Abstract
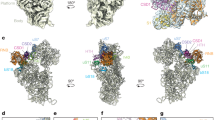
The bacterial ribosome is recycled into subunits by two conserved proteins, elongation factor G (EF-G) and the ribosome recycling factor (RRF). The molecular basis for ribosome recycling by RRF and EF-G remains unclear. Here, we report the crystal structure of a posttermination Thermus thermophilus 70S ribosome complexed with EF-G, RRF and two transfer RNAs at a resolution of 3.5 Å. The deacylated tRNA in the peptidyl (P) site moves into a previously unsuspected state of binding (peptidyl/recycling, p/R) that is analogous to that seen during initiation. The terminal end of the p/R-tRNA forms nonfavorable contacts with the 50S subunit while RRF wedges next to central inter-subunit bridges, illuminating the active roles of tRNA and RRF in dissociation of ribosomal subunits. The structure uncovers a missing snapshot of tRNA as it transits between the P and exit (E) sites, providing insights into the mechanisms of ribosome recycling and tRNA translocation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Atomic coordinates and structure factors have been deposited in the PDB under the accession code 6UCQ.
References
Janosi, L., Hara, H., Zhang, S. & Kaji, A. Ribosome recycling by ribosome recycling factor (RRF)—an important but overlooked step of protein biosynthesis. Adv. Biophys. 32, 121–201 (1996).
Savelsbergh, A., Rodnina, M. V. & Wintermeyer, W. Distinct functions of elongation factor G in ribosome recycling and translocation. RNA 15, 772–780 (2009).
Borg, A., Pavlov, M. & Ehrenberg, M. Complete kinetic mechanism for recycling of the bacterial ribosome. RNA 22, 10–21 (2016).
Fujiwara, T., Ito, K., Yamami, T. & Nakamura, Y. Ribosome recycling factor disassembles the post-termination ribosomal complex independent of the ribosomal translocase activity of elongation factor G. Mol. Microbiol. 53, 517–528 (2004).
Prabhakar, A., Capece, M. C., Petrov, A., Choi, J. & Puglisi, J. D. Post-termination ribosome intermediate acts as the gateway to ribosome recycling. Cell Rep. 20, 161–172 (2017).
Chen, Y., Kaji, A., Kaji, H. & Cooperman, B. S. The kinetic mechanism of bacterial ribosome recycling. Nucl. Acids Res. 45, 10168–10177 (2017).
Zavialov, A. V., Hauryliuk, V. V. & Ehrenberg, M. Splitting of the posttermination ribosome into subunits by the concerted action of RRF and EF-G. Mol. Cell 18, 675–686 (2005).
Peske, F., Rodnina, M. V. & Wintermeyer, W. Sequence of steps in ribosome recycling as defined by kinetic analysis. Mol. Cell 18, 403–412 (2005).
Karimi, R., Pavlov, M. Y., Buckingham, R. H. & Ehrenberg, M. Novel roles for classical factors at the interface between translation termination and initiation. Mol. Cell 3, 601–609 (1999).
Yokoyama, T. et al. Structural insights into initial and intermediate steps of the ribosome-recycling process. EMBO J. 31, 1836–1846 (2012).
Fu, Z. et al. Key intermediates in ribosome recycling visualized by time-resolved cryoelectron microscopy. Structure 24, 2092–2101 (2016).
Gao, N., Zavialov, A. V., Ehrenberg, M. & Frank, J. Specific interaction between EF-G and RRF and its implication for GTP-dependent ribosome splitting into subunits. J. Mol. Biol. 374, 1345–1358 (2007).
Borovinskaya, M. A. et al. Structural basis for aminoglycoside inhibition of bacterial ribosome recycling. Nat. Struct. Mol. Biol. 14, 727–732 (2007).
Weixlbaumer, A. et al. Crystal structure of the ribosome recycling factor bound to the ribosome. Nat. Struct. Mol. Biol. 14, 733–737 (2007).
Dunkle, J. A. et al. Structures of the bacterial ribosome in classical and hybrid states of tRNA binding. Science 332, 981–984 (2011).
Pavlov, M. Y., Antoun, A., Lovmar, M. & Ehrenberg, M. Complementary roles of initiation factor 1 and ribosome recycling factor in 70S ribosome splitting. EMBO J. 27, 1706–1717 (2008).
Agrawal, R. K. et al. Visualization of ribosome-recycling factor on the Escherichia coli 70S ribosome: functional implications. Proc. Natl Acad. Sci. USA 101, 8900–8905 (2004).
Zhou, J., Lancaster, L., Donohue, J. P. & Noller, H. F. How the ribosome hands the A-site tRNA to the P site during EF-G-catalyzed translocation. Science 345, 1188–1191 (2014).
Zhou, J., Lancaster, L., Donohue, J. P. & Noller, H. F. Crystal structures of EF-G-ribosome complexes trapped in intermediate states of translocation. Science 340, 1236086 (2013).
Tourigny, D. S., Fernandez, I. S., Kelley, A. C. & Ramakrishnan, V. Elongation factor G bound to the ribosome in an intermediate state of translocation. Science 340, 1235490 (2013).
Valle, M. et al. Locking and unlocking of ribosomal motions. Cell 114, 123–134 (2003).
Fischer, N., Konevega, A. L., Wintermeyer, W., Rodnina, M. V. & Stark, H. Ribosome dynamics and tRNA movement by time-resolved electron cryomicroscopy. Nature 466, 329–333 (2010).
Moazed, D. & Noller, H. F. Intermediate states in the movement of transfer RNA in the ribosome. Nature 342, 142–148 (1989).
Ramrath, D. J. et al. Visualization of two transfer RNAs trapped in transit during elongation factor G-mediated translocation. Proc. Natl Acad. Sci. USA 110, 20964–20969 (2013).
Lin, J., Gagnon, M. G., Bulkley, D. & Steitz, T. A. Conformational changes of elongation factor G on the ribosome during tRNA translocation. Cell 160, 219–227 (2015).
Pulk, A. & Cate, J. H. Control of ribosomal subunit rotation by elongation factor G. Science 340, 1235970 (2013).
Gao, Y. G. et al. The structure of the ribosome with elongation factor G trapped in the posttranslocational state. Science 326, 694–699 (2009).
Salsi, E., Farah, E., Netter, Z., Dann, J. & Ermolenko, D. N. Movement of elongation factor G between compact and extended conformations. J. Mol. Biol. 427, 454–467 (2015).
Bulkley, D. et al. The antibiotics dityromycin and GE82832 bind protein S12 and block EF-G-catalyzed translocation. Cell Rep. 6, 357–365 (2014).
Brandi, L. et al. Structural and functional characterization of the bacterial translocation inhibitor GE82832. FEBS Lett. 586, 3373–3378 (2012).
Gagnon, M. G., Lin, J., Bulkley, D. & Steitz, T. A. Crystal structure of elongation factor 4 bound to a clockwise ratcheted ribosome. Science 345, 684–687 (2014).
Gagnon, M. G., Lin, J. & Steitz, T. A. Elongation factor 4 remodels the A-site tRNA on the ribosome. Proc. Natl Acad. Sci. USA 113, 4994–4999 (2016).
Polikanov, Y. S., Melnikov, S. V., Soll, D. & Steitz, T. A. Structural insights into the role of rRNA modifications in protein synthesis and ribosome assembly. Nat. Struct. Mol. Biol. 22, 342–344 (2015).
Koripella, R. K., Sharma, M. R., Risteff, P., Keshavan, P. & Agrawal, R. K. Structural insights into unique features of the human mitochondrial ribosome recycling. Proc. Natl Acad. Sci. USA 116, 8283–8288 (2019).
Gao, N. et al. Mechanism for the disassembly of the posttermination complex inferred from cryo-EM studies. Mol. Cell 18, 663–674 (2005).
Sternberg, S. H., Fei, J., Prywes, N., McGrath, K. A. & Gonzalez, R. L. Jr. Translation factors direct intrinsic ribosome dynamics during translation termination and ribosome recycling. Nat. Struct. Mol. Biol. 16, 861–868 (2009).
Guo, P., Zhang, L., Zhang, H., Feng, Y. & Jing, G. Domain II plays a crucial role in the function of ribosome recycling factor. Biochem. J. 393, 767–777 (2006).
Rhodin, M. H. & Dinman, J. D. A flexible loop in yeast ribosomal protein L11 coordinates P-site tRNA binding. Nucl. Acids Res. 38, 8377–8389 (2010).
Budkevich, T. V. et al. Regulation of the mammalian elongation cycle by subunit rolling: a eukaryotic-specific ribosome rearrangement. Cell 158, 121–131 (2014).
Selmer, M. et al. Structure of the 70S ribosome complexed with mRNA and tRNA. Science 313, 1935–1942 (2006).
Allen, G. S., Zavialov, A., Gursky, R., Ehrenberg, M. & Frank, J. The cryo-EM structure of a translation initiation complex from Escherichia coli. Cell 121, 703–712 (2005).
Fernandez, I. S. et al. Molecular architecture of a eukaryotic translational initiation complex. Science 342, 1240585 (2013).
Simonetti, A. et al. Structure of the 30S translation initiation complex. Nature 455, 416–420 (2008).
Sprink, T. et al. Structures of ribosome-bound initiation factor 2 reveal the mechanism of subunit association. Sci. Adv. 2, e1501502 (2016).
Kaledhonkar, S. et al. Late steps in bacterial translation initiation visualized using time-resolved cryo-EM. Nature 570, 400–404 (2019).
Rao, A. R. & Varshney, U. Specific interaction between the ribosome recycling factor and the elongation factor G from Mycobacterium tuberculosis mediates peptidyl-tRNA release and ribosome recycling in Escherichia coli. EMBO J. 20, 2977–2986 (2001).
Iwakura, N. et al. Chemical and structural characterization of a model post-termination complex (PoTC) for the ribosome recycling reaction: evidence for the release of the mRNA by RRF and EF-G. PLoS ONE 12, e0177972 (2017).
Korostelev, A., Trakhanov, S., Laurberg, M. & Noller, H. F. Crystal structure of a 70S ribosome-tRNA complex reveals functional interactions and rearrangements. Cell 126, 1065–1077 (2006).
Zhang, J. & Ferre-D’Amare, A. R. The tRNA elbow in structure, recognition and evolution. Life 6, E3 (2016).
Khade, P. & Joseph, S. Functional interactions by transfer RNAs in the ribosome. FEBS Lett. 584, 420–426 (2010).
Atkins, J. F. & Bjork, G. R. A gripping tale of ribosomal frameshifting: extragenic suppressors of frameshift mutations spotlight P-site realignment. Microbiol. Mol. Biol. Rev. 73, 178–210 (2009).
Herr, A. J., Nelson, C. C., Wills, N. M., Gesteland, R. F. & Atkins, J. F. Analysis of the roles of tRNA structure, ribosomal protein L9, and the bacteriophage T4 gene 60 bypassing signals during ribosome slippage on mRNA. J. Mol. Biol. 309, 1029–1048 (2001).
Wang, L. et al. Allosteric control of the ribosome by small-molecule antibiotics. Nat. Struct. Mol. Biol. 19, 957–963 (2012).
Bock, L. V. et al. Energy barriers and driving forces in tRNA translocation through the ribosome. Nat. Struct. Mol. Biol. 20, 1390–1396 (2013).
Pellegrino, S. et al. Structural insights into the role of diphthamide on elongation factor 2 in mRNA reading-frame maintenance. J. Mol. Biol. 430, 2677–2687 (2018).
Natchiar, S. K., Myasnikov, A. G., Kratzat, H., Hazemann, I. & Klaholz, B. P. Visualization of chemical modifications in the human 80S ribosome structure. Nature 551, 472–477 (2017).
Junemann, R. et al. In vivo deuteration of transfer RNAs: overexpression and large-scale purification of deuterated specific tRNAs. Nucl. Acids Res. 24, 907–913 (1996).
Gagnon, M. G., Seetharaman, S. V., Bulkley, D. & Steitz, T. A. Structural basis for the rescue of stalled ribosomes: structure of YaeJ bound to the ribosome. Science 335, 1370–1372 (2012).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Cryst. 26, 795–800 (1993).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D. Biol. Crystallogr. 58, 1948–1954 (2002).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
The PyMOL molecular graphics system, v.2.1 (Schrodinger L.L.C., 2015).
Acknowledgements
We thank M. Garcia-Blanco and H. Lander for critical reading of the manuscript and valuable suggestions. We are grateful to Y. Polikanov and M. White for their kind help and advice with the Oxford 700 series cryostream cooler. We also thank the staff of the Advanced Photon Source NE-CAT beamline 24-ID-C and of the Shanghai Synchrotron Radiation Facility beamline BL17U for help with data collection. This work was supported by grants from the National Key R&D Program of China (no. 2017YFA0504602) and the National Natural Science Foundation of China (no. 31770784) (to J.L.), and by startup funds from the University of Texas Medical Branch (to M.G.G.), by a McLaughlin Fellowship from the Institute for Human Infections and Immunity at the University of Texas Medical Branch (to M.G.G.), by a Rising Science and Technology Acquisition and Retention Program award (to M.G.G.) and by an endowment from Sealy and Smith Foundation to Sealy Center for Structural Biology at the University of Texas Medical Branch. This work is based on research conducted at the Northeastern Collaborative Access Team beamlines, which are funded by the National Institute of General Medical Sciences from the National Institutes of Health (grant no. P41-GM103403 to NE-CAT). The Pilatus 6M detector on 24-ID-C beamline is funded by the NIH–ORIP HEI Grant (no. S10-RR029205 to NE-CAT). This research used resources of the Advanced Photon Source, a US Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory (contract no. DE-AC02-06CH11357).
Author information
Authors and Affiliations
Contributions
D.Z. and T.T. performed experiments, analyzed the data and made the figures. J.L. and M.G.G. designed the study; purified ribosomes, protein factors and tRNAPhe; performed experiments, determined the structure, analyzed the data and wrote the manuscript with input from all authors. All authors discussed and commented on the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Inês Chen was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Difference Fourier map of the 70S ribosome recycling complex and conformation of the ribosome.
a, Unbiased Fo – Fc difference electron density map contoured at 2.0 σ following the first round of refinement. Both deacylated tRNAs in the P and E sites, together with the mRNA, RRF, and EF-G were clearly visible. b, In the 70S ribosome recycling complex (orange), the 30S subunit rotates counter-clockwise by ~2° compared to the classical state ribosome (blue).
Extended Data Fig. 2 Conformation of RRF on the ribosome in the recycling complex.
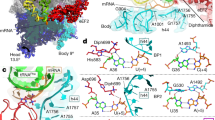
a, Interactions between domain I of RRF and the ribosomal RNA. Shown are parts of H74 and H80 of the 23S rRNA (white), the CCA-end of the p/R-tRNA (pink), and domain I of RRF (blue). Key interacting residues in RRF are shown. b, Compared with the E. coli 70S ribosome bound to RRF (PDB 4V5413), domain II of RRF in the current structure rotates by ~28° and becomes wedged between the 23S rRNA helix H69 (orange) and ribosomal protein uS12 (magenta).
Extended Data Fig. 3 Difference Fourier map of the CCA-end of the p/R-tRNA.
a, Unbiased Fo – Fc map contoured at 2.0 σ of the 3′ end of the p/R-tRNA showing its location relative to the 23S rRNA helices H74 and H80. b, Rotated view of panel a with ribosomal elements omitted for clarity.
Extended Data Fig. 4 Conformation of anticodon stem loop (ASL) of tRNAs in the recycling complex.
a, Unstable state of binding of the p/R-tRNA ASL in the P site of the ribosome recycling complex. The unbiased Fo – Fc difference Fourier map contoured at 2.0 σ of the ASL of the p/R-tRNA shows that the electron density is much clearer for the nucleotide bases than for the phosphate backbone. b, Average Wilson B-factor values for the phosphate backbone (blue bars) and the nucleotide bases (orange bars) of the p/R-tRNA ASL. c, Movements of ASLs in the P and E site in the recycling complex compared to tRNAs bound in the classical state (gray).
Extended Data Fig. 5 Conformation of the p/R-tRNA compared to that of the p/I state during initiation.
Superimposition of the current recycling complex with that of a eukaryotic initiation 80S complex bound to eIF5B (orange) (PDB 4V8Z42) shows that like the IF2-70S initiation complex (Fig. 5), the conformation of the p/R-tRNA (magenta) resembles that of the p/I-tRNA (yellow). The same molecular mimic is observed between RRF/EF-G and eIF5B as that seen with IF2.
Supplementary information
Supplementary Video 1
Structure of the bacterial ribosome recycling complex. Animation showing the interactions between EF-G and RRF with the ribosome and the p/R-tRNA. The conformation of tRNAs in the recycling complex is compared with that of tRNAs bound in the canonical e/E and p/P states. The position of domain II of RRF is compared to a previous ribosome-bound RRF structure in the absence of EF-G (PDB 4V54, ref. 13). The RRF-mediated displacement of the P-site tRNA from the p/P to the p/R state is guided by ribosomal protein uL5 shown in green.
Rights and permissions
About this article
Cite this article
Zhou, D., Tanzawa, T., Lin, J. et al. Structural basis for ribosome recycling by RRF and tRNA. Nat Struct Mol Biol 27, 25–32 (2020). https://doi.org/10.1038/s41594-019-0350-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0350-7
This article is cited by
-
Quality control of protein synthesis in the early elongation stage
Nature Communications (2023)
-
Structural basis for clearing of ribosome collisions by the RQT complex
Nature Communications (2023)
-
Molecular basis of the pleiotropic effects by the antibiotic amikacin on the ribosome
Nature Communications (2023)
-
Antibiotic thermorubin tethers ribosomal subunits and impedes A-site interactions to perturb protein synthesis in bacteria
Nature Communications (2023)
-
The expanding world of tRNA modifications and their disease relevance
Nature Reviews Molecular Cell Biology (2021)