Abstract
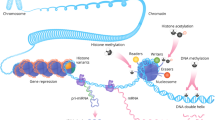
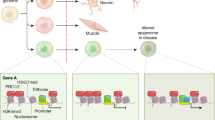
An increasing number of epilepsies are being attributed to variants in genes with epigenetic functions. The products of these genes include factors that regulate the structure and function of chromatin and the placing, reading and removal of epigenetic marks, as well as other epigenetic processes. In this Review, we provide an overview of the various epigenetic processes, structuring our discussion around five function-based categories: DNA methylation, histone modifications, histone–DNA crosstalk, non-coding RNAs and chromatin remodelling. We provide background information on each category, describing the general mechanism by which each process leads to altered gene expression. We also highlight key clinical and mechanistic aspects, providing examples of genes that strongly associate with epilepsy within each class. We consider the practical applications of these findings, including tissue-based and biofluid-based diagnostics and precision medicine-based treatments. We conclude that variants in epigenetic genes are increasingly found to be causally involved in the epilepsies, with implications for disease mechanisms, treatments and diagnostics.
Key points
-
The term epigenetics refers to potentially heritable changes in gene expression that do not involve alterations in the DNA sequence; the key epigenetic processes include DNA methylation, histone modifications and the actions of certain non-coding RNAs.
-
Various monogenic forms of epilepsy have been attributed to pathogenic variants in genes encoding factors that regulate chromatin access and the deposition, reading and removal of epigenetic marks.
-
Epigenetics-related epilepsies are often accompanied by a range of comorbidities, including intellectual disability.
-
Insights from experimental studies in cell and animal models are helping us to understand how epigenetic alterations give rise to neuronal hyperexcitability.
-
The findings of this research might yield diagnostic or prognostic biomarkers or treatment strategies, including precision medicine-based treatments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Fiest, K. M. et al. Prevalence and incidence of epilepsy. Neurology 88, 296–303 (2017).
Thakran, S. et al. Genetic landscape of common epilepsies: advancing towards precision in treatment. Int. J. Mol. Sci. 21, 7784 (2020).
Wang, J. et al. Epilepsy-associated genes. Seizure 44, 11–20 (2017).
Oyrer, J. et al. Ion channels in genetic epilepsy: from genes and mechanisms to disease-targeted therapies. Pharmacol. Rev. 70, 142–173 (2018).
Kobow, K. et al. Epigenetics explained: a topic “primer” for the epilepsy community by the ILAE genetics/epigenetics task force. Epileptic Disord. 22, 127–141 (2020).
Steinlein, O. K. et al. A missense mutation in the neuronal nicotinic acetylcholine receptor α4 subunit is associated with autosomal dominant nocturnal frontal lobe epilepsy. Nat. Genet. 11, 201–203 (1995).
Reid, C. A., Berkovic, S. F. & Petrou, S. Mechanisms of human inherited epilepsies. Prog. Neurobiol. 87, 41–57 (2009).
Allen, A. S. et al. De novo mutations in epileptic encephalopathies. Nature 501, 217–221 (2013).
Poduri, A., Evrony, G. D., Cai, X. & Walsh, C. A. Somatic mutation, genomic variation, and neurological disease. Science 341, 1237758 (2013).
Epi4K consortium & Epilepsy Phenome/Genome Project. Ultra-rare genetic variation in common epilepsies: a case-control sequencing study. Lancet Neurol. 16, 135–143 (2017).
Dibbens, L. M. et al. Familial and sporadic 15q13.3 microdeletions in idiopathic generalized epilepsy: precedent for disorders with complex inheritance. Hum. Mol. Genet. 18, 3626–3631 (2009).
Helbig, I. et al. 15q13.3 microdeletions increase risk of idiopathic generalized epilepsy. Nat. Genet. 41, 160–162 (2009).
Leu, C. et al. Polygenic burden in focal and generalized epilepsies. Brain 142, 3473–3481 (2019).
Waddington, C. H. The epigenotype. 1942. Int. J. Epidemiol. 41, 10–13 (2012).
Deichmann, U. Epigenetics: the origins and evolution of a fashionable topic. Dev. Biol. 416, 249–254 (2016).
Luger, K., Mäder, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 389, 251–260 (1997).
Morrison, O. & Thakur, J. Molecular complexes at euchromatin, heterochromatin and centromeric chromatin. Int. J. Mol. Sci. 22, 6922 (2021).
Gräff, J., Kim, D., Dobbin, M. M. & Tsai, L.-H. Epigenetic regulation of gene expression in physiological and pathological brain processes. Physiol. Rev. 91, 603–649 (2011).
Biswas, S. & Rao, C. M. Epigenetic tools (The Writers, The Readers and The Erasers) and their implications in cancer therapy. Eur. J. Pharmacol. 837, 8–24 (2018).
Amir, R. E. et al. Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2. Nat. Genet. 23, 185–188 (1999).
Hotchkiss, R. D. The quantitative separation of purines, pyrimidines, and nucleosides by paper chromatography. J. Biol. Chem. 175, 315–332 (1948).
Jabbari, K. & Bernardi, G. Cytosine methylation and CpG, TpG (CpA) and TpA frequencies. Gene 333, 143–149 (2004).
Moore, L. D., Le, T. & Fan, G. DNA methylation and its basic function. Neuropsychopharmacology 38, 23–38 (2013).
Bird, A. P. CpG-rich islands and the function of DNA methylation. Nature 321, 209–213 (1986).
Cedar, H. DNA methylation and gene activity. Cell 53, 3–4 (1988).
Hark, A. T. et al. CTCF mediates methylation-sensitive enhancer-blocking activity at the H19/Igf2 locus. Nature 405, 486–489 (2000).
Ragione, F. D., Vacca, M., Fioriniello, S., Pepe, G. & D’Esposito, M. MECP2, a multi-talented modulator of chromatin architecture. Brief. Funct. Genomics 15, 420–431 (2016).
Eden, S., Hashimshony, T., Keshet, I., Cedar, H. & Thorne, A. W. DNA methylation models histone acetylation. Nature 394, 842 (1998).
Hashimshony, T., Zhang, J., Keshet, I., Bustin, M. & Cedar, H. The role of DNA methylation in setting up chromatin structure during development. Nat. Genet. 34, 187–192 (2003).
Almouzni, G. & Cedar, H. Maintenance of epigenetic information. Cold Spring Harb. Perspect. Biol. 8, a019372 (2016).
Day, J. J. & Sweatt, J. D. DNA methylation and memory formation. Nat. Neurosci. 13, 1319–1323 (2010).
Sega, A. G. et al. De novo pathogenic variants in neuronal differentiation factor 2 (NEUROD2) cause a form of early infantile epileptic encephalopathy. J. Med. Genet. 56, 113–122 (2019).
Mis, E. K. et al. Expansion of NEUROD2 phenotypes to include developmental delay without seizures. Am. J. Med. Genet. Part A 185, 1076–1080 (2021).
Olson, J. M. et al. NeuroD2 is necessary for development and survival of central nervous system neurons. Dev. Biol. 234, 174–187 (2001).
Telley, L. et al. Sequential transcriptional waves direct the differentiation of newborn neurons in the mouse neocortex. Science 351, 1443–1446 (2016).
Runge, K. et al. Disruption of NEUROD2 causes a neurodevelopmental syndrome with autistic features via cell-autonomous defects in forebrain glutamatergic neurons. Mol. Psychiatry 26, 6125–6148 (2021).
Fong, A. P. et al. Genetic and epigenetic determinants of neurogenesis and myogenesis. Dev. Cell 22, 721–735 (2012).
Hahn, M. A. et al. Reprogramming of DNA methylation at NEUROD2-bound sequences during cortical neuron differentiation. Sci. Adv. 5, eaax0080 (2019).
Chen, F. et al. The transcription factor NeuroD2 coordinates synaptic innervation and cell intrinsic properties to control excitability of cortical pyramidal neurons. J. Physiol. 594, 3729–3744 (2016).
Konishi, Y., Matsu-Ura, T., Mikoshiba, K. & Tamura, T. A. Stimulation of gene expression of NeuroD-related factor in the mouse brain following pentylenetetrazol-induced seizures. Mol. Brain Res. 97, 129–136 (2001).
Lewis, J. D. et al. Purification, sequence, and cellular localization of a novel chromosomal protein that binds to methylated DNA. Cell 69, 905–914 (1992).
Marafi, D. et al. Spectrum and time course of epilepsy and the associated cognitive decline in MECP2 duplication syndrome. Neurology 92, E108–E114 (2019).
Sandweiss, A. J., Brandt, V. L. & Zoghbi, H. Y. Advances in understanding of Rett syndrome and MECP2 duplication syndrome: prospects for future therapies. Lancet Neurol. 19, 689–698 (2020).
Castells, A. A. et al. Unraveling molecular pathways altered in MeCP2-related syndromes, in the search for new potential avenues for therapy. Biomedicines 9, 148 (2021).
Smirnov, K., Stroganova, T., Molholm, S. & Sysoeva, O. Reviewing evidence for the relationship of EEG abnormalities and RTT phenotype paralleled by insights from animal studies. Int. J. Mol. Sci. 22, 5308 (2021).
Tarquinio, D. C. et al. Longitudinal course of epilepsy in Rett syndrome and related disorders. Brain 140, 306–318 (2017).
Henriksen, M. W. et al. Epilepsy in classic Rett syndrome: course and characteristics in adult age. Epilepsy Res. 145, 134–139 (2018).
Marballi, K. & MacDonald, J. L. Proteomic and transcriptional changes associated with MeCP2 dysfunction reveal nodes for therapeutic intervention in Rett syndrome. Neurochem. Int. 148, 105076 (2021).
Marano, D., Fioriniello, S., D’Esposito, M. & Della Ragione, F. Transcriptomic and epigenomic landscape in Rett syndrome. Biomolecules 11, 967 (2021).
Ehrhart, F. et al. Rett syndrome–biological pathways leading from MECP2 to disorder phenotypes. Orphanet J. Rare Dis. 11, 158 (2016).
Zhang, Y. et al. Loss of MeCP2 in cholinergic neurons causes part of RTT-like phenotypes via α7 receptor in hippocampus. Cell Res. 26, 728–742 (2016).
Adebayo, O. L. et al. Intensified mitochondrial hydrogen peroxide release occurs in all brain regions, affects male as well as female Rett mice, and constitutes a life-long burden. Arch. Biochem. Biophys. 696, 108666 (2020).
Allfrey, V. G., Faulkner, R. & Mirsky, A. E. Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc. Natl Acad. Sci. USA 51, 786–794 (1964).
Bannister, A. J. & Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 21, 381–395 (2011).
Berson, A., Nativio, R., Berger, S. L. & Bonini, N. M. Epigenetic regulation in neurodegenerative diseases. Trends Neurosci. 41, 587–598 (2018).
Jakovcevski, M. & Akbarian, S. Epigenetic mechanisms in neurological disease. Nat. Med. 18, 1194–1204 (2012).
Fuks, F. DNA methylation and histone modifications: teaming up to silence genes. Curr. Opin. Genet. Dev. 15, 490–495 (2005).
Kleefstra, T. et al. Loss-of-function mutations in euchromatin histone methyl transferase 1 (EHMT1) cause the 9q34 subtelomeric deletion syndrome. Am. J. Hum. Genet. 79, 370–377 (2006).
Kleefstra, T. & de Leeuw, N. Kleefstra Syndrome (GeneReviews, 2019).
Tachibana, M., Matsumura, Y., Fukuda, M., Kimura, H. & Shinkai, Y. G9a/GLP complexes independently mediate H3K9 and DNA methylation to silence transcription. EMBO J. 27, 2681–2690 (2008).
Tachibana, M. et al. Histone methyltransferases G9a and GLP form heteromeric complexes and are both crucial for methylation of euchromatin at H3-K9. Genes Dev. 19, 815–826 (2005).
Bian, C., Chen, Q. & Yu, X. The zinc finger proteins ZNF644 and WIZ regulate the G9a/GLP complex for gene repression. eLife 4, e05606 (2015).
Meng, T.-G. et al. PRC2 and EHMT1 regulate H3K27me2 and H3K27me3 establishment across the zygote genome. Nat. Commun. 11, 6354 (2020).
Lu, X. et al. GLP-catalyzed H4K16me1 promotes 53BP1 recruitment to permit DNA damage repair and cell survival. Nucleic Acids Res. 47, 10977–10993 (2019).
Davis, B. A. et al. Impairments in sensory-motor gating and information processing in a mouse model of Ehmt1 haploinsufficiency. Brain Neurosci. Adv. 4, 239821282092864 (2020).
Negwer, M. et al. EHMT1 regulates parvalbumin-positive interneuron development and GABAergic input in sensory cortical areas. Brain Struct. Funct. 225, 2701–2716 (2020).
Frega, M. et al. Neuronal network dysfunction in a model for Kleefstra syndrome mediated by enhanced NMDAR signaling. Nat. Commun. 10, 4928 (2019).
Frega, M. et al. Distinct pathogenic genes causing intellectual disability and autism exhibit a common neuronal network hyperactivity phenotype. Cell Rep. 30, 173–186.e6 (2020).
O’Donnell-Luria, A. H. et al. Heterozygous variants in KMT2E cause a spectrum of neurodevelopmental disorders and epilepsy. Am. J. Hum. Genet. 104, 1210–1222 (2019).
Conforti, R. et al. ODLURO syndrome: personal experience and review of the literature. Radiol. Med. 126, 316–322 (2021).
Li, Y. et al. Case report: de novo variants of KMT2E cause O’Donnell-Luria-Rodan syndrome: additional cases and literature review. Front. Pediatr. 9, 641841 (2021).
Sharawat, I. K., Panda, P. K. & Dawman, L. Clinical characteristics and genotype-phenotype correlation in children with KMT2E gene-related neurodevelopmental disorders: report of two new cases and review of published literature. Neuropediatrics 52, 98–104 (2021).
Zhang, X., Novera, W., Zhang, Y. & Deng, L.-W. MLL5 (KMT2E): structure, function, and clinical relevance. Cell. Mol. Life Sci. 74, 2333–2344 (2017).
Emerling, B. M. et al. MLL5, a homolog of Drosophila trithorax located within a segment of chromosome band 7q22 implicated in myeloid leukemia. Oncogene 21, 4849–4854 (2002).
Sebastian, S. et al. MLL5, a trithorax homolog, indirectly regulates H3K4 methylation, represses cyclin A2 expression, and promotes myogenic differentiation. Proc. Natl Acad. Sci. USA 106, 4719–4724 (2009).
Jensen, L. R. et al. Mutations in the JARID1C gene, which is involved in transcriptional regulation and chromatin remodeling, cause X-linked mental retardation. Am. J. Hum. Genet. 76, 227–236 (2005).
Tzschach, A. et al. Novel JARID1C/SMCX mutations in patients with X-linked mental retardation. Hum. Mutat. 27, 389 (2006).
Carmignac, V. et al. Further delineation of the female phenotype with KDM5C disease causing variants: 19 new individuals and review of the literature. Clin. Genet. 98, 43–55 (2020).
Iwase, S. et al. The X-linked mental retardation gene SMCX/JARID1C defines a family of histone H3 lysine 4 demethylases. Cell 128, 1077–1088 (2007).
Tahiliani, M. et al. The histone H3K4 demethylase SMCX links REST target genes to X-linked mental retardation. Nature 447, 601–605 (2007).
Grafodatskaya, D. et al. Multilocus loss of DNA methylation in individuals with mutations in the histone H3 lysine 4 demethylase KDM5C. BMC Med. Genomics 6, 1 (2013).
Brookes, E. et al. Mutations in the intellectual disability gene KDM5C reduce protein stability and demethylase activity. Hum. Mol. Genet. 24, 2861–2872 (2015).
Iwase, S. et al. A mouse model of X-linked intellectual disability associated with impaired removal of histone methylation. Cell Rep. 14, 1000–1009 (2016).
Hiraide, T. et al. De novo variants in SETD1B are associated with intellectual disability, epilepsy and autism. Hum. Genet. 137, 95–104 (2018).
Den, K. et al. A novel de novo frameshift variant in SETD1B causes epilepsy. J. Hum. Genet. 64, 821–827 (2019).
Roston, A. et al. SETD1B-associated neurodevelopmental disorder. J. Med. Genet. 58, 196–204 (2021).
Hiraide, T. et al. De novo variants in SETD1B cause intellectual disability, autism spectrum disorder, and epilepsy with myoclonic absences. Epilepsia Open 4, 476–481 (2019).
Weerts, M. J. A. et al. Delineating the molecular and phenotypic spectrum of the SETD1B-related syndrome. Genet. Med. 23, 2122–2137 (2021).
Lee, J. H., Tate, C. M., You, J. S. & Skalnik, D. G. Identification and characterization of the human Set1B histone H3-Lys4 methyltransferase complex. J. Biol. Chem. 282, 13419–13428 (2007).
Bledau, A. S. et al. The H3K4 methyltransferase Setd1a is first required at the epiblast stage, whereas Setd1b becomes essential after gastrulation. Development 141, 1022–1035 (2014).
Krzyzewska, I. M. et al. A genome-wide DNA methylation signature for SETD1B-related syndrome. Clin. Epigenetics 11, 156 (2019).
Costa, F. F. Non-coding RNAs, epigenetics and complexity. Gene 410, 9–17 (2008).
Wei, J. W., Huang, K., Yang, C. & Kang, C. S. Non-coding RNAs as regulators in epigenetics (review). Oncol. Rep. 37, 3–9 (2017).
Gomes, A., Nolasco, S. & Soares, H. Non-coding RNAs: multi-tasking molecules in the cell. Int. J. Mol. Sci. 14, 16010–16039 (2013).
Kim, D. H., Sætrom, P., Snøve, O. & Rossi, J. J. MicroRNA-directed transcriptional gene silencing in mammalian cells. Proc. Natl Acad. Sci. USA 105, 16230–16235 (2008).
Alles, J. et al. An estimate of the total number of true human miRNAs. Nucleic Acids Res. 47, 3353–3364 (2019).
Gonzalez, S., Pisano, D. G. & Serrano, M. Mechanistic principles of chromatin remodeling guided by siRNAs and miRNAs. Cell Cycle 7, 2601–2608 (2008).
Tuddenham, L. et al. The cartilage specific microRNA-140 targets histone deacetylase 4 in mouse cells. FEBS Lett. 580, 4214–4217 (2006).
Yuan, J. H. et al. The histone deacetylase 4/SP1/microrna-200a regulatory network contributes to aberrant histone acetylation in hepatocellular carcinoma. Hepatology 54, 2025–2035 (2011).
Fabbri, M. et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc. Natl Acad. Sci. USA 104, 15805–15810 (2007).
Benetti, R. et al. A mammalian microRNA cluster controls DNA methylation and telomere recombination via Rbl2-dependent regulation of DNA methyltransferases. Nat. Struct. Mol. Biol. 15, 268–279 (2008).
Sinkkonen, L. et al. MicroRNAs control de novo DNA methylation through regulation of transcriptional repressors in mouse embryonic stem cells. Nat. Struct. Mol. Biol. 15, 259–267 (2008).
Henshall, D. C. et al. MicroRNAs in epilepsy: pathophysiology and clinical utility. Lancet Neurol. 15, 1368–1376 (2016).
Kan, A. A. et al. Genome-wide microRNA profiling of human temporal lobe epilepsy identifies modulators of the immune response. Cell. Mol. Life Sci. 69, 3127–3145 (2012).
Raoof, R. et al. Dual-center, dual-platform microRNA profiling identifies potential plasma biomarkers of adult temporal lobe epilepsy. EBioMedicine 38, 127–141 (2018).
Peron, A. et al. Phenotypes in adult patients with Rett syndrome: results of a 13-year experience and insights into healthcare transition. J. Med. Genet. 59, 39–45 (2022).
Guerrini, R. & Parrini, E. Epilepsy in Rett syndrome, and CDKL5- and FOXG1-gene-related encephalopathies. Epilepsia 53, 2067–2078 (2012).
Wong, L.-C. et al. FOXG1-related syndrome: from clinical to molecular genetics and pathogenic mechanisms. Int. J. Mol. Sci. 20, 4176 (2019).
Miyoshi, G. et al. FoxG1 regulates the formation of cortical GABAergic circuit during an early postnatal critical period resulting in autism spectrum disorder-like phenotypes. Nat. Commun. 12, 3773 (2021).
Weise, S. C. et al. FOXG1 regulates PRKAR2B transcriptionally and posttranscriptionally via miR200 in the adult hippocampus. Mol. Neurobiol. 56, 5188–5201 (2019).
Testa, G. et al. A triheptanoin-supplemented diet rescues hippocampal hyperexcitability and seizure susceptibility in Foxg1+/− mice. Neuropharmacology 148, 305–310 (2019).
Testa, G. et al. Cortical seizures in Foxg1+/− mice are accompanied by Akt/S6 overactivation, excitation/inhibition imbalance and impaired synaptic transmission. Int. J. Mol. Sci. 20, 4127 (2019).
Patriarchi, T. et al. Imbalance of excitatory/inhibitory synaptic protein expression in iPSC-derived neurons from FOXG1+/− patients and in Foxg1+/− mice. Eur. J. Hum. Genet. 24, 871–880 (2016).
Avansini, S. H. et al. Dysregulation of NEUROG2 plays a key role in focal cortical dysplasia. Ann. Neurol. 83, 623–635 (2018).
Leventer, R. J., Guerrini, R. & Dobyns, W. B. Malformations of cortical development and epilepsy. Dialogues Clin. Neurosci. 10, 47–62 (2008).
Blümcke, I. et al. Toward a better definition of focal cortical dysplasia: an iterative histopathological and genetic agreement trial. Epilepsia 62, 1416–1428 (2021).
Heng, J. I. T. et al. Neurogenin 2 controls cortical neuron migration through regulation of Rnd2. Nature 455, 114–118 (2008).
Noack, F. et al. Multimodal profiling of the transcriptional regulatory landscape of the developing mouse cortex identifies Neurog2 as a key epigenome remodeler. Nat. Neurosci. 25, 154–167 (2022).
Schuurmans, C. et al. Sequential phases of cortical specification involve neurogenin-dependent and -independent pathways. EMBO J. 23, 2892–2902 (2004).
Sun, Y. et al. Neurogenin promotes neurogenesis and inhibits glial differentiation by independent mechanisms. Cell 104, 365–376 (2001).
Kessaris, N., Pringle, N. & Richardson, W. D. Ventral neurogenesis and the neuron-glial switch. Neuron 31, 677–680 (2001).
Lamparello, P. et al. Developmental lineage of cell types in cortical dysplasia with balloon cells. Brain 130, 2267–2276 (2007).
Bertrand, N., Castro, D. S. & Guillemot, F. Proneural genes and the specification of neural cell types. Nat. Rev. Neurosci. 3, 517–530 (2002).
Mizuguchi, R. et al. Combinatorial roles of Olig2 and Neurogenin2 in the coordinated induction of pan-neuronal and subtype-specific properties of motoneurons. Neuron 31, 757–771 (2001).
Ge, W. et al. Coupling of cell migration with neurogenesis by proneural bHLH factors. Proc. Natl Acad. Sci. USA 103, 1319–1324 (2006).
Cepeda, C. et al. Epileptogenesis in pediatric cortical dysplasia: the dysmature cerebral developmental hypothesis. Epilepsy Behav. 9, 219–235 (2006).
Heasman, S. J. & Ridley, A. J. Mammalian Rho GTPases: new insights into their functions from in vivo studies. Nat. Rev. Mol. Cell Biol. 9, 690–701 (2008).
Hand, R. et al. Phosphorylation of Neurogenin2 specifies the migration properties and the dendritic morphology of pyramidal neurons in the neocortex. Neuron 48, 45–62 (2005).
Clapier, C. R. & Cairns, B. R. The biology of chromatin remodeling complexes. Annu. Rev. Biochem. 78, 273–304 (2009).
Timpano, S. & Picketts, D. J. Neurodevelopmental disorders caused by defective chromatin remodeling: phenotypic complexity is highlighted by a review of ATRX function. Front. Genet. 11, 885 (2020).
Gibbons, R. J., Picketts, D. J., Villard, L. & Higgs, D. R. Mutations in a putative global transcriptional regulator cause X-linked mental retardation with α-thalassemia (ATR-X syndrome). Cell 80, 837–845 (1995).
Weatherall, D. J. et al. Hemoglobin H disease and mental retardation. N. Engl. J. Med. 305, 607–612 (1981).
Gibbons, R. Alpha thalassaemia-mental retardation, X linked. Orphanet J. Rare Dis. 1, 15 (2006).
Villard, L. et al. Splicing mutation in the ATR-X gene can lead to a dysmorphic mental retardation phenotype without α-thalassemia. Am. J. Hum. Genet. 58, 499–505 (1996).
Gibbons, R. J. et al. Mutations in ATRX, encoding a SWI/SNF-like protein, cause diverse changes in the pattern of DNA methylation. Nat. Genet. 24, 368–371 (2000).
Schenkel, L. C. et al. Identification of epigenetic signature associated with alpha thalassemia/mental retardation X-linked syndrome. Epigenetics Chromatin 10, 10 (2017).
Udugama, M. et al. Histone variant H3.3 provides the heterochromatic H3 lysine 9 tri-methylation mark at telomeres. Nucleic Acids Res. 43, 10227 (2015).
Goldberg, A. D. et al. Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell 140, 678–691 (2010).
Voon, H. P. J. et al. ATRX plays a key role in maintaining silencing at interstitial heterochromatic loci and imprinted genes. Cell Rep. 11, 405–418 (2015).
Nan, X. et al. Interaction between chromatin proteins MECP2 and ATRX is disrupted by mutations that cause inherited mental retardation. Proc. Natl Acad. Sci. USA 104, 2709–2714 (2007).
Kernohan, K. D. et al. ATRX partners with cohesin and MeCP2 and contributes to developmental silencing of imprinted genes in the brain. Dev. Cell 18, 191–202 (2010).
Bérubé, N. G. et al. The chromatin-remodeling protein ATRX is critical for neuronal survival during corticogenesis. J. Clin. Invest. 115, 258–267 (2005).
Seah, C. et al. Neuronal death resulting from targeted disruption of the Snf2 protein ATRX is mediated by p53. J. Neurosci. 28, 12570–12580 (2008).
Watson, L. A. et al. Atrx deficiency induces telomere dysfunction, endocrine defects, and reduced life span. J. Clin. Invest. 123, 2049–2063 (2013).
Shioda, N. et al. Aberrant calcium/calmodulin-dependent protein kinase II (CaMKII) activity is associated with abnormal dendritic spine morphology in the ATRX mutant mouse brain. J. Neurosci. 31, 346–358 (2011).
Nogami, T. et al. Reduced expression of the ATRX gene, a chromatin-remodeling factor, causes hippocampal dysfunction in mice. Hippocampus 21, 678–687 (2011).
Gugustea, R., Tamming, R. J., Martin-Kenny, N., Bérubé, N. G. & Leung, L. S. Inactivation of ATRX in forebrain excitatory neurons affects hippocampal synaptic plasticity. Hippocampus 30, 565–581 (2020).
Tamming, R. J. et al. Mosaic expression of Atrx in the mouse central nervous system causes memory deficits. Dis. Model. Mech. 10, 119–126 (2017).
Gibbons, R. J. et al. Mutations in the chromatin-associated protein ATRX. Hum. Mutat. 29, 796–802 (2008).
Cardoso, C. et al. ATR-X mutations cause impaired nuclear location and altered DNA binding properties of the XNP/ATR-X protein. J. Med. Genet. 37, 746–751 (2000).
McDowell, T. L. et al. Localization of a putative transcriptional regulator (ATRX) at pericentromeric heterochromatin and the short arms of acrocentric chromosomes. Proc. Natl Acad. Sci. USA 96, 13983–13988 (1999).
Howard, M. T. et al. Attenuation of an amino-terminal premature stop codon mutation in the ATRX gene by an alternative mode of translational initiation. J. Med. Genet. 41, 951–956 (2004).
Bérubé, N. G. et al. Patient mutations alter ATRX targeting to PML nuclear bodies. Eur. J. Hum. Genet. 16, 192–201 (2008).
Rauch, A. et al. Range of genetic mutations associated with severe non-syndromic sporadic intellectual disability: an exome sequencing study. Lancet 380, 1674–1682 (2012).
Carvill, G. L. et al. Targeted resequencing in epileptic encephalopathies identifies de novo mutations in CHD2 and SYNGAP1. Nat. Genet. 45, 825–830 (2013).
Suls, A. et al. De novo loss-of-function mutations in CHD2 cause a fever-sensitive myoclonic epileptic encephalopathy sharing features with Dravet syndrome. Am. J. Hum. Genet. 93, 967–975 (2013).
Thomas, R. H. et al. CHD2 myoclonic encephalopathy is frequently associated with self-induced seizures. Neurology 84, 951–958 (2015).
Carvill, G. L. & Mefford, H. C. CHD2-Related Neurodevelopmental Disorders (GeneReviews, 1993).
Woodage, T., Basrai, M. A., Baxevanis, A. D., Hieter, P. & Collins, F. S. Characterization of the CHD family of proteins. Proc. Natl Acad. Sci. USA 94, 11472–11477 (1997).
Wilson, M. M., Henshall, D. C., Byrne, S. M. & Brennan, G. P. Chd2-related CNS pathologies. Int. J. Mol. Sci. 22, 588 (2021).
Lamar, K.-M. J. & Carvill, G. L. Chromatin remodeling proteins in epilepsy: lessons from CHD2-associated epilepsy. Front. Mol. Neurosci. 11, 208 (2018).
Flanagan, J. F. et al. Double chromodomains cooperate to recognize the methylated histone H3 tail. Nature 438, 1181–1185 (2005).
Luijsterburg, M. S. et al. PARP1 links CHD2-mediated chromatin expansion and H3.3 deposition to DNA repair by non-homologous end-joining. Mol. Cell 61, 547–562 (2016).
Kim, Y. J. et al. Chd2 is necessary for neural circuit development and long-term memory. Neuron 100, 1180–1193.e6 (2018).
Aref-Eshghi, E. et al. Evaluation of DNA methylation episignatures for diagnosis and phenotype correlations in 42 Mendelian neurodevelopmental disorders. Am. J. Hum. Genet. 108, 1161–1163 (2021).
Balestrini, S. et al. Real-life survey of pitfalls and successes of precision medicine in genetic epilepsies. J. Neurol. Neurosurg. Psychiatry 92, 1044–1052 (2021).
Beltrán-Corbellini, Á. et al. Epilepsy genetics and precision medicine in adults: a new landscape for developmental and epileptic encephalopathies. Front. Neurol. 13, 777115 (2022).
Younus, I. & Reddy, D. S. Epigenetic interventions for epileptogenesis: a new frontier for curing epilepsy. Pharmacol. Ther. 177, 108–122 (2017).
Cheng, Y. et al. Targeting epigenetic regulators for cancer therapy: mechanisms and advances in clinical trials. Signal Transduct. Target. Ther. 4, 62 (2019).
Deutsch, S. I. et al. Sodium butyrate, an epigenetic interventional strategy, attenuates a stress-induced alteration of MK-801’s pharmacologic action. Eur. Neuropsychopharmacol. 18, 565–568 (2008).
Deutsch, S. I., Mastropaolo, J., Burket, J. A. & Rosse, R. B. An epigenetic intervention interacts with genetic strain differences to modulate the stress-induced reduction of flurazepam’s antiseizure efficacy in the mouse. Eur. Neuropsychopharmacol. 19, 398–401 (2009).
Liu, X. S. et al. Editing DNA methylation in the mammalian genome. Cell 167, 233–247.e17 (2016).
Reschke, C. R. et al. Systemic delivery of antagomirs during blood-brain barrier disruption is disease-modifying in experimental epilepsy. Mol. Ther. 29, 2041–2052 (2021).
Campbell, A. et al. Antagomir-mediated suppression of microRNA-134 reduces kainic acid-induced seizures in immature mice. Sci. Rep. 11, 340 (2021).
Mueller, C. et al. SOD1 suppression with adeno-associated virus and microRNA in familial ALS. N. Engl. J. Med. 383, 151–158 (2020).
Esrick, E. B. et al. Post-transcriptional genetic silencing of BCL11A to treat sickle cell disease. N. Engl. J. Med. 384, 205–215 (2021).
International, T., Against, L., Consortium, E. & Epilepsies, C. Genome-wide mega-analysis identifies 16 loci and highlights diverse biological mechanisms in the common epilepsies. Nat. Commun. 9, 5269 (2018).
Enright, N., Simonato, M. & Henshall, D. C. Discovery and validation of blood microRNAs as molecular biomarkers of epilepsy: ways to close current knowledge gaps. Epilepsia Open. 3, 427–436 (2018).
Kaur, S. & Christodoulou, J. MECP2 Disorders (GeneReviews, 1993).
Van Esch, H. MECP2 Duplication Syndrome (GeneReviews, 1993).
Mullegama, S. V, Mendoza-Londono, R. & Elsea, S. H. MBD5 Haploinsufficiency (GeneReviews, 1993).
Myers, K. A. et al. Phenotypic spectrum of seizure disorders in MBD5-associated neurodevelopmental disorder. Neurol. Genet. 7, e579 (2021).
Ichino, L. et al. MBD5 and MBD6 couple DNA methylation to gene silencing through the J-domain protein SILENZIO. Science 372, 1434–1439 (2021).
Velmans, C. et al. O’Donnell-Luria-Rodan syndrome: description of a second multinational cohort and refinement of the phenotypic spectrum. J. Med. Genet. 59, 697–705 (2022).
Koolen, D. A., Morgan, A. & de Vries, B. B. Koolen-de Vries Syndrome (GeneReviews, 1993).
Fichera, M. et al. Mutations in ACTL6B, coding for a subunit of the neuron-specific chromatin remodeling complex nBAF, cause early onset severe developmental and epileptic encephalopathy with brain hypomyelination and cerebellar atrophy. Hum. Genet. 138, 187–198 (2019).
Bell, S. et al. Mutations in ACTL6B cause neurodevelopmental deficits and epilepsy and lead to loss of dendrites in human neurons. Am. J. Hum. Genet. 104, 815–834 (2019).
Stevenson, R. E. Alpha-Thalassemia X-Linked Intellectual Disability Syndrome (GeneReviews, 1993).
Cossée, M. et al. ARX polyalanine expansions are highly implicated in familial cases of mental retardation with infantile epilepsy and/or hand dystonia. Am. J. Med. Genet. A 155A, 98–105 (2011).
Chatron, N. et al. The epilepsy phenotypic spectrum associated with a recurrent CUX2 variant. Ann. Neurol. 83, 926–934 (2018).
Zarate, Y. A., Kaylor, J. & Fish, J. SATB2-Associated Syndrome (GeneReviews, 1993).
Sweetser, D. A. et al. Pitt-Hopkins Syndrome (GeneReviews, 1993).
Reijnders, M. R. et al. PURA-Related Neurodevelopmental Disorders (GeneReviews, 1993).
Symonds, J. D. et al. Heterozygous truncation mutations of the SMC1A gene cause a severe early onset epilepsy with cluster seizures in females: detailed phenotyping of 10 new cases. Epilepsia 58, 565–575 (2017).
Cooley Coleman, J. A. et al. Comprehensive investigation of the phenotype of MEF2C-related disorders in human patients: a systematic review. Am. J. Med. Genet. A 185, 3884–3894 (2021).
Schoch, K. et al. A recurrent de novo variant in NACC1 causes a syndrome characterized by infantile epilepsy, cataracts, and profound developmental delay. Am. J. Hum. Genet. 100, 343–351 (2017).
Acknowledgements
The authors thank G. Cavalleri for helpful advice on genome-wide association studies. The authors thank colleagues and members of the Neurobiology Commission of the International League Against Epilepsy. The authors gratefully acknowledge the following funders: Deutsche Forschungsgemeinschaft (grant no. FOR 2715), a research grant from Science Foundation Ireland (grant no. 16/RC/3948) co-funded under the European Regional Development Fund and by FutureNeuro industry partners, and EPICLUSTER, which is supported by the European Brain Research Area (EBRA) project. EBRA has received funding from the European Union’s Horizon 2020 research and innovation programme under grant agreement no. 825348.
Author information
Authors and Affiliations
Contributions
All authors researched data for the article, contributed substantially to discussion of the content and reviewed and/or edited the manuscript before submission. K.M.J.V.L., G.L.C., A.J.B., A.M.G., K.K., I.L.-C., C.A.R., E.A.v.V. and D.C.H. wrote the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Neurology thanks S. Balestrini, F. Lubin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Review criteria
Genes were selected on the basis of PubMed searches using the terms “epilepsy” AND “epigenetic”, and the searches were combined with the terms “mutation” or “variant”. The genes and their pathogenic variants were prioritized according to relevance. Epigenetic genes that have been implicated in the severe, early-onset paediatric epilepsies, in which gene discovery has been the most robust and effective in establishing bona fide causative genetic variants for these conditions, were included. An example of how epigenetic processes might be perturbed more broadly in epilepsy, including in the more common focal epilepsies, was also included, using the example of the miR-34a–NEUROG2 cascade that has been implicated in focal cortical dysplasia.
Glossary
- Polygenic risk scores
-
The cumulative risk assessment for an individual to develop a particular medical condition, based on the collective influence of multiple genetic variants.
- X chromosome inactivation
-
(XCI). The X chromosome is gene-rich and, as females have two X chromosomes and males have only one, potential discrepancies in gene dosage between the sexes are addressed through inactivation of one of the X chromosomes. This process occurs randomly, and all daughter cells will have the same X chromosome inactivated.
- Histone variant exchange
-
Histone variants confer different structural properties to the nucleosome. Exchange of these histone variants can promote or weaken nucleosome stability and/or permit more or less DNA to be wrapped around the nucleosome.
- Chromodomain
-
A functional protein domain commonly found in chromatin remodellers and other proteins that associate with chromatin.
- Telomeric
-
Towards the telomeres — the very ends of the linear chromosome.
- Pericentromeric
-
The centromere is the region of the chromosome to which the microtubules of the mitotic spindle are attached during cell division. Pericentromeric regions lie either side of the centromere.
- Chromocentres
-
Large heterochromatic regions of densely packed DNA, mostly satellite DNA and other repetitive regions, as well as histone proteins.
Rights and permissions
About this article
Cite this article
Van Loo, K.M.J., Carvill, G.L., Becker, A.J. et al. Epigenetic genes and epilepsy — emerging mechanisms and clinical applications. Nat Rev Neurol 18, 530–543 (2022). https://doi.org/10.1038/s41582-022-00693-y
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41582-022-00693-y
This article is cited by
-
Chromatin modifiers in human disease: from functional roles to regulatory mechanisms
Molecular Biomedicine (2024)
-
The 100 most-cited manuscripts in epilepsy epigenetics: a bibliometric analysis
Child's Nervous System (2023)
-
Further advances in epilepsy
Journal of Neurology (2023)