside analogue inhibitors of HIV-1 reverse transcriptase. Virus Res. 134, 124–146 (2008).
Selmi, B., Deval, J., Boretto, J. & Canard, B. Nucleotide analogue binding, catalysis and primer unblocking in the mechanisms of HIV-1 reverse transcriptase-mediated resistance to nucleoside analogues. Antivir. Ther. 8, 143–154 (2003).
Shaw, T., Bartholomeusz, A. & Locarnini, S. HBV drug resistance: Mechanisms, detection and interpretation. J. Hepatol. 44, 593–606 (2006).
Luber, A. D. Genetic barriers to resistance and impact on clinical response. MedGenMed 7, 69 (2005).
Kim, J. E. et al. Naturally occurring mutations in the reverse transcriptase region of hepatitis B virus polymerase from treatment-naive Korean patients infected with genotype C2. World J. Gastroenterol. 23, 4222–4232 (2017).
Whalley, S. A. et al. Kinetics of acute hepatitis B virus infection in humans. J. Exp. Med. 193, 847–854 (2001).
Khudyakov, Y. Coevolution and HBV drug resistance. Antivir. Ther. 15, 505–515 (2010).
Thai, H. et al. Convergence and coevolution of hepatitis B virus drug resistance. Nat. Commun. 3, 789 (2012).
Schweitzer, A., Horn, J., Mikolajczyk, R. T., Krause, G. & Ott, J. J. Estimations of worldwide prevalence of chronic hepatitis B virus infection: A systematic review of data published between 1965 and 2013. Lancet 386, 1546–1555 (2015).
Summers, J. & Mason, W. S. Replication of the genome of a hepatitis B-like virus by reverse transcription of an RNA intermediate. Cell 29, 403–415 (1982).
Heathcote, E. J. et al. Three-year efficacy and safety of tenofovir disoproxil fumarate treatment for chronic hepatitis B. Gastroenterology 140, 132–143 (2011).
Kitrinos, K. M. et al. No detectable resistance to tenofovir disoproxil fumarate after 6 years of therapy in patients with chronic hepatitis B. Hepatology 59, 434–442 (2014).
Marcellin, P. et al. Regression of cirrhosis during treatment with tenofovir disoproxil fumarate for chronic hepatitis B: A 5-year open-label follow-up study. Lancet 381, 468–475 (2013).
Snow-Lampart, A. et al. No resistance to tenofovir disoproxil fumarate detected after up to 144 weeks of therapy in patients monoinfected with chronic hepatitis B virus. Hepatology 53, 763–773 (2011).
Gordon, S. C. et al. Efficacy of tenofovir disoproxil fumarate at 240 weeks in patients with chronic hepatitis B with high baseline viral load. Hepatology 58, 505–513 (2013).
Lovett, G. C. et al. Efficacy and safety of tenofovir in chronic hepatitis B: Australian real world experience. World J. Hepatol. 9, 48–56 (2017).
Hall, M. A. Correlation-Based Feature Selection for Machine Learning Doctor of Philosophy Thesis (The University of Waikato, New Zeland, 1999).
Dopazo, J. & Carazo, J. M. Phylogenetic reconstruction using an unsupervised growing neural network that adopts the topology of a phylogenetic tree. J. Mol. Evol. 44, 226–233 (1997).
Herrero, J., Valencia, A. & Dopazo, J. A hierarchical unsupervised growing neural network for clustering gene expression patterns. Bioinformatics 17, 126–136 (2001).
Guckian, K. M. et al. Factors contributing to aromatic stacking in water: Evaluation in the context of DNA. J. Am. Chem. Soc. 122, 2213–2222 (2000).
Kidd-Ljunggren, K., Miyakawa, Y. & Kidd, A. H. Genetic variability in hepatitis B viruses. J. Gen. Virol. 83, 1267–1280 (2002).
Kramvis, A. Genotypes and genetic variability of hepatitis B virus. Intervirology 57, 141–150 (2014).
Jeffreys, H. Some tests of significance, treated by the theory of probability. Math. Proc. Camb. Philos. Soc. 31, 203–222 (1935).
Jeffreys, H. Theory of Probability 3rd edn. (Oxford University Press, Oxford, 1961).
Choi, Y. M., Lee, S. Y. & Kim, B. J. Naturally occurring hepatitis B virus reverse transcriptase mutations related to potential antiviral drug resistance and liver disease progression. World J. Gastroenterol. 24, 1708–1724 (2018).
Kay, A. & Zoulim, F. Hepatitis B virus genetic variability and evolution. Virus Res. 127, 164–176 (2007).
Wu, Q. et al. Evolution and mutations of hepatitis B virus quasispecies in genotype B and C during vertical transmission. J. Med. Virol. 88, 1018–1026 (2016).
Xu, X. et al. Modeling the functional state of the reverse transcriptase of hepatitis B virus and its application to probing drug-protein interaction. BMC Bioinform. 17(8), 280 (2016).
Humphrey, W., Dalke, A. & Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph 14(33–38), 27–38 (1996).
Campo, D. S. et al. Coordinated evolution of the hepatitis B virus polymerase. Silico Biol. 11, 175–182 (2011).
Engell, C. A., Pham, V. P., Holzman, R. S. & Aberg, J. A. Virologic outcome of using tenofovir/emtricitabine to treat hepatitis B in HIV-coinfected patients. ISRN Gastroenterol. 2011, 405390 (2011).
Lada, O. et al. Long-term outcome of primary non-responders to tenofovir therapy in HIV/HBV-co-infected patients: impact of HBV genotype G. Liver Int. 32, 93–101 (2012).
Bihl, F. et al. HBV genotypes and response to tenofovir disoproxil fumarate in HIV/HBV-coinfected persons. BMC Gastroenterol. 15, 79 (2015).
Murakami, E. et al. Effect of tenofovir disoproxil fumarate on drug-resistant HBV clones. J. Infect. 72, 91–102 (2016).
Wright, D. W. et al. A polymorphism at position 400 in the connection subdomain of HIV-1 reverse transcriptase affects sensitivity to NNRTIs and RNaseH activity. PLoS ONE 8, e74078 (2013).
Evans, A. et al. Programmed death 1 expression during antiviral treatment of chronic hepatitis B: Impact of hepatitis B e-antigen seroconversion. Hepatology 48, 759–769 (2008).
Fried, M. W. et al. HBeAg and hepatitis B virus DNA as outcome predictors during therapy with peginterferon alfa-2a for HBeAg-positive chronic hepatitis B. Hepatology 47, 428–434 (2008).
Liaw, Y. F. et al. 2-Year GLOBE trial results: Telbivudine is superior to lamivudine in patients with chronic hepatitis B. Gastroenterology 136, 486–495 (2009).
Cui, X. J., Cho, Y. K. & Song, B. C. Influence of the basal core promoter and precore mutation on replication of hepatitis B virus and antiviral susceptibility of different genotypes. J. Med. Virol. 87, 601–608 (2015).
Chan, H. L. et al. Effects of tenofovir disoproxil fumarate in hepatitis B e antigen-positive patients with normal levels of alanine aminotransferase and high levels of hepatitis B virus DNA. Gastroenterology 146, 1240–1248 (2014).
Ramachandran, S. et al. Evaluation of intra-host variants of the entire hepatitis B virus genome. PLoS ONE 6, e25232 (2011).
Bandelt, H. J., Forster, P. & Rohl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 16, 37–48 (1999).
Korb, K. & Nicholson, A. Bayesian Artificial Intelligence (Chapman & Hall/CRC Press, Cambridge, 2004).
Jouffe, L. Nouvelle classe de méthodes d'apprentissage de réseaux bayésiens. In Proceedings: Extraction et gestion des connaissances (EGC'2002), 345–356 (Montpellier, France, 2002).
Jouffe, L. & Munteanu, P. in Proceedings of the 10th International Symposium on Applied Stochastic Models and Data Analysis, 591–596 (Compiègne, France, 2001).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Kullback, S. & Leibler, R. A. On information and sufficiency. Ann. Math. Stat. 22, 79–86 (1951).
Webb, A. R. Statistical Pattern Recognition Ch. 9 307–318 (Wiley, New York, 2002).
Bishop, C. M. Pattern Recognition and Machine Learning 48–66 (Springer Science+Business Media LLC Publishers, New York, 2006).
Hall, M. A. in Proceedings of the Seventeenth International Conference on Machine Learning, 359–366 (Morgan Kaufmann, 2000).
Witten, I. H. & Frank, E. Data Mining: Practical Machine Learning Tools and Techniques 422–423 (Morgan Kaufmann, Amsterdam, 2005).
Sprites, P., Glymour, C. & Scheines, R. Causation, Prediction, and Search, Lecture Notes in Statistics Vol. 81, 87–102 (Springer, New York, 1993).
Molina, L. C. et al. in Proceedings of the 2002 IEEE International Conference on Data Mining, 306 (Maebashi City, Japan, 2002).
Peng, C., Xiao, S., Nie, Z., Wang, Z. & Wang, F. Applying Bayes’ theorem in medical expert systems. IEEE Eng. Med. Biol. 15, 76–79 (1996).
Libbrecht, M. W. & Noble, W. S. Machine learning applications in genetics and genomics. Nat. Rev. Genet. 16, 321–332 (2015).
Berthold, M. R. et al. KNIME: The Konstanz information miner. In Data Analysis, Machine Learning and Applications. Studies in Classification, Data Analysis, and Knowledge Organization (eds Preisach, C. et al.) (Springer, Berlin, 2008). https://doi.org/10.1007/978-3-540-78246-9_38.
Acknowledgements
We thank Mike Purdy (Centers for Disease Control and Prevention) for providing Perl-scripts for sequence data processing; Gilberto Vaughan, Suma Ramachandran and Lili Punkova (Centers for Disease Control and Prevention) for EPLD sequencing support. This study was supported by CDC intramural funding. Publication costs were funded by an internal program of CDC. At the time of this study, X.X.’s work was supported by the APHL-CDC Bioinformatics Fellowship Program.
Author information
Authors and Affiliations
Contributions
H.T. and J.L. performed research; K.K., A.G. and H.L.Y.C. worked on clinical data acquisition, analyzed clinical outcome data and provided patient annotation and serum specimens; H.T., G.X. and L.G.-R. performed DNA sequence analysis; J.L. performed machine-learning experiments; X.X. performed MD simulations and protein 3D-structure modeling; H.T., J.L. and Y.K. analyzed data and wrote manuscript.
Corresponding author
Ethics declarations
Competing interests
H.T., J.L., X.X., G.X., L.G.-R. and Y.K. (Centers for Disease Control and Prevention) declare no competing interests. CDC Disclaimer: the findings and conclusions of this manuscript are those of the authors and do not necessarily represent the official views of the Centers for Disease Control and Prevention. K.K. was an employee and stockholder at Gilead Sciences at the time the work was completed. A.G. is an employee and stockholder at Gilead Sciences. H.L.Y.C. is an advisor of Abbvie, Aligos, Arbutus, Gilead Sciences, Intellia, Janssen, MedImmune, Merck, Roche, Vir Biotechnology, Vaccitech, VenatoRx; and a speaker for Gilead Sciences, Mylan and Roche.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Thai, H., Lara, J., Xu, X. et al. Complex genetic encoding of the hepatitis B virus on-drug persistence. Sci Rep 10, 15574 (2020). https://doi.org/10.1038/s41598-020-72467-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-72467-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.