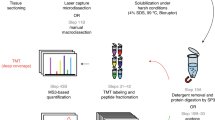
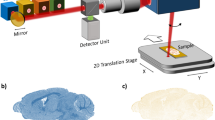
Abstract
The landscape of tissue-based imaging modalities is constantly and rapidly evolving. While formalin-fixed, paraffin-embedded material is still useful for histological imaging, the fixation process irreversibly changes the molecular composition of the sample. Therefore, many imaging approaches require fresh-frozen material to get meaningful results. This is particularly true for molecular imaging techniques such as mass spectrometry imaging, which are widely used to probe the spatial arrangement of the tissue metabolome. As high-quality fresh-frozen tissues are limited in their availability, any sample preparation workflow they are subjected to needs to ensure morphological and molecular preservation of the tissues and be compatible with as many of the established and emerging imaging techniques as possible to obtain the maximum possible insights from the tissues. Here we describe a universal sample preparation workflow, from the initial step of freezing the tissues to the cold embedding in a new hydroxypropyl methylcellulose/polyvinylpyrrolidone-enriched hydrogel and the generation of thin tissue sections for analysis. Moreover, we highlight the optimized storage conditions that limit molecular and morphological degradation of the sections. The protocol is compatible with human and plant tissues and can be easily adapted for the preparation of alternative sample formats (e.g., three-dimensional cell cultures). The integrated workflow is universally compatible with histological tissue analysis, mass spectrometry imaging and imaging mass cytometry, as well as spatial proteomic, genomic and transcriptomic tissue analysis. The protocol can be completed within 4 h and requires minimal prior experience in the preparation of tissue samples for multimodal imaging experiments.
Key points
-
This protocol presents the use of an hydroxypropyl methylcellulose/polyvinylpyrrolidone-rich hydrogel for the cold embedding and sectioning of animal and plant tissues as well as three-dimensional cell cultures.
-
The new embedding material achieves superior morphological preservation of the tissues and makes the specimens compatible with a wide variety of downstream applications, including mass spectrometry imaging, histology and molecular biology methods. The method facilitates the creation of multitissue blocks, limiting batch effects.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
Data availability
All relevant data are included in the main body of this paper or in the Extended Data. Source data are provided with this paper.
References
Doria, M. L. et al. Epithelial ovarian carcinoma diagnosis by desorption electrospray ionization mass spectrometry imaging. Sci. Rep. 6, 39219 (2016).
Groseclose, M. R., Laffan, S. B., Frazier, K. S., Hughes-Earle, A. & Castellino, S. Imaging MS in toxicology: an investigation of juvenile rat nephrotoxicity associated with dabrafenib administration. J. Am. Soc. Mass Spectrom. 26, 887–898 (2015).
Goodwin, R. J., Pennington, S. R. & Pitt, A. R. Protein and peptides in pictures: imaging with MALDI mass spectrometry. Proteomics 8, 3785–3800 (2008).
McDonnell, L. A. et al. Peptide and protein imaging mass spectrometry in cancer research. J. Proteom. 73, 1921–1944 (2010).
Schwartz, S. A., Reyzer, M. L. & Caprioli, R. M. Direct tissue analysis using matrix-assisted laser desorption/ionization mass spectrometry: practical aspects of sample preparation. J. Mass Spectrom. 38, 699–708 (2003).
Peukert, M. et al. Spatially resolved analysis of small molecules by matrix-assisted laser desorption/ionization mass spectrometric imaging (MALDI–MSI). N. Phytol. 193, 806–815 (2012).
Dannhorn, A. et al. Universal sample preparation unlocking multimodal molecular tissue imaging. Anal. Chem. 92, 11080–11088 (2020).
Bonnel, D. et al. Multivariate analyses for biomarkers hunting and validation through on-tissue bottom-up or in-source decay in MALDI–MSI: application to prostate cancer. Anal. Bioanal. Chem. 401, 149–165 (2011).
Stoeckli, M., Staab, D., Schweitzer, A., Gardiner, J. & Seebach, D. Imaging of a beta-peptide distribution in whole-body mice sections by MALDI mass spectrometry. J. Am. Soc. Mass Spectrom. 18, 1921–1924 (2007).
Everest-Dass, A. V. et al. N-glycan MALDI imaging mass spectrometry on formalin-fixed paraffin-embedded tissue enables the delineation of ovarian cancer tissues. Mol. Cell. Proteom. 15, 3003–3016 (2016).
Dannhorn, A. et al. Evaluation of formalin-fixed and FFPE tissues for spatially resolved metabolomics and drug distribution studies. Pharmaceuticals 15, 1307 (2022).
Berghmans, E. et al. Implementation of MALDI mass spectrometry imaging in cancer proteomics research: applications and challenges. J. Pers. Med. 10, 54 (2020).
Grgic, A., Krestensen, K. K. & Heeren, R. M. A. Optimized protocol for MALDI MSI of N-glycans using an on-tissue digestion in fresh frozen tissue sections. Sci. Rep. 13, 2776 (2023).
Shariatgorji, M. et al. Pyrylium salts as reactive matrices for MALDI–MS imaging of biologically active primary amines. J. Am. Soc. Mass Spectrom. 26, 934–939 (2015).
Shariatgorji, R. et al. Spatial visualization of comprehensive brain neurotransmitter systems and neuroactive substances by selective in situ chemical derivatization mass spectrometry imaging. Nat. Protoc. 16, 3298–3321 (2021).
Bednařík, A., Bölsker, S., Soltwisch, J. & Dreisewerd, K. An on-tissue Paternò–Büchi reaction for localization of carbon–carbon double bonds in phospholipids and glycolipids by matrix-assisted laser-desorption–ionization mass-spectrometry imaging. Angew. Chem. Int. Ed. 57, 12092–12096 (2018).
Flint, L. E. et al. Characterization of an aggregated three-dimensional cell culture model by multimodal mass spectrometry imaging. Anal. Chem. 92, 12538–12547 (2020).
Flint, L. E. et al. Comparison of osteosarcoma aggregated tumour models with human tissue by multimodal mass spectrometry imaging. Metabolites 11, 506 (2021).
Najumudeen, A. K. et al. The amino acid transporter SLC7A5 is required for efficient growth of KRAS-mutant colorectal cancer. Nat. Genet. 53, 16–26 (2021).
Adua, S. J. et al. Brain metastatic outgrowth and osimertinib resistance are potentiated by RhoA in EGFR-mutant lung cancer. Nat. Commun. 13, 7690 (2022).
Prag, H. A. et al. Ischemia-selective cardioprotection by malonate for ischemia/reperfusion injury. Circ. Res. 131, 528–541 (2022).
Vande Voorde, J. et al. Metabolic profiling stratifies colorectal cancer and reveals adenosylhomocysteinase as a therapeutic target. Nat. Metab. 5, 1303–1318 (2023).
Graziano, V. et al. Defining the spatial distribution of extracellular adenosine revealed a myeloid-dependent immunosuppressive microenvironment in pancreatic ductal adenocarcinoma. J. Immunother. Cancer 11, e006457 (2023).
Cuddihy, J. et al. Lactate dehydrogenase activity staining demonstrates time-dependent immune cell infiltration in human ex-vivo burn-injured skin. Sci. Rep. 11, 21249 (2021).
Dannhorn, A. et al. Evaluation of UV-C decontamination of clinical tissue sections for spatially resolved analysis by mass spectrometry imaging (MSI). Anal. Chem. 93, 2767–2775 (2021).
Isberg, O. G. et al. Automated cancer diagnostics via analysis of optical and chemical images by deep and shallow learning. Metabolites 12, 455 (2022).
Dannhorn, A. et al. Correlating mass spectrometry imaging and liquid chromatography–tandem mass spectrometry for tissue-based pharmacokinetic studies. Metabolites 12, 261 (2022).
Hamm, G. et al. Pharmacological inhibition of MERTK induces in vivo retinal degeneration: a multimodal imaging ocular safety assessment. Arch. Toxicol. 96, 613–624 (2022).
Mottahedin, A. et al. Targeting succinate metabolism to decrease brain injury upon mechanical thrombectomy treatment of ischemic stroke. Redox Biol. 59, 102600 (2023).
Race, A. M. et al. Deep learning-based annotation transfer between molecular imaging modalities: An automated workflow for multimodal data integration. Anal. Chem. 93, 3061–3071 (2021).
Strittmatter, N. et al. Method to visualize the intratumor distribution and impact of gemcitabine in pancreatic ductal adenocarcinoma by multimodal imaging. Anal. Chem. 94, 1795–1803 (2022).
Kucheriavaia, D. et al. Toward revealing microcystin distribution in mouse liver tissue using MALDI–MS imaging. Toxins 13, 709 (2021).
Strittmatter, N. et al. Holistic characterization of a Salmonella typhimurium infection model using integrated molecular imaging. J. Am. Soc. Mass Spectrom. 32, 2791–2802 (2021).
Rittel, M. F. et al. Spatial omics imaging of fresh-frozen tissue and routine FFPE histopathology of a single cancer needle core biopsy: a freezing device and multimodal workflow. Cancers 15, 2676 (2023).
Strittmatter, N. et al. Method to investigate the distribution of water-soluble drug-delivery systems in fresh frozen tissues using imaging mass cytometry. Anal. Chem. 93, 3742–3749 (2021).
Boughton, B. A., Thinagaran, D., Sarabia, D., Bacic, A. & Roessner, U. Mass spectrometry imaging for plant biology: a review. Phytochem. Rev. 15, 445–488 (2016).
Zemaitis, K. J. et al. Expanded coverage of phytocompounds by mass spectrometry imaging using on-tissue chemical derivatization by 4-APEBA. Anal. Chem. 95, 12701–12709 (2023).
Lukowski, J. K. et al. An optimized approach and inflation media for obtaining complimentary mass spectrometry-based omics data from human lung tissue. Front. Mol. Biosci. https://doi.org/10.3389/fmolb.2022.1022775 (2022).
Yang, J. et al. Optimization of Zebrafish larvae sectioning for mass spectrometry imaging. Pharmaceuticals 15, 1230 (2022).
Zhang, Y.-X., Zhang, Y.-D. & Shi, Y.-P. A reliable and effective sample preparation protocol of MALDI–TOF–MSI for lipids imaging analysis in hard and dry cereals. Food Chem. 398, 133911 (2023).
Ma, L. et al. Sample preparation optimization of insects and zebrafish for whole-body mass spectrometry imaging. Anal. Bioanal. Chem. 414, 4777–4790 (2022).
Hou, Y. et al. Applications of spatially resolved omics in the field of endocrine tumors. Front. Endocrinol. https://doi.org/10.3389/fendo.2022.993081 (2023).
Granborg, J. R., Handler, A. M. & Janfelt, C. Mass spectrometry imaging in drug distribution and drug metabolism studies—principles, applications and perspectives. Trends Anal. Chem. 146, 116482 (2022).
Dilmetz, B. A. et al. Novel technical developments in mass spectrometry imaging in 2020: a mini review. Anal. Sci. Adv. 2, 225–237 (2021).
Iakab, S. A., Ràfols, P., Correig-Blanchar, X. & García-Altares, M. Perspective on multimodal imaging techniques coupling mass spectrometry and vibrational spectroscopy: picturing the best of both worlds. Anal. Chem. 93, 6301–6310 (2021).
Tuck, M. et al. Multimodal imaging based on vibrational spectroscopies and mass spectrometry imaging applied to biological tissue: a multiscale and multiomics review. Anal. Chem. 93, 445–477 (2021).
Patterson, N. H., Tuck, M., Van de Plas, R. & Caprioli, R. M. Advanced registration and analysis of MALDI imaging mass spectrometry measurements through autofluorescence microscopy. Anal. Chem. https://doi.org/10.1021/acs.analchem.8b02884 (2018).
Ushida, K., Asai, N., Uchiyama, K., Enomoto, A. & Takahashi, M. Development of a method to preliminarily embed tissue samples using low melting temperature fish gelatin before sectioning: a technical note. Pathol. Int. 68, 241–245 (2018).
Strohalm, M. et al. Poly[N -(2-hydroxypropyl)methacrylamide]-based tissue-embedding medium compatible with MALDI mass spectrometry imaging experiments. Anal. Chem. 83, 5458–5462 (2011).
Yang, C. et al. Polyacrylamide gel as a new embedding medium for the enhancement of metabolite MALDI imaging. Chem. Commun. 59, 3842–3845 (2023).
Fala, M. et al. Comparison of 13C MRI of hyperpolarized [1-13C]pyruvate and lactate with the corresponding mass spectrometry images in a murine lymphoma model. Magn. Reson. Med. 85, 3027–3035 (2021).
Goodwin, R. J. Sample preparation for mass spectrometry imaging: small mistakes can lead to big consequences. J. Proteom. 75, 4893–4911 (2012).
Steu, S. et al. A procedure for tissue freezing and processing applicable to both intra-operative frozen section diagnosis and tissue banking in surgical pathology. Virchows Arch. 452, 305–312 (2008).
Nilsson, A. et al. Investigating nephrotoxicity of polymyxin derivatives by mapping renal distribution using mass spectrometry imaging. Chem. Res. Toxicol. 28, 1823–1830 (2015).
Shabihkhani, M. et al. The procurement, storage, and quality assurance of frozen blood and tissue biospecimens in pathology, biorepository, and biobank settings. Clin. Biochem. 47, 258–266 (2014).
Choy Buentello, D., Koch, L. S., Trujillo-de Santiago, G., Alvarez, M. M. & Broersen, K. Use of standard U-bottom and V-bottom well plates to generate neuroepithelial embryoid bodies. PLoS ONE 17, e0262062 (2022).
Bakker, B. et al. Preparing ductal epithelial organoids for high-spatial-resolution molecular profiling using mass spectrometry imaging. Nat. Protoc. 17, 962–979 (2022).
Broutier, L. et al. Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation. Nat. Protoc. 11, 1724–1743 (2016).
Li, H. & Hummon, A. B. Imaging mass spectrometry of three-dimensional cell culture systems. Anal. Chem. 83, 8794–8801 (2011).
Liu, X. et al. MALDI–MSI of immunotherapy: mapping the EGFR-targeting antibody cetuximab in 3D colon-cancer cell cultures. Anal. Chem. 90, 14156–14164 (2018).
Xie, P., Zhang, H., Wu, P., Chen, Y. & Cai, Z. Three-dimensional mass spectrometry imaging reveals distributions of lipids and the drug metabolite associated with the enhanced growth of colon cancer cell spheroids treated with triclosan. Anal. Chem. 94, 13667–13675 (2022).
Swales, J. G. et al. Quantitation of endogenous metabolites in mouse tumors using mass-spectrometry imaging. Anal. Chem. 90, 6051–6058 (2018).
Maynard, K. R. et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat. Neurosci. 24, 425–436 (2021).
Lukowski, J. K. et al. Storage conditions of human kidney tissue sections affect spatial lipidomics analysis reproducibility. J. Am. Soc. Mass Spectrom. 31, 2538–2546 (2020).
Asslan, M., Lauzon, N., Beus, M., Maysinger, D. & Rousseau, S. Mass spectrometry imaging in zebrafish larvae for assessing drug safety and metabolism. Anal. Bioanal. Chem. 413, 5135–5146 (2021).
Iakab, S.-A. et al. 3D-mass spectrometry imaging of micro-scale 3D cell culture models in cancer research. Preprint at bioRxiv https://doi.org/10.1101/2022.12.05.519157 (2022).
Archer, C. R. et al. Characterization and validation of a human 3D cardiac microtissue for the assessment of changes in cardiac pathology. Sci. Rep. 8, 10160 (2018).
Acknowledgements
The authors thank the Biotechnology and Biological Sciences Research Council for the case funding for A.D. (BB/N504038/1). The authors also acknowledge the Cancer Research UK Grand Challenge Rosetta Consortium for discussions in support of their research.
Author information
Authors and Affiliations
Contributions
A.D. and E.K. developed the embedding medium and procedure for tissues. L.F., F.G. and A.R.H. adapted the methodology for the embedding of 3D cell cultures. A.C. and A.R.H. provided the organoids for the appropriate protocol steps. S.A.J. provided the expertise for the freezing procedures of fresh tissue samples. G.P., S.T.B., O.J.S., J.B., Z.T. and R.J.A.G. provided feedback discussion for the methodology. Z.T. and R.J.A.G. secured the funding for the work. All authors reviewed and edited the manuscript drafted by A.D.
Corresponding author
Ethics declarations
Competing interests
A.D., L.F., F.G., A.C., A.R.H., S.A.J., S.T.B. and R.J.A.G. are full-time salaried employees, own stocks of AstraZeneca and performed this study as part of their regular duties.
Peer review
Peer review information
Nature Protocols thanks Junhai Yang, Xiaodong Wang and Miriam Rittel for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Dannhorn, A. et al. Anal. Chem. 92, 11080–11088 (2020): https://doi.org/10.1021/acs.analchem.0c00826
Flint, L. E. et al. Anal. Chem. 92, 12538–12547 (2020): https://doi.org/10.1021/acs.analchem.0c02389
Prag, H. A. et al. Circ. Res. 131, 528–541 (2022): https://doi.org/10.1161/circresaha.121.320717
Vande Voorde, J. et al. Nat. Metab. 5, 1303–1318 (2023): https://doi.org/10.1038/s42255-023-00857-0
Najumudeen, A. K. et al. Nat. Genet. 53, 16–26 (2021): https://doi.org/10.1038/s41588-020-00753-3
Extended data
Extended Data Fig. 1 Impact of choice of cryogen on sample integrity.
a, mouse kidney snap frozen in dry ice chilled isopentane with fracture line through the organ. b, mouse kidney snap frozen in dry ice chilled isopropanol after washing in dry ice chilled isopentane without visible fractures.
Extended Data Fig. 2 Troubleshooting of the freezing of the multi-tissue blocks.
a, Multi-tissue block that was fully frozen before being washed in the isopentane dry ice slurry. b, Multi-tissue block that was not fully frozen before being washed in the isopentane dry ice slurry and that fractured due to pressure building up during rapid freezing. The fracture line is indicated with ▲.
Extended Data Fig. 3 Identification of the sectioning level for 3D cell culture blocks.
a, Top layer of the red-dyed embedding medium. b, Interface layer between the two dyed embedding media in which the 3D cell cultures are positioned and sections of the tissues should be collected. c, tissue-free, blue-dyed base medium after exhaustion of the sample.
Extended Data Fig. 4 Troubleshooting of the sectioning temperature.
a, Section of a multi-tissue block collected at the appropriate temperature without visible fractures of the tissues or sticking to the surfaces of the cryomicrotome. b, Section collected at too low temperature with visible fracturing of the tissues within. c, Section of a multi-tissue block collected at too high temperature with visible damage to the embedding medium and tissue sections.
Extended Data Fig. 5 Troubleshooting of a damaged blade or anti-rolling plate.
a,b, Section of a multi-tissue block with intact blade and the resulting H&E stained tissue section. c,d, Section of the same block collected with a faulty blade and visibly affected tissue morphology on a H&E stained tissue as indicated by ▲. The faulty blade scraped the tissue resulting in bands where the morphological structures were torn apart, rendering them impossible to identify. Scale bars in b and d, 200 µm.
Source data
Source Data Fig. 9
Unprocessed fluorescent microscopy data used to generate Fig. 9.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dannhorn, A., Kazanc, E., Flint, L. et al. Morphological and molecular preservation through universal preparation of fresh-frozen tissue samples for multimodal imaging workflows. Nat Protoc 19, 2685–2711 (2024). https://doi.org/10.1038/s41596-024-00987-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-024-00987-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.