Abstract
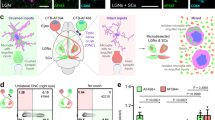
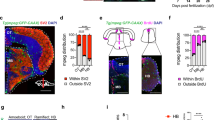
Oligodendrocyte precursor cells (OPCs) sculpt neural circuits through the phagocytic engulfment of synapses during development and adulthood. However, existing techniques for analyzing synapse engulfment by OPCs have limited accuracy. Here we describe the quantification of synapse engulfment by OPCs via a two-pronged cell biological approach that combines high-confidence and high-throughput methodologies. Firstly, an adeno-associated virus encoding a pH-sensitive, fluorescently tagged synaptic marker is expressed in neurons in vivo to differentially label presynaptic inputs, depending upon whether they are outside of or within acidic phagolysosomal compartments. When paired with immunostaining for OPC markers in lightly fixed tissue, this approach quantifies the engulfment of synapses by around 30–50 OPCs in each experiment. The second method uses OPCs isolated from dissociated brain tissue that are then fixed, incubated with fluorescent antibodies against presynaptic proteins, and analyzed by flow cytometry, enabling the quantification of presynaptic material within tens of thousands of OPCs in <1 week. The integration of both methods extends the current imaging-based assays, originally designed to quantify synaptic phagocytosis by other brain cells such as microglia and astrocytes, by enabling the quantification of synaptic engulfment by OPCs at individual and populational levels. With minor modifications, these approaches can be adapted to study synaptic phagocytosis by numerous glial cell types in the brain. The protocol is suitable for users with expertise in both confocal microscopy and flow cytometry. The imaging-based and flow cytometry-based protocols require 5 weeks and 2 d to complete, respectively.
Key points
-
An adeno-associated virus encoding a pH-sensitive, fluorescently tagged synaptic marker is expressed in neurons in vivo and combined with immunostaining for oligodendrocyte precursor cells (OPCs) in fixed tissue to quantify synaptic engulfment within individual OPCs. The measurements can be integrated with data from the flow cytometry-based quantification of synapses within OPCs pooled from across the mouse brain.
-
This protocol extends the limited capabilities in throughput and accuracy of existing imaging-based approaches.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Raw microscopy images (.czi files), processed microscopy images (.ims files), and flow cytometry data (.fcs files) have all been deposited and are available in the following Zenodo repository (https://doi.org/10.5281/zenodo.10625432).
Code availability
All R code used for analysis of flow cytometry data can be accessed from our Zenodo repository (https://doi.org/10.5281/zenodo.10625432).
References
Hooks, B. M. & Chen, C. Circuitry underlying experience-dependent plasticity in the mouse visual system. Neuron 106, 21–36 (2020).
Katz, L. C. & Shatz, C. J. Synaptic activity and the construction of cortical circuits. Science 274, 1133–1138 (1996).
Badimon, A. et al. Negative feedback control of neuronal activity by microglia. Nature 586, 417–423 (2020).
Lee, J. H. et al. Astrocytes phagocytose adult hippocampal synapses for circuit homeostasis. Nature 590, 612–617 (2021).
Wang, C. et al. Microglia mediate forgetting via complement-dependent synaptic elimination. Science 367, 688–694 (2020).
Schafer, D. P. et al. Microglia sculpt postnatal neural circuits in an activity and complement-dependent manner. Neuron 74, 691–705 (2012).
Paolicelli, R. C. et al. Synaptic pruning by microglia is necessary for normal brain development. Science 333, 1456–1458 (2011).
Gunner, G. et al. Sensory lesioning induces microglial synapse elimination via ADAM10 and fractalkine signaling. Nat. Neurosci. 22, 1075–1088 (2019).
Auguste, Y. S. S. et al. Oligodendrocyte precursor cells engulf synapses during circuit remodeling in mice. Nat. Neurosci. 25, 1273–1278 (2022).
Buchanan, J. et al. Oligodendrocyte precursor cells ingest axons in the mouse neocortex. Proc. Natl Acad. Sci. USA 119, e2202580119 (2022).
Xiao, Y., Petrucco, L., Hoodless, L. J., Portugues, R. & Czopka, T. Oligodendrocyte precursor cells sculpt the visual system by regulating axonal remodeling. Nat. Neurosci. 25, 280–284 (2022).
Buchanan, J., da Costa, N. M. & Cheadle, L. Emerging roles of oligodendrocyte precursor cells in neural circuit development and remodeling. Trends Neurosci. 46, 628–639 (2023).
Bergles, D. E., Roberts, J. D., Somogyi, P. & Jahr, C. E. Glutamatergic synapses on oligodendrocyte precursor cells in the hippocampus. Nature 405, 187–191 (2000).
Lin, S. C. & Bergles, D. E. Synaptic signaling between GABAergic interneurons and oligodendrocyte precursor cells in the hippocampus. Nat. Neurosci. 7, 24–32 (2004).
Spitzer, S. O. et al. Oligodendrocyte progenitor cells become regionally diverse and heterogeneous with age. Neuron 101, 459–471 e455 (2019).
Marisca, R. et al. Functionally distinct subgroups of oligodendrocyte precursor cells integrate neural activity and execute myelin formation. Nat. Neurosci. 23, 363–374 (2020).
Huang, W. et al. Origins and proliferative states of human oligodendrocyte precursor Cells. Cell 182, 594–608 e511 (2020).
Beiter, R. M. et al. Evidence for oligodendrocyte progenitor cell heterogeneity in the adult mouse brain. Sci. Rep. 12, 12921 (2022).
Brioschi, S. et al. Detection of synaptic proteins in microglia by flow cytometry. Front. Mol. Neurosci. 13, 149 (2020).
Dissing-Olesen, L. et al. FEAST: a flow cytometry-based toolkit for interrogating microglial engulfment of synaptic and myelin proteins. Nat. Commun. 14, 6015 (2023).
Kirby, L. et al. Oligodendrocyte precursor cells present antigen and are cytotoxic targets in inflammatory demyelination. Nat. Commun. 10, 3887 (2019).
Yuen, T. J. et al. Oligodendrocyte-encoded HIF function couples postnatal myelination and white matter angiogenesis. Cell 158, 383–396 (2014).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Hammill, D. CytoExploreR: interactive analysis of cytometric data. GitHub https://github.com/DillonHammill/CytoExploreR (2021).
Finak, G., Perez, J. M., Weng, A. & Gottardo, R. Optimizing transformations for automated, high throughput analysis of flow cytometry data. BMC Bioinforma. 11, 546 (2010).
Clayton, B. L. L. & Tesar, P. J. Oligodendrocyte progenitor cell fate and function in development and disease. Curr. Opin. Cell Biol. 73, 35–40 (2021).
Pfeiffer, F., Sherafat, A. & Nishiyama, A. The impact of fixation on the detection of oligodendrocyte precursor cell morphology and vascular associations. Cells 10, 1302 (2021).
Mori, T., Wakabayashi, T., Takamori, Y., Kitaya, K. & Yamada, H. Phenotype analysis and quantification of proliferating cells in the cortical gray matter of the adult rat. Acta Histochem. Cytochem. 42, 1–8 (2009).
Acknowledgements
We acknowledge and thank the following individuals for their contributions: U. Vrudhula for contributions to early imaging-based engulfment protocols; A.-S. Nichitiu for previously validating the pSynDig construct; P. Moody from the CSHL Flow Cytometry Core; E. Wee from the CSHL Microscopy Core and M.J. Gastinger from Imaris/Andor. This work was supported by the following funding sources (to L.C.): R00MH120051, DP2MH132943, R01NS131486, Rita Allen Scholar Award, McKnight Scholar Award, Klingenstein-Simons Fellowship Award in Neuroscience, Pershing Square Innovation Fund and a Brain and Behavior Foundation NARSAD grant. L.C. is a Howard Hughes Medical Institute Freeman Hrabowski Scholar.
Author information
Authors and Affiliations
Contributions
J.A.K., A.M.X. and L.C. wrote the paper. For the material, reagents and protocol sections, the components describing the microscopy-based approach were written by J.A.K. and the components related to flow cytometry were written by A.M.X. A.F. designed and produced the pSynDig construct. Y.S.S.A. designed the analysis of pSynDig data. A.F., J.A.K., S.T. and Y.A. contributed to optimizing the pSynDig OPC engulfment assay analysis. All figures were created by J.A.K., A.M.X. and L.C. Supervision and funding provided by L.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Tara DeSilva, Xiaoping Tong and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Auguste, Y. S. S. et al. Nat. Neurosci. 25, 1273–1278 (2022): https://doi.org/10.1038/s41593-022-01170-x
Supplementary information
Supplementary Information
Supplementary Figs. 1, 2
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kahng, J.A., Xavier, A.M., Ferro, A. et al. High-confidence and high-throughput quantification of synapse engulfment by oligodendrocyte precursor cells. Nat Protoc (2024). https://doi.org/10.1038/s41596-024-01048-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41596-024-01048-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.