Abstract
The evolution of the obligate human pathogen Neisseria gonorrhoeae has been shaped by selective pressures from diverse host niche environments and antibiotics. The varying prevalence of antibiotic resistance across N. gonorrhoeae lineages suggests that underlying metabolic differences may influence the likelihood of acquisition of specific resistance mutations. We hypothesized that the requirement for supplemental CO2, present in approximately half of isolates, reflects one such example of metabolic variation. Here, using a genome-wide association study and experimental investigations, we show that CO2 dependence is attributable to a single substitution in a β-carbonic anhydrase, CanB. CanB19E is necessary and sufficient for growth in the absence of CO2, and the hypomorphic CanB19G variant confers CO2 dependence. Furthermore, ciprofloxacin resistance is correlated with CanB19G in clinical isolates, and the presence of CanB19G increases the likelihood of acquisition of ciprofloxacin resistance. Together, our results suggest that metabolic variation has affected the acquisition of fluoroquinolone resistance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data sets generated during and/or analysed in this study can be found in the source data associated with this work. RNA-seq reads are available publicly at SRA Bioproject PRJNA869861. The NCCP11945 genome is accessible via RefSeq accession NC_011035.1. Genomic data, metadata and accession numbers are publicly available in previously published work69. Source data are provided with this paper.
Code availability
Code used in this study has been published69.
References
Rowley, J. et al. Chlamydia, gonorrhoea, trichomoniasis and syphilis: global prevalence and incidence estimates, 2016. Bull. World Health Organ. 97, 548–562P (2019).
Marazzo, J. M. & Apicella, M. A. in Principles and Practice of Infectious Disease (eds. Bennett, J. E. et al.) Ch. 214 (Saunders, 2015).
Ma, K. C. et al. Adaptation to the cervical environment is associated with increased antibiotic susceptibility in Neisseria gonorrhoeae. Nat. Commun. 11, 4126 (2020).
Lewis, L. A. & Ram, S. Complement interactions with the pathogenic Neisseriae: clinical features, deficiency states, and evasion mechanisms. FEBS Lett. 594, 2670–2694 (2020).
Golparian, D. et al. Genomic evolution of Neisseria gonorrhoeae since the preantibiotic era (1928-2013): antimicrobial use/misuse selects for resistance and drives evolution. BMC Genomics 21, 116 (2020).
Gransden, W. R., Warren, C. A., Phillips, I., Hodges, M. & Barlow, D. Decreased susceptibility of Neisseria gonorrhoeae to ciprofloxacin. Lancet (Lond., Engl.) 335, 51 (1990).
Unemo, M. & Shafer, W. M. Antimicrobial resistance in Neisseria gonorrhoeae in the 21st century: past, evolution, and future. Clin. Microbiol. Rev. 27, 587–613 (2014).
Sexually Transmitted Disease Surveillance – 2020 (CDC, 2020).
WHO Gonococcal AMR Surveillance Programme. WHO https://www.who.int/data/gho/data/themes/topics/who-gonococcal-amr-surveillance-programme-who-gasp (2022).
Rubin, D. H. F., Ross, J. D. C. & Grad, Y. H. The frontiers of addressing antibiotic resistance in Neisseria gonorrhoeae. Transl. Res.https://doi.org/10.1016/j.trsl.2020.02.002 (2020).
Ito, M. et al. Emergence and spread of Neisseria gonorrhoeae clinical isolates harboring mosaic-like structure of penicillin-binding protein 2 in Central Japan. Antimicrobial Agents Chemother. 49, 137–143 (2005).
Sánchez-Busó, L. et al. The impact of antimicrobials on gonococcal evolution. Nat. Microbiol. 4, 1941–1950 (2019).
Mortimer, T. D. et al. The distribution and spread of susceptible and resistant Neisseria gonorrhoeae across demographic groups in a major metropolitan center. Clin. Infect. Dis.: Off. Publ. Infect. Dis. Soc. Am. 73, e3146–e3155 (2021).
Williamson, D. A. et al. Bridging of Neisseria gonorrhoeae lineages across sexual networks in the HIV pre-exposure prophylaxis era. Nat. Commun. 10, 3988 (2019).
Lopatkin, A. J. et al. Clinically relevant mutations in core metabolic genes confer antibiotic resistance. Sci. (N. Y., N. Y.) 371, eaba0862 (2021).
Pinheiro, F., Warsi, O., Andersson, D. I. & Lässig, M. Metabolic fitness landscapes predict the evolution of antibiotic resistance. Nat. Ecol. Evol. 5, 677–687 (2021).
Spence, J. M., Wright, L. & Clark, V. L. Laboratory maintenance of Neisseria gonorrhoeae. Curr. Protoc. Microbiol. 4, Unit 4A.1 (2008); https://doi.org/10.1002/9780471729259.mc04a01s8
Platt, D. J. Carbon dioxide requirement of Neisseria gonorrhoeae growing on a solid medium. J. Clin. Microbiol 4, 129–132 (1976).
Huang, S. et al. Crystal structure of carbonic anhydrase from Neisseria gonorrhoeae and its complex with the inhibitor acetazolamide. J. Mol. Biol. 283, 301–310 (1998).
Elleby, B., Chirica, L. C., Tu, C., Zeppezauer, M. & Lindskog, S. Characterization of carbonic anhydrase from Neisseria gonorrhoeae. Eur. J. Biochem 268, 1613–1619 (2001).
Hewitt, C. S. et al. Structure-activity relationship studies of acetazolamide-based carbonic anhydrase inhibitors with activity against Neisseria gonorrhoeae. ACS Infect. Dis. 7, 1969–1984 (2021).
Remmele, C. W. et al. Transcriptional landscape and essential genes of Neisseria gonorrhoeae. Nucleic Acid Res. 42, 10579–10595 (2014).
Merlin, C., Masters, M., McAteer, S. & Coulson, A. Why is carbonic anhydrase essential to Escherichia coli? J. Bacteriol. 185, 6415–6424 (2003).
Teufel, F. et al. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01156-3 (2022).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Burghout, P. et al. A single amino acid substitution in the MurF UDP-MurNAc-pentapeptide synthetase renders Streptococcus pneumoniae dependent on CO2 and temperature. Mol. Microbiol. 89, 494–506 (2013).
Dillard, J. P. Genetic manipulation of Neisseria gonorrhoeae. Curr. Protoc. Microbiol. 23, Unit 4A.2 (2011); https://doi.org/10.1002/9780471729259.mc04a02s23
Tuttle, D. M. & Scherp, H. W. Studies on the carbon dioxide requirement of Neisseria meningitidis. J. Bacteriol. 64, 171–182 (1952).
Fan, S. H. et al. MpsAB is important for Staphylococcus aureus virulence and growth at atmospheric CO(2) levels. Nat. Commun. 10, 3627 (2019).
Linhares, I. M., Summers, P. R., Larsen, B., Giraldo, P. C. & Witkin, S. S. Contemporary perspectives on vaginal pH and lactobacilli. Am. J. Obstet. Gynecol. 204, 120.e121–125 (2011).
Muir, A. et al. Construction of a complete set of Neisseria meningitidis mutants and its use for the phenotypic profiling of this human pathogen. Nat. Commun. 11, 5541 (2020).
Kampmeier, R. H. Introduction of sulfonamide therapy for gonorrhea. Sex. Trans. Dis. 10, 81–84 (1983).
Zampieri, M. et al. Metabolic constraints on the evolution of antibiotic resistance. Mol. Syst. Biol. 13, 917 (2017).
Yao, J., Bruhn, D. F., Frank, M. W., Lee, R. E. & Rock, C. O. Activation of exogenous fatty acids to acyl-acyl carrier protein cannot bypass FabI inhibition in Neisseria. J. Biol. Chem. 291, 171–181 (2016).
Kitzenberg, D. A. et al. Adenosine awakens metabolism to enhance growth-independent killing of tolerant and persister bacteria across multiple classes of antibiotics. mBio 13, e0048022 (2022).
Firestine, S. M., Poon, S. W., Mueller, E. J., Stubbe, J. & Davisson, V. J. Reactions catalyzed by 5-aminoimidazole ribonucleotide carboxylases from Escherichia coli and Gallus gallus: a case for divergent catalytic mechanisms. Biochem. 33, 11927–11934 (1994).
Andersson, D. I. & Hughes, D. Antibiotic resistance and its cost: is it possible to reverse resistance? Nat. Rev. Microbiol. 8, 260–271 (2010).
MacLean, R. C. & San Millan, A. The evolution of antibiotic resistance. Sci. (N. Y., N. Y.) 365, 1082–1083 (2019).
Sousa, A., Magalhães, S. & Gordo, I. Cost of antibiotic resistance and the geometry of adaptation. Mol. Biol. Evol. 29, 1417–1428 (2012).
Sommer, M. O. A., Munck, C., Toft-Kehler, R. V. & Andersson, D. I. Prediction of antibiotic resistance: time for a new preclinical paradigm? Nat. Rev. Microbiol. 15, 689–696 (2017).
Vincent, L. R. et al. In vivo-selected compensatory mutations restore the fitness cost of mosaic pena alleles that confer ceftriaxone resistance in Neisseria gonorrhoeae. mBio 9, e01905–e01917 (2018).
Kohler, P. L., Hamilton, H. L., Cloud-Hansen, K. & Dillard, J. P. AtlA functions as a peptidoglycan lytic transglycosylase in the Neisseria gonorrhoeae type IV secretion system. J. Bacteriol. 189, 5421–5428 (2007).
Ramsey, M. E., Hackett, K. T., Kotha, C. & Dillard, J. P. New complementation constructs for inducible and constitutive gene expression in Neisseria gonorrhoeae and Neisseria meningitidis. Appl. Environ. Microbiol 78, 3068–3078 (2012).
Norrander, J., Kempe, T. & Messing, J. Construction of improved M13 vectors using oligodeoxynucleotide-directed mutagenesis. Gene 26, 101–106 (1983).
Edwards, R. A., Keller, L. H. & Schifferli, D. M. Improved allelic exchange vectors and their use to analyze 987P fimbria gene expression. Gene 207, 149–157 (1998).
Maness, M. J. & Sparling, P. F. Multiple antibiotic resistance due to a single mutation in Neisseria gonorrhoeae. J. Infect. Dis. 128, 321–330 (1973).
Cohen, M. S. et al. Human experimentation with Neisseria gonorrhoeae: rationale, methods, and implications for the biology of infection and vaccine development. J. Infect. Dis. 169, 532–537 (1994).
Faruki, H. & Sparling, P. F. Genetics of resistance in a non-beta-lactamase-producing gonococcus with relatively high-level penicillin resistance. Antimicrobial. Agents. Chemother. 30, 856–860 (1986).
Johnson, S. R., Steiner, B. M. & Perkins, G. H. Cloning and characterization of the catalase gene of Neisseria gonorrhoeae: use of the gonococcus as a host organism for recombinant DNA. Infect. Immun. 64, 2627–2634 (1996).
Guyer, M. S., Reed, R. R., Steitz, J. A. & Low, K. B. Identification of a sex-factor-affinity site in E. coli as gamma delta. Cold Spring Harb. Symp. Quant. Biol. 45 Pt 1, 135–140 (1981).
Palace, S. G. et al. RNA polymerase mutations cause cephalosporin resistance in clinical Neisseria gonorrhoeae isolates. eLife 9, e51407 (2020).
Kellogg, D. S. Jr., Peacock, W. L. Jr., Deacon, W. E., Brown, L. & Pirkle, D. I. Neisseria gonorrhoeae. I. virulence genetically linked to cloncal variation. J. Bacteriol. 85, 1274–1279 (1963).
Heng, L. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at arXiv.1303.3997 (2013).
García-Alcalde, F. et al. Qualimap: evaluating next-generation sequencing alignment data. Bioinformatics 28, 2678–2679 (2012).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Page, A. J. et al. Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics 31, 3691–3693 (2015).
Thorpe, H. A., Bayliss, S. C., Sheppard, S. K. & Feil, E. J. Piggy: a rapid, large-scale pan-genome analysis tool for intergenic regions in bacteria. Gigascience 7, 1–11 (2018).
Croucher, N. J. et al. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 43, e15 (2015).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–w296 (2021).
Lees, J. A., Galardini, M., Bentley, S. D., Weiser, J. N. & Corander, J. pyseer: a comprehensive tool for microbial pangenome-wide association studies. Bioinformatics 34, 4310–4312 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Stein, D. C., Danaher, R. J. & Cook, T. M. Characterization of a gyrB mutation responsible for low-level nalidixic acid resistance in Neisseria gonorrhoeae. Antimicrobial Agents Chemother. 35, 622–626 (1991).
Bushnell, B., Rood, J. & Singer, E. BBMerge – accurate paired shotgun read merging via overlap. PLoS ONE 12, e0185056 (2017).
Ma, K. C. et al. Increased antibiotic susceptibility in Neisseria gonorrhoeae through adaptation to the cervical environment. Preprint at bioRxiv https://doi.org/10.1101/2020.01.07.896696 (2020).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Wade, J. J. & Graver, M. A. A fully defined, clear and protein-free liquid medium permitting dense growth of Neisseria gonorrhoeae from very low inocula. FEMS Microbiol. Lett. 273, 35–37 (2007).
Mazoyer, A., Drouilhet, R., Despréaux, S. & Ycart, B. flan: an R package for inference on mutation models. R J. 9, 334–351 (2017).
Acknowledgments
We thank the members of the Grad Laboratory and the Waldor Laboratory for their invaluable feedback; the members of the gonococcal subgroup, particularly A. Bandekar, for their invaluable contributions; S. Palace for her thoughts on experimental design; and the Microbial Genome Sequencing Center (https://www.migscenter.com/) and SeqCenter (https://www.seqcenter.com/) for their work on sequencing strains. This work was supported by grants NIH R01 AI132606 and R01 AI153521 and by the Smith Family Foundation Odyssey award (Y.H.G.) and R01 AI 042347-24 (M.K.W.). Authors are further funded by grants NIH F30 AI160911-01 (D.H.F.R.), NIH T32 GM007753 (D.H.F.R.), NIH F31 AI156949-01 (K.H.) and NIH T32 AI132120-01.
Author information
Authors and Affiliations
Contributions
D.H.F.R. and K.C.M. performed the GWAS and statistical analyses. D.H.F.R. and K.A.W. performed the anaerobic experiments. D.H.F.R. and K.H. performed the macrophage experiments. D.H.F.R. performed the remainder of the experimental work. All authors (D.H.F.R., K.C.M., K.A.W., K.H., M.K.W. and Y.H.G.) contributed to data interpretation. Y.H.G. supervised and managed the study. D.H.F.R. and Y.H.G. wrote the manuscript. All authors reviewed and edited the final manuscript. All authors were responsible for the decision to submit for publication.
Corresponding author
Ethics declarations
Competing interests
Y.H.G. is on the scientific advisory board of Day Zero Diagnostics and consulted for GlaxoSmithKline. Y.H.G. has received funding from Merck and Pfizer. None of these competing interests has a bearing on this project. The other authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Jukka Corander, Zeynep Baharoglu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Undirected genetic approaches identify a variant of CanB as causative for CO2-dependence.
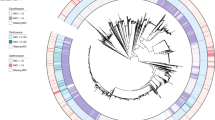
(a) Schematic of undirected transformation of FA19 (parental CanB19G) to identify causative factors for CO2-dependence. (b) Plating efficiency in the presence and absence of CO2 following undirected transformation of gDNA from N. gonorrhoeae strain FA6140 into the N. gonorrhoeae lab strain FA19 (N = 10, 10, 9 from left to right from two independent experiments, error bars represent SEM). Significance (from left to right, p = 0.00018, p = 0.00028) determined by two-sided Mann-Whitney U test. (c) Putative reaction catalyzed by NGO2079/CanB and downstream metabolic products. (d) SNPs present in whole-genome sequencing of two transformants in (b). Arrow indicates the SNP identified in Fig. 1. (e) (Top) Alphafold predicted homodimeric structures of the CanB19E variant (teal) and the CanB19G variant (green). (Bottom) Magnified Alphafold predicted structures overlaying the glutamate (maroon) and glycine (red) at position 19. (f) Proportion of sequenced isolates by year with the CanB19G variant. *p < 0.05, **p < 0.01, ***p < 0.001. Figure created with BioRender.com.
Extended Data Fig. 2 The CanB19E variant has an advantage in the absence of CO2.
(a) Plating efficiency at different CO2 concentrations for N. gonorrhoeae strains 28BL (parental CanB19E) and FA19 (parental CanB19G) with isogenic CanB mutants (N = 6 for 0.1% CO2, N = 3 for air, from two independent experiments, error bars represent SEM). (b) Competition experiment in the presence of CO2 between FA19 CanB19G and FA19 CanBE19G with (left) the FA19 CanB19G strain kanamycin-labeled and (right) the FA19 CanBE19G kanamycin-labeled. The proportion of colony forming units (CFUs) that are kanamycin resistant is graphed against time (N = 3, representative of two independent experiments, error bars represent SEM). (c) Similar to (b), timecourse of competition between isogenic CanB FA19 strains in the presence of media supplementation and the absence of supplemental CO2 (N = 3, representative of two independent experiments, error bars represent SEM). (d) As in (c), competition between unlabeled FA19 CanBG19E and kanamycin-labeled FA19 CanB19G with selected metabolite supplementation assayed at 16 hours in the absence of supplemental CO2 (N = 6, from two independent experiments, error bars represent SEM). (e) Western blot of E. coli MG1655Δcan complemented with IPTG-inducible CanB variants C-terminally FLAG-tagged with G6PD as loading control (representative of two independent experiments).
Extended Data Fig. 3 The 19E variant of NGO2079 is not advantaged at moderately low pH or within macrophages.
The 19E variant of NGO2079 is not advantaged at moderately low pH or within macrophages. (a) Gentamicin intracellular protection assay in RAW 264.7 macrophages as measured by CFUs (N = 3/timepoint, representative of two independent experiments, error bars represent SEM). (b) Competition experiment as in Extended Data Fig. 2b in pH-adjusted Graver-Wade media (N = 3, error bars represent SEM).
Extended Data Fig. 4 CanB19E and CanB19G isogenic pairs have similar MICs.
(a) MICs for sulfamethoxazole, trimethoprim, and trimethoprim/sulfamethoxazole for the FA19 and 28BL pairs isogenic CanB variants. (b) Sulfamethoxazole susceptibility of MG1655Δcan complemented with isogenic CanB variants and induced with IPTG. (c) MICs for clinically relevant antibiotics for FA19 CanB19G and FA19 CanBE19G as determined by E-test and agar dilution plating. (d) gyrA mutations for spontaneous ciprofloxacin escapees or clean mutants of FA19 CanB19G and FA19 CanB19E, along with associated ciprofloxacin MIC.
Extended Data Fig. 5 Loss of CanB does not appear to confer a loss of natural competence.
Loss of CanB does not appear to confer a loss of natural competence. (a) Plating efficiency in the presence and absence of CO2 for a knockout of CanB in the N. gonorrhoeae lab strain FA1090 (N = 6, from two independent experiments, error bars represent SEM). Significance (p = 0.0006) by two-sided Mann-Whitney U. (b) Natural competence of strains of N. gonorrhoeae as measured by uptake of a nalidixic acid resistance-conferring integrating plasmid (N = 3, representative of two independent experiments, error bars represent SEM). (c and d) As in (b), natural competence as measured by (c) uptake of KanR-conferring plasmid DR1 and (d) a PCR product containing the allele encoding for GyrA91F/95G (N = 6, representative of two independent experiments, error bars represent SEM). *p < 0.05, **p < 0.01, ***p < 0.001.
Extended Data Fig. 6 The CanB19G variant is associated with ciprofloxacin resistance across countries and does not lead to a hypermutator phenotype.
(a) Violin plots of drug MICs for ~10,000 N. gonorrhoeae clinical isolates. Black bar represents median, dotted lines represent 25th/75th percentiles. (b) Average log2 of ciprofloxacin MIC in µg/mL by country and by allele of CanB. Weighted average of all datasets is represented by black dots/line. Size of data point represents size of dataset. Error bars represent standard deviation as determined by weighted variance. Significance (p = 0.0120) by unpaired two-sided t-test. (c) Calculated mutation rates of resistance acquisition for to rifampin by fluctuation analysis of isogenic CanB strains. (N = 192, representative of two independent experiments, error bars represent 95% confidence interval). Significance (p = 0.029) by unpaired two-sided t-test (N = 192, representative of two independent experiments, error bars represent SD). (d) Cumulative distribution of mutants in experiments in (c) along with data on the inoculum size and final population size. *p < 0.05, **p < 0.01, ***p < 0.001.
Extended Data Fig. 7 The CanB19G variant does not affect killing by ciprofloxacin, but CanB19G provides an advantage in the presence of gyrA mutations across multiple gyrA alleles and strain backgrounds.
The CanB19G variant does not affect killing by ciprofloxacin, but CanB19G provides an advantage in the presence of gyrA mutations across multiple gyrA alleles and strain backgrounds. (a) Kill curve of isogenic CanB FA19 at 2x MIC and 32x MIC (N = 3, representative of two independent experiments). (b) Doubling time of replicates in Fig. 3e as determined by linear regression on log-transformed CFU/mL counts from 2–10 hours of growth (N = 12, from two independent experiments, error bars represent SEM). Significance (p = 1.9e-5) by unpaired two-sided t-test. (c) Growth of CanB isogenic isogenic 28BL strains bearing ciprofloxacin resistance-determining GyrA91F/95G (N = 3, representative of two independent experiments). Significance determined by unpaired two-sided t-test. (d) Competition between spontaneously ciprofloxacin resistant isogenic CanB strains (see Extended Data Fig. 2) and susceptible parental strains (N = 3, representative of two independent experiments). (e) Growth curves of CanB isogenic FA19 strains with spontaneous ciprofloxacin resistance-determining gyrA alleles (N = 3, representative of two independent experiments). (f and g) Competition between (f) FA19 CanBG19E and FA19 CanBG19E GyrA91F (N = 6, except UMP and 100 µM Adenosine, N = 4) and (g) FA19 CanB19G and FA19 CanB19G GyrA91F (N = 3) after 16 hours along with media supplementation in liquid GCP-K (representative of two independent experiments, error bars represent SEM). Significance (from left to right, p = 0.0002, p = 5.7e-13, p = 3.86e-5), by one-way ANOVA with Dunnett’s multiple comparisons test. *p < 0.05, **p < 0.01, ***p < 0.001.
Extended Data Fig. 8 The relationship between CanB19E and CO2-dependence depends on the acyl-ACP synthetase AasN in the PenA34 lineage of N. gonorrhoeae.
(a) Plating efficiency in the absence and presence of supplemental CO2 of clinical N. gonorrhoeae isolate NY0195 following the introduction of mutations in CanB and AasN (N = 6, from two independent experiments, error bars represent SEM). Significance (from left to right, p = 0.00034 p = 0.00028) by two-sided Mann-Whitney U test. (b) Maximum-likelihood tree as in Fig. 1a overlaid with tracks indicating AasN allele and the ceftriaxone resistance determinant mosaic PenA34 allele. *p < 0.05, **p < 0.01, ***p < 0.001.
Supplementary information
Supplementary Table 1
RNA-seq DESeq2 results.
Supplementary Table 2
Strains, plasmids and primers in this study.
Source data
Source Data Fig. 1
Statistical source sata.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rubin, D.H.F., Ma, K.C., Westervelt, K.A. et al. CanB is a metabolic mediator of antibiotic resistance in Neisseria gonorrhoeae. Nat Microbiol 8, 28–39 (2023). https://doi.org/10.1038/s41564-022-01282-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01282-x
This article is cited by
-
Gut virome and diabetes: discovering links, exploring therapies
Archives of Microbiology (2024)