Abstract
RNA structure determination is essential to understand how RNA carries out its diverse biological functions. In cells, RNA isoforms are readily expressed with partial variations within their sequences due, for example, to alternative splicing, heterogeneity in the transcription start site, RNA processing or differential termination/polyadenylation. Nanopore dimethyl sulfate mutational profiling (Nano-DMS-MaP) is a method for in situ isoform-specific RNA structure determination. Unlike similar methods that rely on short sequencing reads, Nano-DMS-MaP employs nanopore sequencing to resolve the structures of long and highly similar RNA molecules to reveal their previously hidden structural differences. This Protocol describes the development and applications of Nano-DMS-MaP and outlines the main considerations for designing and implementing a successful experiment: from bench to data analysis. In cell probing experiments can be carried out by an experienced molecular biologist in 3–4 d. Data analysis requires good knowledge of command line tools and Python scripts and requires a further 3–5 d.
Key points
-
Nano-DMS-MaP is a method for in situ isoform-specific RNA structure determination. It employs nanopore sequencing to resolve the structures of long and highly similar RNA molecules, revealing previously hidden structural differences.
-
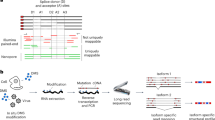
Compared with short-read sequencing, in which it is difficult to uniquely map individual reads to highly similar transcript isoforms, Nano-DMS-MaP uses long-read Nanopore sequencing, enabling unambiguous assignment of reads to transcript isoforms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The data used to generate the anticipated results were originally published in ref. 22. All sequencing data are available at Sequence Read Archive (SRP424422, Bioproject ID PRJNA938445).
Code availability
Code used for the Nano-DMS-MaP analysis is accessible via the Smyth lab Github (https://github.com/smyth-lab/Nano-DMS-MaP) and is available for reuse under the Massachusetts Institute of Technology (MIT) License.
References
Vicens, Q. & Kieft, J. S. Thoughts on how to think (and talk) about RNA structure. Proc. Natl Acad. Sci. USA 119, e2112677119 (2022).
Mortimer, S. A., Kidwell, M. A. & Doudna, J. A. Insights into RNA structure and function from genome-wide studies. Nat. Rev. Genet. 15, 469–479 (2014).
Spitale, R. C. & Incarnato, D. Probing the dynamic RNA structurome and its functions. Nat. Rev. Genet. 24, 178–196 (2023).
Reyes, A. & Huber, W. Alternative start and termination sites of transcription drive most transcript isoform differences across human tissues. Nucleic Acids Res. 46, 582–592 (2018).
Shabalina, S. A., Ogurtsov, A. Y., Spiridonov, N. A. & Koonin, E. V. Evolution at protein ends: major contribution of alternative transcription initiation and termination to the transcriptome and proteome diversity in mammals. Nucleic Acids Res. 42, 7132–7144 (2014).
Ehresmann, C. et al. Probing the structure of RNAs in solution. Nucleic Acids Res. 15, 9109–9128 (1987).
Mailler, E., Paillart, J.-C. J.-C., Marquet, R., Smyth, R. P. R. P. & Vivet-Boudou, V. The evolution of RNA structural probing methods: from gels to next-generation sequencing. Wiley Interdiscip. Rev. RNA 10, e1518 (2019).
Morgan, B. S., Forte, J. E. & Hargrove, A. E. Insights into the development of chemical probes for RNA. Nucleic Acids Res. 46, 8025–8037 (2018).
Spitale, R. C. et al. RNA SHAPE analysis in living cells. Nat. Chem. Biol. 9, 18–20 (2013).
Merino, E. J., Wilkinson, K. A., Coughlan, J. L. & Weeks, K. M. RNA structure analysis at single nucleotide resolution by selective 2′-hydroxyl acylation and primer extension (SHAPE). J. Am. Chem. Soc. 127, 4223–4231 (2005).
Wilkinson, K. A., Merino, E. J. & Weeks, K. M. RNA SHAPE chemistry reveals nonhierarchical interactions dominate equilibrium structural transitions in tRNA(Asp) transcripts. J. Am. Chem. Soc. 127, 4659–4667 (2005).
Lee, B. et al. Comparison of SHAPE reagents for mapping RNA structures inside living cells. RNA 23, 169–174 (2017).
Siegfried, N. A., Busan, S., Rice, G. M., Nelson, J. A. E. & Weeks, K. M. RNA motif discovery by SHAPE and mutational profiling (SHAPE-MaP). Nat. Methods 11, 959–965 (2014).
Smola, M. J., Calabrese, J. M. & Weeks, K. M. Detection of RNA–protein interactions in living cells with SHAPE. Biochemistry 54, 6867–6875 (2015).
McGinnis, J. L. et al. In-cell SHAPE reveals that free 30S ribosome subunits are in the inactive state. Proc. Natl Acad. Sci. USA 112, 2425–2430 (2015).
Martin, S., Blankenship, C., Rausch, J. W. & Sztuba-Solinska, J. Using SHAPE-MaP to probe small molecule-RNA interactions. Methods 167, 105–116 (2019).
Kubota, M., Tran, C. & Spitale, R. C. Progress and challenges for chemical probing of RNA structure inside living cells. Nat. Chem. Biol. 11, 933–941 (2015).
Lempereur, L. et al. Conformation of yeast 18S rRNA. Direct chemical probing of the 5′ domain in ribosomal subunits and in deproteinized RNA by reverse transcriptase mapping of dimethyl sulfate-accessible. Nucleic Acids Res. 13, 8339–8357 (1985).
Zubradt, M. et al. DMS-MaPseq for genome-wide or targeted RNA structure probing in vivo. Nat. Methods 14, 75–82 (2017).
Krokhotin, A., Mustoe, A. M., Weeks, K. M. & Dokholyan, N. V. Direct identification of base-paired RNA nucleotides by correlated chemical probing. RNA 23, 6–13 (2017).
Rouskin, S., Zubradt, M., Washietl, S., Kellis, M. & Weissman, J. S. Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo. Nature 505, 701–705 (2014).
Bohn, P., Gribling-Burrer, A.-S., Ambi, U. B. & Smyth, R. P. Nano-DMS-MaP allows isoform-specific RNA structure determination. Nat. Methods 20, 849–859 (2023).
Peattie, D. A. & Gilbert, W. Chemical probes for higher-order structure in RNA. Proc. Natl Acad. Sci. USA 77, 4679–4682 (1980).
Mitchell, D., Cotter, J., Saleem, I. & Mustoe, A. M. Mutation signature filtering enables high-fidelity RNA structure probing at all four nucleobases with DMS. Nucleic Acids Res. 51, 8744–8757 (2023).
Onel, B., Wu, G., Sun, D., Lin, C. & Yang, D. Electrophoretic mobility shift assay and dimethyl sulfate footprinting for characterization of G-quadruplexes and G-quadruplex-protein complexes. Methods Mol. Biol. 2035, 201–222 (2019).
England, W. E., Garfio, C. M. & Spitale, R. C. Chemical approaches to analyzing RNA structure transcriptome-wide. Chembiochem 22, 1114–1121 (2021).
Inoue, T. & Cech, T. R. Secondary structure of the circular form of the Tetrahymena rRNA intervening sequence: a technique for RNA structure analysis using chemical probes and reverse transcriptase. Proc. Natl Acad. Sci. USA 82, 648–652 (1985).
Sobczak, K. & Krzyzosiak, W. J. RNA structure analysis assisted by capillary electrophoresis. Nucleic Acids Res. 30, e124 (2002).
Li, B., Tambe, A., Aviran, S. & Pachter, L. PROBer provides a general toolkit for analyzing sequencing-based toeprinting assays. Cell Syst. 4, 568–574.e7 (2017).
Kertesz, M. et al. Genome-wide measurement of RNA secondary structure in yeast. Nature 467, 103–107 (2010).
Underwood, J. G. et al. FragSeq: transcriptome-wide RNA structure probing using high-throughput sequencing. Nat. Methods 7, 995–1001 (2010).
Lucks, J. B. et al. Multiplexed RNA structure characterization with selective 2′-hydroxyl acylation analyzed by primer extension sequencing (SHAPE-Seq). Proc. Natl Acad. Sci. USA 108, 11063–8 (2011).
Ding, Y. et al. In vivo genome-wide profiling of RNA secondary structure reveals novel regulatory features. Nature 505, 696–700 (2014).
Talkish, J. et al. Mod-seq: high-throughput sequencing for chemical probing of RNA structure. RNA 20, 713–720 (2014).
Incarnato, D., Neri, F., Anselmi, F. & Oliviero, S. Genome-wide profiling of mouse RNA secondary structures reveals key features of the mammalian transcriptome. Genome Biol. 15, 491 (2014).
Spitale, R. C. et al. Structural imprints in vivo decode RNA regulatory mechanisms. Nature 519, 486–490 (2015).
Busan, S. & Weeks, K. M. Accurate detection of chemical modifications in RNA by mutational profiling (MaP) with ShapeMapper 2. RNA 24, 143–148 (2018).
Loughrey, D., Watters, K. E., Settle, A. H. & Lucks, J. B. SHAPE-Seq 2.0: systematic optimization and extension of high-throughput chemical probing of RNA secondary structure with next generation sequencing. Nucleic Acids Res. 42, 1–10 (2014).
Sorefan, K. et al. Reducing ligation bias of small RNAs in libraries for next generation sequencing. Silence 3, 4 (2012).
Aviran, S. et al. Modeling and automation of sequencing-based characterization of RNA structure. Proc. Natl Acad. Sci. USA 108, 11069–11074 (2011).
Tomezsko, P. J. et al. Determination of RNA structural diversity and its role in HIV-1 RNA splicing. Nature 582, 438–442 (2020).
Olson, S. W. et al. Discovery of a large-scale, cell-state-responsive allosteric switch in the 7SK RNA using DANCE-MaP. Mol. Cell 82, 1708–1723.e10 (2022).
Morandi, E. et al. Genome-scale deconvolution of RNA structure ensembles. Nat. Methods 18, 249–252 (2021).
Homan, P. J. et al. Single-molecule correlated chemical probing of RNA. Proc. Natl Acad. Sci. USA 111, 13858–13863 (2014).
Yang, M. et al. In vivo single-molecule analysis reveals COOLAIR RNA structural diversity. Nature 609, 394–399 (2022).
Aw, J. G. A. et al. Determination of isoform-specific RNA structure with nanopore long reads. Nat. Biotechnol. 39, 336–346 (2021).
Bizuayehu, T. T. et al. Long-read single-molecule RNA structure sequencing using nanopore. Nucleic Acids Res. https://doi.org/10.1093/nar/gkac775 (2022).
Stephenson, W. et al. Direct detection of RNA modifications and structure using single-molecule nanopore sequencing. Cell Genomics 2, 100097 (2022).
Mitchell, D., Assmann, S. M. & Bevilacqua, P. C. Probing RNA structure in vivo. Curr. Opin. Struct. Biol. 59, 151–158 (2019).
Smola, M. J., Rice, G. M., Busan, S., Siegfried, N. A. & Weeks, K. M. Selective 2′-hydroxyl acylation analyzed by primer extension and mutational profiling (SHAPE-MaP) for direct, versatile and accurate RNA structure analysis. Nat. Protoc. 10, 1643–1669 (2015).
Tijerina, P., Mohr, S. & Russell, R. DMS footprinting of structured RNAs and RNA-protein complexes. Nat. Protoc. 2, 2608–2623 (2007).
Assmann, S. M., Chou, H.-L. & Bevilacqua, P. C. Rock, scissors, paper: how RNA structure informs function. Plant Cell 35, 1671–1707 (2023).
Tian, S. & Das, R. RNA structure through multidimensional chemical mapping. Q. Rev. Biophys. 49, e7 (2016).
Gilmer, O. et al. Chemical and enzymatic probing of viral RNAs: from infancy to maturity and beyond. Viruses 13, 1894 (2021).
Ye, L. et al. Short- and long-range interactions in the HIV-1 5′ UTR regulate genome dimerization and packaging. Nat. Struct. Mol. Biol. 29, 306–319 (2022).
Lan, T. C. T. et al. Secondary structural ensembles of the SARS-CoV-2 RNA genome in infected cells. Nat. Commun. 13, 1–14 (2022).
Hu, Y. et al. DiffSplice: the genome-wide detection of differential splicing events with RNA-seq. Nucleic Acids Res. 41, e39 (2013).
Engström, P. G. et al. Systematic evaluation of spliced alignment programs for RNA-seq data. Nat. Methods 10, 1185–1191 (2013).
Prjibelski, A. D. et al. Accurate isoform discovery with IsoQuant using long reads. Nat. Biotechnol. 41, 915–918 (2023).
Zhang, Y., Lu, L. & Li, X. Detection technologies for RNA modifications. Exp. Mol. Med. 54, 1601–1616 (2022).
Zee, A. et al. Sequencing Illumina libraries at high accuracy on the ONT MinION using R2C2. Genome Res. 32, 2092–2106 (2022).
Zhao, C., Liu, F. & Pyle, A. M. An ultraprocessive, accurate reverse transcriptase encoded by a metazoan group II intron. RNA 24, 183–195 (2018).
Guo, L.-T. et al. Sequencing and structure probing of long RNAs using MarathonRT: a next-generation reverse transcriptase. J. Mol. Biol. 432, 3338–3352 (2020).
Zhao, C. & Pyle, A. M. Crystal structures of a group II intron maturase reveal a missing link in spliceosome evolution. Nat. Struct. Mol. Biol. 23, 558–565 (2016).
Guo, L.-T., Olson, S., Patel, S., Graveley, B. R. & Pyle, A. M. Direct tracking of reverse-transcriptase speed and template sensitivity: implications for sequencing and analysis of long RNA molecules. Nucleic Acids Res. 50, 6980–6989 (2022).
Mohr, S. et al. Thermostable group II intron reverse transcriptase fusion proteins and their use in cDNA synthesis and next-generation RNA sequencing. RNA 19, 958–970 (2013).
Ip, C. L. C. et al. MinION Analysis and Reference Consortium: phase 1 data release and analysis. F1000Res. 4, 1075 (2015).
Luo, J. et al. Systematic benchmarking of nanopore Q20+ kit in SARS-CoV-2 whole-genome sequencing. Front. Microbiol. 13, 973367 (2022).
Ni, Y., Liu, X., Simeneh, Z. M., Yang, M. & Li, R. Benchmarking of Nanopore R10.4 and R9.4.1 flow cells in single-cell whole-genome amplification and whole-genome shotgun sequencing. Comput. Struct. Biotechnol. J. 21, 2352–2364 (2023).
Kharytonchyk, S. et al. Transcriptional start site heterogeneity modulates the structure and function of the HIV-1 genome. Proc. Natl Acad. Sci. USA 113, 13378–13383 (2016).
Purcell, D. F. & Martin, M. A. Alternative splicing of human immunodeficiency virus type 1 mRNA modulates viral protein expression, replication, and infectivity. J. Virol. 67, 6365–6378 (1993).
Nguyen Quang, N. et al. Dynamic nanopore long-read sequencing analysis of HIV-1 splicing events during the early steps of infection. Retrovirology 17, 1–24 (2020).
Bernacchi, S. et al. HIV-1 Pr55Gag binds genomic and spliced RNAs with different affinity and stoichiometry. RNA Biol. 14, 90–103 (2017).
Abd El-Wahab, E. W. et al. Specific recognition of the HIV-1 genomic RNA by the Gag precursor. Nat. Commun. 5, 4304 (2014).
Smyth, R. P. et al. Mutational interference mapping experiment (MIME) for studying RNA structure and function. Nat. Methods 12, 866–872 (2015).
Brown, J. D. et al. Structural basis for transcriptional start site control of HIV-1 RNA fate. Science 368, 413–417 (2020).
Liao, C. et al. Spacer prioritization in CRISPR–Cas9 immunity is enabled by the leader RNA. Nat. Microbiol. 7, 530–541 (2022).
Sun, L. et al. RNA structure maps across mammalian cellular compartments. Nat. Struct. Mol. Biol. 26, 322–330 (2019).
Aviran, S. & Incarnato, D. Computational approaches for RNA structure ensemble deconvolution from structure probing data: deconvolution of RNA structure ensembles. J. Mol. Biol. 434, 167635 (2022).
Pekarek, L. et al. Cis-mediated interactions of the SARS-CoV-2 frameshift RNA alter its conformations and affect function. Nucleic Acids Res. 51, 728–743 (2023).
Zhu, C. et al. An intranasal ASO therapeutic targeting SARS-CoV-2. Nat. Commun. 13, 4503 (2022).
Lin, Y., Schmidt, B. F., Bruchez, M. P. & McManus, C. J. Structural analyses of NEAT1 lncRNAs suggest long-range RNA interactions that may contribute to paraspeckle architecture. Nucleic Acids Res. 46, 3742–3752 (2018).
Smola, M. J. et al. SHAPE reveals transcript-wide interactions, complex structural domains, and protein interactions across the Xist lncRNA in living cells. Proc. Natl Acad. Sci. USA 113, 10322–10327 (2016).
Lu, Z. et al. Structural modularity of the XIST ribonucleoprotein complex. Nat. Commun. 11, 6163 (2020).
Kim, D. et al. The architecture of SARS-CoV-2 transcriptome. Cell 181, 914–921.e10 (2020).
Schmidt, N. et al. SND1 binds SARS-CoV-2 negative-sense RNA and promotes viral RNA synthesis through NSP9. Cell 186, 4834–4850.e23 (2023).
Chaisson, M. J. P. et al. Resolving the complexity of the human genome using single-molecule sequencing. Nature 517, 608–611 (2015).
Al’Khafaji, A. M. et al. High-throughput RNA isoform sequencing using programmed cDNA concatenation. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-01815-7 (2023).
Deigan, K. E., Li, T. W., Mathews, D. H. & Weeks, K. M. Accurate SHAPE-directed RNA structure determination. Proc. Natl Acad. Sci. USA 106, 97–102 (2009).
Rice, G. M., Leonard, C. W. & Weeks, K. M. RNA secondary structure modeling at consistent high accuracy using differential SHAPE. RNA 20, 846–854 (2014).
Breaker, R. R. Riboswitches and the RNA world. Cold Spring Harb. Perspect. Biol. 4, a003566 (2012).
Chen, Y. et al. Context-aware transcript quantification from long-read RNA-seq data with Bambu. Nat. Methods 20, 1187–1195 (2023).
Hamada, M., Ono, Y., Asai, K. & Frith, M. C. Training alignment parameters for arbitrary sequencers with LAST-TRAIN. Bioinformatics 33, 926–928 (2017).
Incarnato, D., Morandi, E., Simon, L. M. & Oliviero, S. RNA Framework: an all-in-one toolkit for the analysis of RNA structures and post-transcriptional modifications. Nucleic Acids Res. 46, e97 (2018).
Wayment-Steele, H. K. et al. RNA secondary structure packages evaluated and improved by high-throughput experiments. Nat. Methods 19, 1234–1242 (2022).
Darty, K., Denise, A. & Ponty, Y. VARNA: interactive drawing and editing of the RNA secondary structure. Bioinformatics 25, 1974–1975 (2009).
Gibbs, J. S., Regier, D. A. & Desrosiers, R. C. Construction and in vitro properties of HIV-1 mutants with deletions in ‘nonessential’ genes. AIDS Res. Hum. Retroviruses 10, 343–350 (1994).
Acknowledgements
This study was funded by the Helmholtz Association (VH-NG-1347 to R.P.S.) and the National Institutes of Health Center for HIV RNA Studies (SUBK00019361 to R.P.S). A.-S.G.-B. was supported with a fellowship from the Peter und Traudl Engelhorn Stiftung and a Post Doc Plus funding (Graduate School of Life Sciences, University of Würzburg).
Author information
Authors and Affiliations
Contributions
All authors conceived the study and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Peer review
Peer review information
Nature Protocols thanks Philip Bevilacqua and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Bohn, P. et al. Nat. Methods 20, 849–859 (2023): https://doi.org/10.1038/s41592-023-01862-7
Extended data
Extended Data Fig. 1 Flowchart of bioinformatic analysis.
Each major stage of the analysis is shown highlighted in colored boxes and QC steps are shown on the right side. Files generated during the different steps of the analysis are depicted in boxes. The tools/analysis steps are shown as triangular arrows, with options highlighted in hexagons.
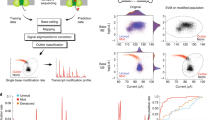
Extended Data Fig. 2 Effect of subsampling on Pearson correlation coefficient between DMS reactivities of different isoforms of both replicates.
(a) Pearson correlation coefficient within multiple subsample iterations at the same subsampling depth reveal isoforms where all variation has been sufficiently sampled (correlation coefficient >0.9 at subsampling rate of 50%), and those where the underlying diversity is not yet fully sampled (correlation coefficient <0.9 at subsampling rate of 50%). (b) Pearson correlation coefficient between replicates 1 and 2 of each sample and isoform at different subsampling depths as a quality control measure of reproducibility of DMS probing data. Plateauing of the correlation coefficient below 0.9 with increasing coverage may indicate that stochastic effects during reverse transcription due to low number of fully reverse transcribed molecules may have resulted in divergent cDNA mutation pools.
Source data
Source Data Fig. 2
Raw data for Fig. 2c.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gribling-Burrer, AS., Bohn, P. & Smyth, R.P. Isoform-specific RNA structure determination using Nano-DMS-MaP. Nat Protoc 19, 1835–1865 (2024). https://doi.org/10.1038/s41596-024-00959-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-024-00959-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.