Abstract
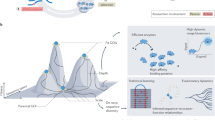
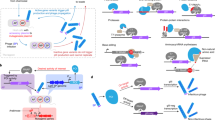
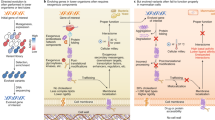
Continuous directed evolution methods allow the key steps of evolution—gene diversification, selection, and replication—to proceed in the laboratory with minimal researcher intervention. As a result, continuous evolution can find solutions much more quickly than traditional discrete evolution methods. Continuous evolution also enables the exploration of longer and more numerous evolutionary trajectories, increasing the likelihood of accessing solutions that require many steps through sequence space and greatly facilitating the iterative refinement of selection conditions and targeted mutagenesis strategies. Here we review the historical advances that have expanded continuous evolution from its earliest days as an experimental curiosity to its present state as a powerful and surprisingly general strategy for generating tailor-made biomolecules, and discuss more recent improvements with an eye to the future.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Mills, D. R., Peterson, R. L. & Spiegelman, S. An extracellular Darwinian experiment with a self-duplicating nucleic acid molecule. Proc. Natl Acad. Sci. USA 58, 217–224 (1967).
Wright, M. C. & Joyce, G. F. Continuous in vitro evolution of catalytic function. Science 276, 614–617 (1997).
McGinness, K. E., Wright, M. C. & Joyce, G. F. Continuous in vitro evolution of a ribozyme that catalyzes three successive nucleotidyl addition reactions. Chem. Biol. 9, 585–596 (2002).
Breaker, R. R., Banerji, A. & Joyce, G. F. Continuous in vitro evolution of bacteriophage RNA polymerase promoters. Biochemistry 33, 11980–11986 (1994).
Kühne, H. & Joyce, G. F. Continuous in vitro evolution of ribozymes that operate under conditions of extreme pH. J. Mol. Evol. 57, 292–298 (2003).
Voytek, S. B. & Joyce, G. F. Emergence of a fast-reacting ribozyme that is capable of undergoing continuous evolution. Proc. Natl Acad. Sci. USA 104, 15288–15293 (2007).
Matsuura, T. & Yomo, T. In vitro evolution of proteins. J. Biosci. Bioeng. 101, 449–456 (2006).
Monod, J. La technique de culture continue: theorie et applications. Ann. Inst. Pasteur 79, 390–410 (1950).
Novick, A. & Szilard, L. Experiments with the Chemostat on spontaneous mutations of bacteria. Proc. Natl Acad. Sci. USA 36, 708–719 (1950).
Bryson, V. & Szybalski, W. Microbial selection. Science 116, 45–51 (1952).
Larsson, G., Enfors, S. O. & Pham, H. The pH-auxostat as a tool for studying microbial dynamics in continuous fermentation. Biotechnol. Bioeng. 36, 224–232 (1990).
Moser, H. The Dynamics of Bacterial Populations Maintained in the Chemostat. (Washington D.C., Carnegie Institution of Washington, 1958).
Bull, J. J. et al. Exceptional convergent evolution in a virus. Genetics 147, 1497–1507 (1997).
Marlière, P. et al. Chemical evolution of a bacterium’s genome. Angew. Chem. Int. Ed. Engl. 50, 7109–7114 (2011).
Toprak, E. et al. Evolutionary paths to antibiotic resistance under dynamically sustained drug selection. Nat. Genet. 44, 101–105 (2011).
Zamenhof, S. & Eichhorn, H. H. Study of microbial evolution through loss of biosynthetic functions: establishment of “defective” mutants. Nature 216, 456–458 (1967).
Lwoff, A. L’évolution Physiologique: Étude des Pertes de Fonctions chez les Microorganismes. (Paris, Hermann, 1944).
Lenski, R. E. & Levin, B. R. Constraints on the coevolution of bacteria and virulent phage: a model, some experiments, and predictions for natural communities. Am. Nat. 125, 585–602 (1985).
Hillis, D. M., Bull, J. J., White, M. E., Badgett, M. R. & Molineux, I. J. Experimental phylogenetics: generation of a known phylogeny. Science 255, 589–592 (1992).
Greener, A., Callahan, M. & Jerpseth, B. An efficient random mutagenesis technique using an E. coli mutator strain. Mol. Biotechnol. 7, 189–195 (1997).
Chou, H. H. & Keasling, J. D. Programming adaptive control to evolve increased metabolite production. Nat. Commun. 4, 2595 (2013).
Pham, H. L. et al. Engineering a riboswitch-based genetic platform for the self-directed evolution of acid-tolerant phenotypes. Nat. Commun. 8, 411 (2017).
Badran, A. H. & Liu, D. R. Development of potent in vivo mutagenesis plasmids with broad mutational spectra. Nat. Commun. 6, 8425 (2015).
Wang, H. H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009).
Mundhada, H. et al. Increased production of l-serine in Escherichia coli through adaptive laboratory evolution. Metab. Eng. 39, 141–150 (2017).
Amiram, M. et al. Evolution of translation machinery in recoded bacteria enables multi-site incorporation of nonstandard amino acids. Nat. Biotechnol. 33, 1272–1279 (2015).
Martin, R. W. et al. Cell-free protein synthesis from genomically recoded bacteria enables multisite incorporation of noncanonical amino acids. Nat. Commun. 9, 1203 (2018).
Barbieri, E. M., Muir, P., Akhuetie-Oni, B. O., Yellman, C. M. & Isaacs, F. J. Precise editing at DNA replication forks enables multiplex genome engineering in eukaryotes. Cell 171, 1453–1467.e13 (2017).
Garst, A. D. et al. Genome-wide mapping of mutations at single-nucleotide resolution for protein, metabolic and genome engineering. Nat. Biotechnol. 35, 48–55 (2017).
Yano, T., Oue, S. & Kagamiyama, H. Directed evolution of an aspartate aminotransferase with new substrate specificities. Proc. Natl Acad. Sci. USA 95, 5511–5515 (1998).
Ravikumar, A., Arzumanyan, G. A., Obadi, M. K. A., Javanpour, A. A. & Liu, C. C. Scalable, continuous evolution of genes at mutation rates above genomic error thresholds. Cell 175, 1957.e13 (2018).
Fabret, C. et al. Efficient gene targeted random mutagenesis in genetically stable Escherichia coli strains. Nucleic Acids Res. 28, E95 (2000).
Camps, M., Naukkarinen, J., Johnson, B. P. & Loeb, L. A. Targeted gene evolution in Escherichia coli using a highly error-prone DNA polymerase I. Proc. Natl Acad. Sci. USA 100, 9727–9732 (2003).
Ravikumar, A., Arrieta, A. & Liu, C. C. An orthogonal DNA replication system in yeast. Nat. Chem. Biol. 10, 175–177 (2014).
Crook, N. et al. In vivo continuous evolution of genes and pathways in yeast. Nat. Commun. 7, 13051 (2016).
Simon, A. J., Morrow, B. R. & Ellington, A. D. Retroelement-based genome editing and evolution. ACS Synth. Biol. 7, 2600–2611 (2018).
Romanini, D. W., Peralta-Yahya, P., Mondol, V. & Cornish, V. W. A heritable recombination system for synthetic Darwinian evolution in yeast. ACS Synth. Biol. 1, 602–609 (2012).
Finney-Manchester, S. P. & Maheshri, N. Harnessing mutagenic homologous recombination for targeted mutagenesis in vivo by TaGTEAM. Nucleic Acids Res. 41, e99 (2013).
Moore, C. L., Papa, L. J. III & Shoulders, M. D. A processive protein chimera introduces mutations across defined DNA regions in vivo. J. Am. Chem. Soc. 140, 11560–11564 (2018).
Hess, G. T. et al. Directed evolution using dCas9-targeted somatic hypermutation in mammalian cells. Nat. Methods 13, 1036–1042 (2016).
Halperin, S. O. et al. CRISPR-guided DNA polymerases enable diversification of all nucleotides in a tunable window. Nature 560, 248–252 (2018).
Esvelt, K. M., Carlson, J. C. & Liu, D. R. A system for the continuous directed evolution of biomolecules. Nature 472, 499–503 (2011).
Smeal, S. W., Schmitt, M. A., Pereira, R. R., Prasad, A. & Fisk, J. D. Simulation of the M13 life cycle I: assembly of a genetically-structured deterministic chemical kinetic simulation. Virology 500, 259–274 (2017).
Carlson, J. C., Badran, A. H., Guggiana-Nilo, D. A. & Liu, D. R. Negative selection and stringency modulation in phage-assisted continuous evolution. Nat. Chem. Biol. 10, 216–222 (2014).
Dickinson, B. C., Packer, M. S., Badran, A. H. & Liu, D. R. A system for the continuous directed evolution of proteases rapidly reveals drug-resistance mutations. Nat. Commun. 5, 5352 (2014).
Packer, M. S., Rees, H. A. & Liu, D. R. Phage-assisted continuous evolution of proteases with altered substrate specificity. Nat. Commun. 8, 956 (2017).
Badran, A. H. et al. Continuous evolution of Bacillus thuringiensis toxins overcomes insect resistance. Nature 533, 58–63 (2016).
Wang, T., Badran, A. H., Huang, T. P. & Liu, D. R. Continuous directed evolution of proteins with improved soluble expression. Nat. Chem. Biol. 14, 972–980 (2018).
Bryson, D. I. et al. Continuous directed evolution of aminoacyl-tRNA synthetases. Nat. Chem. Biol. 13, 1253–1260 (2017).
Hubbard, B. P. et al. Continuous directed evolution of DNA-binding proteins to improve TALEN specificity. Nat. Methods 12, 939–942 (2015).
Hu, J. H. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556, 57–63 (2018).
Miller, S.M. et al. Continuous evolution of SpCas9 variants compatible with non-G PAMs. Nat. Biotechnol. (2020).
Thuronyi, B.W. et al. Continuous evolution of base editors with expanded target compatibility and improved activity. Nat. Biotechnol. 37, 1070–1079 (2019).
Berman, C. M. et al. An adaptable platform for directed evolution in human cells. J. Am. Chem. Soc. 140, 18093–18103 (2018).
English, J. G. et al. VEGAS as a platform for facile directed evolution in mammalian cells. Cell 178, 748–761.e7 (2019).
Butterfield, G. L. et al. Evolution of a designed protein assembly encapsulating its own RNA genome. Nature 552, 415–420 (2017).
Voigt, C. A., Martinez, C., Wang, Z. G., Mayo, S. L. & Arnold, F. H. Protein building blocks preserved by recombination. Nat. Struct. Biol. 9, 553–558 (2002).
Wu, Z., Kan, S. B. J., Lewis, R. D., Wittmann, B. J. & Arnold, F. H. Machine learning-assisted directed protein evolution with combinatorial libraries. Proc. Natl Acad. Sci. USA 116, 8852–8858 (2019).
Wong, J. T. Membership mutation of the genetic code: loss of fitness by tryptophan. Proc. Natl Acad. Sci. USA 80, 6303–6306 (1983).
Wang, L. & Schultz, P. G. A general approach for the generation of orthogonal tRNAs. Chem. Biol. 8, 883–890 (2001).
Monk, J. W. et al. Rapid and inexpensive evaluation of nonstandard amino acid incorporation in Escherichia coli. ACS Synth. Biol. 6, 45–54 (2017).
Hammerling, M. J. et al. Bacteriophages use an expanded genetic code on evolutionary paths to higher fitness. Nat. Chem. Biol. 10, 178–180 (2014).
Drienovská, I., Mayer, C., Dulson, C. & Roelfes, G. A designer enzyme for hydrazone and oxime formation featuring an unnatural catalytic aniline residue. Nat. Chem. 10, 946–952 (2018).
Mayer, C., Dulson, C., Reddem, E., Thunnissen, A. W. H. & Roelfes, G. Directed evolution of a designer enzyme featuring an unnatural catalytic amino acid. Angew. Chem. Int. Ed. Engl. 58, 2083–2087 (2019).
Kanigowska, P., Shen, Y., Zheng, Y., Rosser, S. & Cai, Y. Smart DNA fabrication using sound waves: applying acoustic dispensing technologies to synthetic biology. J. Lab. Autom. 21, 49–56 (2016).
Terekhov, S. S. et al. Microfluidic droplet platform for ultrahigh-throughput single-cell screening of biodiversity. Proc. Natl Acad. Sci. USA 114, 2550–2555 (2017).
Wong, B. G., Mancuso, C. P., Kiriakov, S., Bashor, C. J. & Khalil, A. S. Precise, automated control of conditions for high-throughput growth of yeast and bacteria with eVOLVER. Nat. Biotechnol. 36, 614–623 (2018).
Callens, C. et al. A multiplex culture system for the long-term growth of fission yeast cells. Yeast 34, 343–355 (2017).
Takahashi, C. N., Miller, A. W., Ekness, F., Dunham, M. J. & Klavins, E. A low cost, customizable turbidostat for use in synthetic circuit characterization. ACS Synth. Biol. 4, 32–38 (2015).
Acknowledgements
D.R.L. gratefully acknowledges support from NIH U01 AI142756 (D.R.L.), RM1 HG009490 (D.R.L.), R01 EB022376 (D.R.L.), and R35 GM118062 (D.R.L.); and HHMI (D.R.L.). We thank A. Badran and K. Zhao for their helpful comments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
D.R.L. is a consultant and co-founder of Prime Medicine, Beam Therapeutics, Pairwise Plants, and Editas Medicine, companies that use genome editing. Complete disclosures are available at https://liugroup.us.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Morrison, M.S., Podracky, C.J. & Liu, D.R. The developing toolkit of continuous directed evolution. Nat Chem Biol 16, 610–619 (2020). https://doi.org/10.1038/s41589-020-0532-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0532-y
This article is cited by
-
Ultrahigh-throughput screening-assisted in vivo directed evolution for enzyme engineering
Biotechnology for Biofuels and Bioproducts (2024)
-
Engineering is evolution: a perspective on design processes to engineer biology
Nature Communications (2024)
-
Quantification of evolved DNA-editing enzymes at scale with DEQSeq
Genome Biology (2023)
-
Engineered bacterial orthogonal DNA replication system for continuous evolution
Nature Chemical Biology (2023)
-
Advances in biosynthesis of higher alcohols in Escherichia coli
World Journal of Microbiology and Biotechnology (2023)