Abstract
Functional trait-based approaches have undergone an extraordinary expansion in phytoplankton ecology. Morpho-functional traits have been shown to vary both within and between populations and species, potentially affecting individual fitness and the network of inter-individual relationships. Here we integrate six fully harmonized phytoplankton morpho-functional trait datasets, characterized by a fine data grain, reporting individual-level data over a large biogeographical area. Datasets refer to transitional water ecosystems, from five biogeographical areas: Northern Atlantic Ocean (Scotland), South-Western Atlantic Ocean (Brazil), South-Western Pacific Ocean (Australia), Indo Pacific Ocean (Maldives) and Mediterranean Sea (Greece and Turkey). The integrated dataset includes 127311 individual phytoplankton records with sampling locations, taxonomic and morphometric information according to Darwin Core standards and semantic annotations. The six FAIR datasets are openly available in the LifeWatch Italy data portal. The datasets have already been used for morpho-functional analyses and hypothesis testing on phytoplankton guilds at different levels of data aggregation and scale, from local to global.
Similar content being viewed by others
Background & Summary
Trait-based approaches have become increasingly popular in community ecology1,2, including phytoplankton communities3,4,5,6, over the last few decades. Phytoplankton are a diverse group of microscopic organisms, accounting for approx. 40% of global primary productivity and are key contributors to the biogeochemical processes7,8. They provide an ideal model system for testing trait-based approaches, due to their relative simplicity and well-defined traits9,10. Phytoplankton morpho-functional traits affect the fitness and competitive success of individual cells, with cascading implications at the population, species and community levels11. Individual trait-based approaches provide the framework for linking individual responses to natural and anthropogenic pressures to community organization and ecosystem functioning12,13. Individual trait-based approaches have been applied in plant communities14,15 and more recently to plankton ones16,17,18. Here, we present an integrated individual-level trait-based phytoplankton dataset that combines six fully harmonized datasets related to transitional water ecosystems from the Northern Atlantic, South-Western Atlantic, South-Western Pacific, Indo Pacific Ocean and the Mediterranean Sea, sampled within the “Phytoplankton Bio-Imaging” project. All data are from a specific transitional water type, i.e., lagoon ecosystems, characterized by being micro-tidal, shallow and nutrient-rich depositional ecosystems19, determining morphometric traits adaptation of phytoplankton guilds when compared to the marine ones20. The lagoon ecosystems in each biogeographic area have been selected as relatively pristine ecosystems with low anthropogenic pressure; therefore, the integrated dataset as well as each individual biogeographic dataset can provide control/reference data for studying phytoplankton morphometric trait responses to anthropogenic pressure. The datasets have been harmonized and are openly accessible through the data portal of the Italian National distributed node (LifeWatch Italy) of the European e-Science Research Infrastructure LifeWatch ERIC. The integrated dataset presented here contributes to enhance the findable, accessible, and interoperable information21 on transitional water phytoplankton morpho-functional traits and complements the existing data resources on marine22,23 and freshwater24,25,26 phytoplankton.
Methods
Sampling and data collection
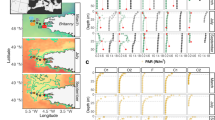
Phytoplankton samples were collected in a single sampling event that took place between July 2010 and November 2012 in 24 transitional water ecosystems distributed across five biogeographical regions: Northern Atlantic Ocean (NAO-Scotland), South-Western Atlantic Ocean (SWAO-Brazil), South-Western Pacific Ocean (SWPO-Australia), Indo Pacific Ocean (IPO-Maldives) and Mediterranean Sea (MED- Greece and Turkey) (Fig. 1). Sampling was carried out according to a hierarchical sampling design: for each ecoregion, three ecosystems were selected and within each of these, a maximum of three habitat categories were chosen and three experimental stations per habitat type were sampled with three replicates each, for a total count of 116 sites and approximately 350 water samples. Habitat types were classified on the basis of sediment granulometry and type of bottom vegetation27 according to the EUNIS habitat type hierarchical classification, version 201228.
Distribution map of the five biogeographical areas included in the dataset: South-Western Atlantic Ocean (SWAO) in brown, Northern Atlantic Ocean (NAO) in green, Mediterranean Sea (MED) in light blue, Indo Pacific Ocean (IPO) in purple and South-Western Pacific Ocean (SWPO) in pink. The red dots identify the phytoplankton sampling stations in each biogeographical area.
Phytoplankton samples were collected with horizontal tows from the subsurface (0.5 m) using a net mesh (6 µm) and fixed with Lugol’s solution (15 mL/L). This sampling technique is not a 100% quantitatively; however, the sampling procedures were standardised following same protocol in every sampling campaign. During the net sampling phase, the net was towed from the boat for a standard length of approximately 1.5–2 m, repeated three times back and forth, with each haul consisting of a linear measure of approximately 10 m. Phytoplankton taxonomic identification, cell abundances estimations and morphometric measurements were performed using an inverted microscope (Nikon T300E, Nikon Eclipse Ti) connected to a video-interactive image analysis system (L.U.C.I.A Version 4.8, Laboratory Imaging), following the Utermöhl method29 at 400x magnification. For each sample, a minimum of 400 cells were counted, measured and identified to the lowest taxonomic level possible, using specific manuals, monographs and phytoplankton Atlas30,31,32,33,34,35,36,37. The taxonomic validation was performed using the World Register of Marine Species (WoRMS)38 and Algae Base39.Where identification to species level was not possible the “Cf.” qualifier was used to indicate a specimen relevant to the species claimed and the numbered “sp.” was used to denote an organism relevant to the identified genus.
After taxonomic identification, cell volumes (expressed in μm3) were estimated according to the species/taxa specific shape association and using the geometric equations for simple and complex shapes recorded in the webservice “Atlas of Shapes” https://www.phytovre.lifewatchitaly.eu/vre/shapes-groups/. The geometric shape was attributed to the shape of the individual cell, even for coenobial, colonial and filamentous species where cells were not observable. The cell and shape views (e.g., lateral, frontal, etc.) with all the corresponding measured linear dimensions were reported in the datasets using alphabetical codes (e.g. length indicated by “a”, “l”, etc.; width indicated by alphabetical code “b”, “d”, etc.), together with information on the presence of internal and external cell structures (Table 1). Cell volumes were also reported in the datasets as “volume equivalent to sphere” and “volume equivalent to cylinder” and calculated using the Nikon image analysis system, based on cell contours and the application of a rod model using minimum and maximum Feret distances as linear dimensions40. Phytoplankton cellular carbon content (pg C) was obtained indirectly by converting cell biovolume to carbon using empirical or theoretically derived equations in accordance with Menden-Deuer and Lessard, 200041.
Data Records
The integrated dataset generated and analyzed for this study includes six datasets42,43,44,45,46,47 published in the LifeWatch Italy data portal https://dataportal.lifewatchitaly.eu/data with their respective DOIs (Table 2). The data were collected and harmonized according to the Phytoplankton Data Template https://www.phytovre.lifewatchitaly.eu/phyto-data-template/ which is based on the Darwin Core standards48 and the Phytoplankton Traits Thesaurus49. The datasets are formatted as column-oriented tables with data reported in semicolon_separated values format (.csv). The associated metadata are described using the Ecological Metadata Language50 (EML 2.2.0) standard in extensible markup language (.xml) format to ensure data understanding and long-term control. Each phytoplankton record is represented by an identifier (catalogNumber) associated with ancillary information (e.g. sampling locations, temporal and spatial information), phytoplankton taxonomic classification and morphological trait data (Table 1). Data variables are numeric and categorical and are expressed in text and numeric formats.
In total 127311 phytoplankton cells, belonging to 306 taxa were counted, measured and taxonomically classified. Summarized information from each dataset is presented in Table 3. The highest abundance was recorded in South-Western Atlantic Ocean area (SWAO), while the lowest number of records was reported in the South-Western Pacific Ocean (SWPO) region. In terms of taxa richness and shape occurrence a rather similar trend occurred in all biogeographical areas, with Northern Atlantic Ocean (NAO) biogeographical area showing the highest diversity in terms of taxa composition and the Indo Pacific Ocean area (IPO) showing the highest diversity in terms of shape occurrence. The distribution of phytoplankton composition by phyla in each biogeographical area showed a noticeable predominance of Ochrophyta in all regions (Fig. 2) mainly represented by the genera Chaetoceros, Pseudo-nitzschia, Ceratoneis, Cyclotella, Thalassionema and Navicula, followed by the Myzozoa represented by 66 different taxa. Other phyla such us Chlorophyta, Cyanobacteria, Cryptophyta, Haptophyta, Euglenozoa, Charophyta, Bacillariophyta and others accounted for less than 10000 individuals in each phyla, for a total of 87 taxa. A total count of 35 different morphological shapes of phytoplankton were described in the integrated dataset. Prism on elliptic base was the most abundant shape in terms of data records in all the biogeographical areas examined (38073 records) except for the South-Western Atlantic Ocean (SWAO) region where cylinder, parallelepiped and prolate spheroid + 2 cylinders were the most dominant shapes (Fig. 3). Complex shapes such as cylinder + 2 cones and cone + half sphere were mainly found in Northern Atlantic Ocean (NAO) and Mediterranean Sea (MED) regions with more than 2000 phytoplankton individuals, while more than 1500 organisms with cubic and gomphonemoid shapes were recorded in Northern Atlantic Ocean (NAO) area. The category “others” which included a mix of 20 different simple and complex shapes, rarely contributed to the overall morphological distribution of phytoplankton, with less than 1000 individuals.
Barplot showing the number of phytoplankton data records and the number of shapes per biogeographical area: South-Western Atlantic Ocean (SWAO) in brown, Northern Atlantic Ocean (NAO) in green, Mediterranean Sea (MED) in light blue, Indo Pacific Ocean (IPO) in purple and South-Western Pacific Ocean (SWPO) in pink. The shape category “Others” refers to less representative complex shapes described according to the Atlas of Shapes: half ellipsoid + cone on elliptic base, prism on triangular base 1, cone, prism on elliptic base + 4 cones, ellipsoid + cone, 2 half ellipsoids, prism on elliptic base + parallelepiped, 2 half ellipsoids + prism on elliptic base, ellipsoid + 2 cones + cylinder, cymbelloid, pyramid on rectangular base, cylinder + 3 cones, half sphere, parallelepiped + 2 cylinders, half ellipsoid + 3 cones, parallelepiped + 6 half cylinders, truncated cone, truncated cone + truncated cone, prism on elliptic base + 2 parallelepipeds and sickle-shaped prism.
Technical Validation
Data curation and technical validation steps were carried out to ensure the accuracy of the data and metadata. During data collection, a standardized sampling protocol already used in previous studies51 and the same sampling design were followed throughout the entire sampling campaign to avoid bias and ensure replicability of the data and information. Secondly, all the samples were collected, identified and measured by a team of qualified researchers and taxonomists who ensured data quality by checking and validating the taxonomic and morphological classification of the phytoplankton and detecting format and nomenclature errors or missing and inconsistent data. Thirdly, taxonomical and morphological information contained in all datasets were checked and technically validated through the use of WoRMS and Algaebase repository and the web services “Atlas of Shapes” and “Trait computation” provided in the Virtual Research Environment “Phyto VRE” of LifeWatch Italy (Fig. 4). After all the curation and validation steps, the data were stored and preserved in the LifeWatch Italy data portal, making them findable, accessible, interoperable and reusable. Finally, the integrated dataset includes information that have already been published in peer-reviewed scientific journals52,53,54,55,56,57,58.
Code availability
The provided datasets were established without the use of any custom code.
References
McGill, B. J., Enquist, B. J., Weiher, E. & Westoby, M. Rebuilding community ecology from functional traits. Trends Ecol. Evol. 21, 178–185 (2006).
Diaz, S. et al. The global spectrum of plant form and function: enhanced species-level trait dataset. Sci. Data 9, 755 (2022).
Edwards, K. F., Litchman, E. & Klausmeier, C. A. Functional traits explain phytoplankton responses to environmental gradients across lakes of the United States. Ecology 94, 1626–1635 (2013).
Stanca, E., Cellamare, M. & Basset, A. Geometric shape as a trait to study phytoplankton distributions in aquatic ecosystems. Hydrobiologia 701, 99–116 (2012).
Vadrucci, M. R., Cabrini, M. & Basset, A. Biovolume determination of phytoplankton guilds in transitional water ecosystems of Mediterranean Ecoregion. Transit. Water. Bull. 1, 83–102 (2007).
Kruk, C. et al. A morphological classification capturing functional variation in phytoplankton. Freshw. Biol. 55, 614–627 (2010).
Falkowski, P. G. The role of phytoplankton photosynthesis in global biogeochemical cycles. Photosynth. Res. 39, 235–258 (1994).
Litchman, E. et al. Global biogeochemical impacts of phytoplankton: a trait‐based perspective. J. Ecol. 103, 1384–1396 (2015).
Litchman, E. & Klausmeier, C. A. Trait-based community ecology of phytoplankton. Annu. Rev. of Ecol. Evol. Syst. 39, 615–639 (2008).
Violle, C. et al. Let the concept of trait be functional! Oikos 116, 882–892 (2007).
Fontana, S., Thomas, M. K., Moldoveanu, M., Spaak, P. & Pomati, F. Individual-level trait diversity predicts phytoplankton community properties better than species richness or evenness. ISME J. 12, 356–366 (2018).
Reiss, J., Bridle, J. R., Montoya, J. M. & Woodward, G. Emerging horizons in biodiversity and ecosystem functioning research. Trends Ecol. Evol. 24, 505–514 (2009).
Webb, C. T., Hoeting, J. A., Ames, G. M., Pyne, M. I. & LeRoy Poff, N. A structured and dynamic framework to advance traits‐based theory and prediction in ecology. Ecol. Lett. 13, 267–283 (2010).
Kattge, J. et al. TRY plant trait database - enhanced coverage and open access. Glob. Chang. Biol. 26, 119–188 (2020).
Siefert, A. & Ritchie, M. E. Intraspecific trait variation drives functional responses of old-field plant communities to nutrient enrichment. Oecologia 181, 245–255 (2016).
Salmaso, N., Naselli‐Flores, L. & Padisak, J. Functional classifications and their application in phytoplankton ecology. Freshw. Biol. 60, 603–619 (2015).
Weithoff, G. & Beisner, B. E. Measures and approaches in trait-based phytoplankton community ecology–from freshwater to marine ecosystems. Front. Mar. Sci. 6, 40 (2019).
Litchman, E., Edwards, K. F. & Boyd, P. W. Toward trait‐based food webs: Universal traits and trait matching in planktonic predator–prey and host–parasite relationships. Limnol. Oceanogr. 66, 3857–3872 (2021).
Basset, A. et al. A unifying approach to understanding transitional waters: fundamental properties emerging from ecotone ecosystems. Estuar. Coast. Shelf Sci. 132, 5–16 (2013).
Roselli, L. & Basset, A. Decoding size distribution patterns in marine and transitional water phytoplankton: from community to species level. PLoS ONE 10, e0127193 (2015).
Wilkinson, M. D. et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci. Data 3, 1–9 (2016).
Lundsor, E. et al. Marine phytoplankton community data and corresponding environmental properties from eastern Norway, 1896-2020, Sci. Data 9, 767 (2022).
Davies, C. H. et al. A database of marine phytoplankton abundance, biomass and species composition in Australian waters. Sci. Data 3, 160043 (2016).
Kremer, C. T., Gillette, J. P., Rudstam, L. G., Brettum, P. & Ptacnik, R. A compendium of cell and natural unit biovolumes for> 1200 freshwater phytoplankton species: Ecological Archives E095‐257. Ecology 95, 2984–2984 (2014).
Laplace-Treyture, C., Derot, J., Prévost, E., Le Mat, A. & Jamoneau, A. Phytoplankton morpho-functional trait dataset from French water-bodies. Sci. Data 8, 40 (2021).
Rimet, F. & Druart, J.-C. A trait database for Phytoplankton of temperate lakes. Ann. Limnol. – Int. J. Lim. 54, (2018).
Roff, J. C. & Taylor, M. E. National frameworks for marine conservation???a hierarchical geophysical approach. Aquat. Conserv.: Mar. Freshw. Ecos. 10, 209–223 (2000).
Evans, D. The EUNIS habitats classification–past, present & future. Rev. Invest. Mar. 19, 28–29 (2012).
Utermöhl, H. Zur vervollkommnung der quantitativen phytoplankton-methodik: Mit 1 Tabelle und 15 abbildungen im Text und auf 1 Tafel. Verh. - Int. Ver. Theor. Angew. Limnol.: Mitte. 9, 1–38 (1958).
Van Heurck, H. Synopsis Des Diatomées De Belgique (Édité Par l’auteu, 1884).
Rampi, L. & Bernhard, M. Key for the determination of Mediterranean pelagic bacillariophyceae. RT/BIO (78-1) Roma 71, (1978).
Sournia, A. Introduction, Cyanophycées, Dictyochophycées, Dinophycées et Raphidophycées. (1986).
Chrétiennot-Dinet, M. J. 1990. Chlorarachniophycées, chlorophycées, chrysophycées, cryptophycées, euglénophycées, eustigmatophycées, prasinophycées, prymnésiophycées, rhodophycées et tribophycées. (1990).
Round, F. E., Crawford, R. M. & Mann, D. G. Diatoms: Biology And Morphology Of The Genera. (Cambridge University Press, 1990).
Tomas, C. R. Identifying Marine Phytoplankton. (Elsevier, 1997).
Al-Kandari, M., Al-Yamani, F. & Al-Rifaie, K. Marine phytoplankton atlas of Kuwait’s waters. Kuwait J. Sci. 351, (2009).
Haraguchi, L. & Odebrecht, C. Dinophysiales (Dinophyceae) in the farthest Southern region of Brazil (Winter 2005, Summer 2007). Biota Neotropica 10, 101–114 (2010).
WoRMS Editorial Board. World Register of Marine Species https://doi.org/10.14284/170 (2013).
Guiry, M. D. & Guiry, G. M. AlgaeBase https://www.algaebase.org (2013).
Roselli, L., Stanca, E., Paparella, F., Mastrolia, A. & Basset, A. Determination of Coscinodiscus cf. granii biovolume by confocal microscopy: comparison of calculation models. J. Plankton Res. 35, 135–145 (2013).
Menden-Deuer, S. & Lessard, E. J. Carbon to volume relationships for dinoflagellates, diatoms, and other protist plankton. Limnol. Oceanogr. 45, 569–579 (2000).
LifeWatch ERIC https://doi.org/10.48372/dc6c5838-0e81-4aac-9442-fe9cb0bdb604 (2017).
LifeWatch ERIC https://doi.org/10.48372/5901dc22-9943-4fe0-9c23-1aeba0d52293 (2017).
LifeWatch ERIC https://doi.org/10.48372/098f6be3-8d79-4797-b0d6-5b22cdec9829 (2017).
LifeWatch ERIC https://doi.org/10.48372/4cf276c3-ba35-44f5-8ef0-a79de3e3bc06 (2017).
LifeWatch ERIC https://doi.org/10.48372/e7e415b4-4d4f-4180-8880-0f9446970f39 (2017).
LifeWatch ERIC https://doi.org/10.48372/4ea04557-8431-4b2e-8dff-c15a11fa937a (2017).
Darwin Core Maintenance Group. Darwin Core Quick Reference Guide, Biodiversity Information Standards https://dwc.tdwg.org/terms/ (2013).
Rosati, I. et al. A thesaurus for phytoplankton trait-based approaches: development and applicability. Ecol. Inform. 42, 129–138 (2017).
Fegraus, E. H., Andelman, S., Jones, M. B. & Schildhauer, M. Maximizing the value of ecological data with structured metadata: an introduction to ecological metadata language (EML) and principles for metadata creation. Bull. Ecol. Soc. Am. 86, 158–168 (2005).
Lugoli, E., Fiocca, A. & Stanca, E. Taxonomic composition of the phytoplankton community of Lesina lagoon (Apulia-Italy). Transit. Water. Bull. 7, 124–132 (2013).
Leonilde, R., Elena, L., Stanca, E., Francesco, C. & Basset, A. Individual trait variation in phytoplankton communities across multiple spatial scales. J. Plankton Res. 39, 577–588 (2017).
Roselli, L. et al. Multi-scale biodiverity patterns in phytoplankton from coastal lagoons: the Eastern Mediterranean. Transit. Water. Bull. 7, 202–219 (2013).
Roselli, L., Bevilacqua, S. & Terlizzi, A. Using null models and species traits to optimize phytoplankton monitoring: An application across oceans and ecosystems. Ecol. Indic. 138 (2022).
Souza, J. S. D. et al. A checklist of phytoplankton species around the equator in Guarairas, Galinhos and Diogo Lopes lagoons (Rio grande do norte, Brazil). Transit. Water. Bull. 7, 220–232 (2013).
Stanca, E. et al. A checklist of phytoplankton species in Faafu atoll (Republic of Maldives). Transit. Water. Bull. 7, 133–144 (2013).
Durante, G., Stanca, E., Roselli, L. & Basset, A. Phytoplankton composition in six Northern Scotland lagoons (Orkney Islands). Transit. Water. Bull. 7, 159–174 (2013).
Ryabov, A. et al. Shape matters: the relationship between cell geometry and diversity in phytoplankton. Ecol. Lett. 24, 847–861 (2021).
Acknowledgements
This study was supported by “POR PUGLIA Progetto Strategico 2009–2012”, “LifeWatchPLUS” CIR01_00028 and the project ITINERIS – Italian Integrated Environmental Research Infrastructure System CUP B53C22002150006 (PNRR for Mission4, Component 2, Notice 3264/2021, IR0000032). We would like to acknowledge all the colleagues, students and researchers who participate to the sampling phase and data collection, harmonization and compilation. We appreciate specifically the work of Stanca E. for phytoplankton identification and measurements and Vaira L. for her assistance with the data and metadata harmonization and publication.
Author information
Authors and Affiliations
Contributions
Basset A. supervised the sampling campaign and sample collection and processing. Data curation was performed by Rosati I. and data harmonization and analyses were performed by Laraib M. The first draft, editing and final review of the manuscript were conducted by Laraib M., Titocci J. and Basset A. Laraib M., Titocci J. have equally contributed to the development of the study and to all steps of development of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Laraib, M., Titocci, J., Rosati, I. et al. An integrated individual-level trait-based phytoplankton dataset from transitional waters. Sci Data 10, 897 (2023). https://doi.org/10.1038/s41597-023-02785-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02785-w