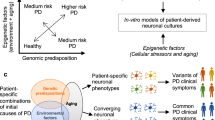
Abstract
We previously reported on two brothers who carry identical compound heterozygous PRKN mutations yet present with significantly different Parkinson’s Disease (PD) clinical phenotypes. Juvenile cases demonstrate that PD is not necessarily an aging-associated disease. Indeed, evidence for a developmental component to PD pathogenesis is accumulating. Thus, we hypothesized that the presence of additional genetic modifiers, including genetic loci relevant to mesencephalic dopamine neuron development, could potentially contribute to the different clinical manifestations of the two brothers. We differentiated human-induced pluripotent stem cells (hiPSCs) derived from the two brothers into mesencephalic neural precursor cells and early postmitotic dopaminergic neurons and performed wholeexome sequencing and transcriptomic and metabolomic analyses. No significant differences in the expression of canonical dopamine neuron differentiation markers were observed. Yet our transcriptomic analysis revealed a significant downregulation of the expression of three neurodevelopmentally relevant cell adhesion molecules, CNTN6, CNTN4 and CHL1, in the cultures of the more severely affected brother. In addition, several HLA genes, known to play a role in neurodevelopment, were differentially regulated. The expression of EN2, a transcription factor crucial for mesencephalic dopamine neuron development, was also differentially regulated. We further identified differences in cellular processes relevant to dopamine metabolism. Lastly, wholeexome sequencing, transcriptomics and metabolomics data all revealed differences in glutathione (GSH) homeostasis, the dysregulation of which has been previously associated with PD. In summary, we identified genetic differences which could potentially, at least partially, contribute to the discordant clinical PD presentation of the two brothers.
Similar content being viewed by others
Introduction
First described over 200 years ago, Parkinson’s Disease (PD) is the second most common neurodegenerative disorder, a progressive disease with a characteristic loss of substantia nigra dopaminergic neurons. PD is believed to result from the interaction of genetic and environmental factors1. To date, more than 20 genes have been associated with increased PD risk and genome wide association studies (GWAS) have further implicated at least 90 risk variants2,3. While the majority of patients suffer from idiopathic PD, meaning no obvious causal genetic variant has been identified, there are indications that at least some forms of idiopathic PD have a complex genetic architecture4. About 5–10% of PD cases are familial and associated with known genetic mutations showing autosomal dominant or recessive inheritance patterns3,5,6.
Mutations in the parkin RBR E3 ubiquitin-protein ligase (PRKN, also known as PARK2) gene are the most common cause of recessively inherited and early-onset PD7,8,9. Over 100 pathogenic PRKN mutations have been described including missense mutations, exon rearrangements and copy number variations10,11,12,13,14. Age of onset and disease progression in patients with PRKN mutations can vary greatly10,15,16,17 and the marked variability of clinical phenotypes, even among family members carrying the same PRKN mutations, suggests that additional factors play a role in disease pathogenesis17,18,19,20.
The occurrence of juvenile cases demonstrates that PD is not necessarily an aging-associated disease. Indeed, evidence is accumulating that there is a developmental component to PD pathogenesis. Minor developmental defects could result in changes in the number of dopamine neurons, their connectivity, insult tolerance or compensatory mechanisms of the dopaminergic circuitry21,22,23,24,25. The developmental hypothesis is supported not only by the occurrence of juvenile PD cases, but also by the examination of PD postmortem brain samples26,27,28 and studies in animal models22,25,29,30,31,32. Indeed, developmental defects relevant to PD have been described for human neuroprogenitor cells as well as postmitotic neurons4,21,26,33,34.
We have previously reported on two brothers (PM and SM) of European ancestry who carry identical compound heterozygous PRKN mutations but present with significant phenotypic discordance with respect to PD onset, disease progression, and clinical symptoms20. The older brother, PM, presented with intermittent bilateral upper extremity action tremors by age 18 and developed marked parkinsonism in his mid-20s. His younger brother, SM, presented with minimal hypomimia and exercise-induced foot dystonia at age 39, but when examined at age 47 he did not meet the UK Brain Bank Criteria for PD35. This difference in clinical presentation between these two brothers is interesting given that they carry identical deletions in PRKN, grew up in the same household, and have spent most of their lives in the same geographical area, suggesting similar exposure to potential environmental risk factors. The divergence in their clinical presentation led us to hypothesize the presence of neurodevelopmentally relevant genetic differences that could potentially play a role in the divergence of disease onset and progression in these two brothers. To test our hypothesis, we compared whole exome sequencing, transcriptomic data, and metabolomic profiling of mitotically active mesencephalic neural precursor cells (floor plate cells) and early postmitotic dopaminergic neurons differentiated from human induced pluripotent stem cells (hiPSCs) from the two brothers.
Results
Validation and characterization of PM and SM hiPSCs
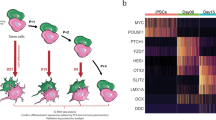
We have previously reported on two brothers (PM and SM) who carry identical compound heterozygous mutations in PRKN, a 40 base pair deletion in exon 3 (rs771529549, p.P113fs) and a deletion encompassing exons 5 and 6 (p.G179_R245del), both predicted to result in deleterious frameshift mutations20,36. Here, the precise deletion in exon 3 was confirmed by wholeexome sequencing (WES, Fig. 1a, Table 1). Moreover, the larger deletion in exon 5 and 6 was refined through copy number variant (CNV) analysis. While the precise breakpoints were not established, the minimum size of the deletion encompassing exons 5 and 6 is over 100,000 base pairs (Fig. 1a, Table 1). hiPSC lines were derived from each brother and all lines were validated through karyotyping and by Pluritest (Supplementary Fig. 1)37. Additional RT-qPCR and immunocytochemical analysis demonstrating the expression of pluripotency markers were previously reported36,38.
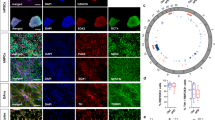
a PRKN gene, transcript variant 1 that encodes a 465 amino acid protein, is shown with the compound heterozygous mutations identified in the brothers: p.P113fs in exon 3 and p.G179_R245del that spans exons 5 and 6. b The expression of β3-tubulin (green) and tyrosine hydroxylase (TH, red) of dopamine neurons differentiated for 27–28 days was assessed by immunocytochemistry. All cultures were counterstained with the nuclear Hoechst stain (light blue). Shown here are cell lines SM15 and PM18, scale bar = 50 µm. c mRNA of dopamine neuron lineage markers expressed by day 11 and day 25 SM (red) and PM (blue) neuronal cultures derived from four SM- and PM hiPSC lines each were quantified by RNA-seq and RT-qPCR. RNA-seq counts were multiplied by a factor of 100 to accommodate logarithmic scaling on the y-axis of the graphs. On day 11 1/4 (PITX3) and 3/4 (TH) of the samples for SM and PM were below the threshold of RT-qPCR detection and are thus not depicted on the graphs. Except for FOXA2 (RNA-seq), the expression levels for all dopamine neuron markers were statistically significantly different between day 11 and day 25, but were not significantly different between SM and PM for RT-qPCR and RNA-seq. Two-way ANOVA followed by Sidak’s multiple comparisons post-hoc test (mean ± SEM; p* < 0.05).
Differentiation of SM and PM hiPSCs into mesencephalic dopamine neurons
hiPSCs were differentiated into mesencephalic dopamine neurons in a two-step process that included a midbrain patterning resulting in day 11 mesencephalic neural precursors (floor plate cells) followed by further differentiation into day 25 early postmitotic mesencephalic dopamine neurons39,40. The differentiations, as assessed by immunocytochemistry were of high efficacy (Fig. 1b, Supplementary Fig. 2)39. Quantification of the expression of additional mesencephalic dopaminergic neuronal markers by RT-qPCR (Fig. 1c), shows decreasing levels for FOXA2 and MSX1, and increasing expression for LMX1A, PITX3 and TH between day 11 and day 25 of differentiation, as previously reported by us and others39,40. The expression of all five markers at both time points were not significantly different between SM and PM lines, suggesting equal efficiency of differentiation to the early mesencephalic lineage for SM and PM hiPSC lines. The data from the RT-qPCR and RNA-seq analysis of dopamine neuron developmental markers correlated significantly (Figs. 1c, 5c).
Genomic variants in genes of interest
Whole exome sequencing (WES) identified variants at 29,732 genomic positions; 20,599 variants were present in both, SM and PM and the remaining 9,133 positions were unique to one of the brothers (PM: 4,271 and SM: 4862, Fig. 2, Supplementary Table 1). Thirty-four of the variants were located within 14 of the established PD-relevant genes (Table 2)3. Of those 34 variants, 21 were identical in SM and PM, including the previously reported c.337_376del frameshift deletion in PRKN (rs771529549). Eight variants were unique to PM and three to SM, all of which were reported to be either benign or tolerated by the ClinVar database, SIFT program or PolyPhen-2 program.
All variants detected for each of the siblings were compared and those unique to SM or PM further evaluated with regards to the type of mutation. The total number of variants that were likely gene disrupting (LGD) for SM (106) and PM (81) are indicated. (heterozy heterozygous, homozy homozygous, hemizy hemizygous).
In addition to variants in established PD-relevant genes, other unique variants being “likely gene-disrupting” (LGD) and predicted to result in stop-gain (nonsense), stop-loss, frameshift, or splicing alterations are highlighted as variants that might have the most significant effect on protein function. PM has 81 unique LGD variants and SM has 106 LGD variants (Fig. 2, Supplementary Table 2). Across both brothers, 39 of the 187 total LGD alterations are within the highly polymorphic HLA genes. Of note, we identified a splicing alteration (rs74853476) in the dopamine β (DBH) gene in PM. This rare variant replaces a nucleotide two base pairs downstream of the first exon of DBH, has a CADD score of 23, and is predicted to be damaging by the MutationTaster program.
Comparison of gene expression of SM and PM dopamine neurons on day 11 and day 25 of differentiation
Levels of gene expression were compared between PM and SM cultures on days 11 and 25 of dopamine neuron differentiation (Fig. 3, Tables 3 and 4, Supplementary Fig. 3). 34 and 31 genes were significantly differentially expressed on days 11 and 25, respectively (FDR < 0.05). 21 of those were differentially expressed at both time points in the same direction, with 13 genes being upregulated and 8 genes downregulated in PM compared to SM. 13 and 10 genes were differentially expressed only on day 11 and day 25, respectively (Fig. 3b, Tables 3 and 4). A heatmap of the differentially expressed genes shows concordance among hiPSC lines derived from the same patient (Fig. 3c).
a Volcano plots illustrate the total number of genes sequenced (black) and genes showing significantly increased or decreased (red) expression levels in PM versus SM (p < 0.01) on day 11 and day 25 of differentiation. The top 10 significant genes are identified on the plots. b Venn diagram indicating that 21 genes are differentially expressed in PM and SM on both days of differentiation, while 13 and 10 genes are differentially expressed on days 11 and 25, respectively (FDR < 0.05) c Heat map of the significantly differentially expressed genes (FDR < 0.05).
Glutathione S-transferase theta 1 (GSTT1) was the most- and glutathione S-transferase mu 1 (GSTM1) was among the most significantly upregulated genes in PM relative to SM cultures on day 11 and day 25, while contactin 4 (CNTN4), contactin 6 (CNTN6) and neural cell adhesion molecule L1-like protein (CHL1) were the most significantly down-regulated genes in PM at both time points (Fig. 3a, Tables 3 and 4).
Each significantly differentially expressed gene identified by RNA-seq was evaluated for potential genomic changes that may be driving its expression levels. Examination of the WES coverage data revealed that SM was homozygous null across the entire length of the GSTT1 and GSTM1 genes, which likely explains the large deficit of expression in SM when compared to PM (Fig. 4a and b). PM has a homozygous deletion encompassing exons 7 and 8 in the guanylate binding protein 3 (GBP3) gene, predicted to result in a frameshift across all transcripts of this gene (Fig. 4c). GBP3 was expressed at significantly lower levels in PM on both, day 11 and day 25 (Tables 3 and 4).
When we overlaid the differentially expressed genes (FDR < 0.05) with the variants from the wholeexome sequencing, 282 variants were identified as being unique to one of the brothers (Supplementary Table 3). For GBP3, we found, in addition to the above-mentioned deletion encompassing exons 7 and 8, three nonsynonymous variants, two of which had CADD scores >10 p.R225W (SM, 0/1) and p.R221Q (SM, 0/1). CHL1, one of the cell adhesion genes significantly downregulated in PM (Tables 3 and 4) showed one nonsynonymous variation in PM but with a very low CADD score (0.003, Supplementary Table 3). The other most significantly down-regulated genes in PM, CNTN4 and CNTN6, showed several different variants. Three of the four CNTN4 variants were distributed among the two brothers (p.P190P, SM 0/1; pN294N, PM1/1, SM 0/1; p.R402R PM 0/1) and had CADD scores between 11–12, while both CNTN6 variants had very low CADD scores (Supplementary Table 3).
The vast majority of variants (224/282) fall in the highly polymorphic HLA genes. HLA-DRB5 was the most differentially expressed HLA gene, with PM showing 69-fold and 335-fold higher expression than SM on days 11 and 25, respectively. We detected a total of 48 different variants in HLA-DRB5, with 16 showing CADD scores > 10; of those, 15 were present in SM and only 1 in PM (Supplementary Table 3). For the other differentially expressed HLA genes, we made the following observations: HLA-B: 9/49 variants have CADD scores >10 and 8/9 of those were present in SM; HLA-DMA: 3/3 variants have CADD scores >10 and 3/3 were present in PM; HLA-DQB1: 11/54 variants have CADD scores >10 and of those 10/11 were present in SM; HLA-DRB1: 10/70 variants had CADD scores >10 and 10/10 of those were present in SM. Overall, for all five of those HLA genes, the individual carrying the majority of variants showed lower expression of the respective HLA gene (Tables 3 and 4 and Supplementary Table 3). In summary, of the 25 genes identified to have brother-specific variants, 19 and 17 genes showed significantly different expression levels between the brothers on day 11 and 25, respectively; of the 16 genes with variants with CADD scores > 10, twelve and eleven genes showed significantly different expression levels between SM and PM on days 11 and 25, respectively.
Using a combination of three in silico predictive programs (SIFT, Polyphen2, MutationTaster) to predict damaging or pathogenic variants, we identified three nonsynonymous variants which had a CADD score greater than 10: GBP3 p.R225W (SM) and ULK4 p.S348G (SM), and MS4A6E p.V47F (heterozygous in PM and homozygous in SM). GBP3 was expressed at significantly higher levels in SM on both day 11 and 25, while ULK4 was expressed significantly higher in SM on day 11 only and MS4A6E was higher in PM on day 25 only (Fig. 5a, b, Tables 3 and 4). Thus, while all three of these alterations are fairly common, with minor allele frequencies in ExAC of 0.12 or higher, they may still potentially be acting as modifiers.
a 13 of the genes identified as differentially expressed in SM and PM by RNA-seq on day 11 and day 25 of differentiation were validated by RT-qPCR. The statistical significance (-log p values) of the differential expression between PM and SM cultures evaluated are graphed. Grey-colored bars lying within the range of non-significant differences as delineated by two orange lines were not significantly differently expressed in PM and SM cultures as assessed by Rt-qPCR (p > 0.05). b Correlation analysis of the expression levels of all genes analyzed by RT-qPCR and RNA-seq including the dopamine-lineage markers (Fig. 1c) across all four SM and PM hiPSC lines are graphed for day 11 and day 25 (Log2(Count) versus Ct; Pearson correlation analysis, two-tailed p-value). RNA-seq counts were multiplied by a factor of 100 to accommodate logarithmic scaling on the x-axis of the graphs (c).
We selected a subset of 13 genes found to be differentially expressed by RNA-seq between PM and SM either on both days (8 genes) or day 11 (2 genes, EN2, ULK4) or day 25 only (3 genes, NOS2, MS4A6E, SLC18A2) and validated the differential expression for all 13 genes by RT-qPCR (Fig. 5a). The RT-qPCR data for all the genes evaluated, including the 5 lineage markers (Fig. 1c) showed a strong correlation with the RNA-seq data (Fig. 5b).
Metabolomics
Our RNA-seq data demonstrates that already on day 11 of differentiation there are important differences in gene expression between SM and PM lines. In order to assess effects downstream of the transcriptional differences in very early stages of dopaminergic differentiation, we performed metabolomics of day 11 cultures. Day 11 cultures are more homogenous than their day 25 counterparts and thus provide a greater opportunity to detect differences in metabolites. Thus, we performed two independent metabolomics analyses of day 11 neural precursor cells (floor plate cells) using two different types of liquid chromatography separation and two different sets of SM and PM hiPSC lines. These analyses revealed significantly different metabolomic profiles between the two brothers (Fig. 6a, d). The relative abundance of metabolites was concordant within different hiPSC lines derived from the same individual (Fig. 6b, e). A similar number of metabolites were elevated or decreased in PM when compared to SM and representative metabolites belonging to pathways altered between the two brothers (see below) are highlighted in the volcano plots (Fig. 6c, f; Supplementary Fig. 4a, b). In particular we observed 4.2 and 1.4-fold lower levels of GSH and GSSG, respectively for PM vs SM, which translates into a 3-fold lower GSH/GSSG ratio for PM as compared to SM floor plate cells (Supplementary Table 4). We further measured 1.6-fold higher level of glutamine in PM than SM cells, similar to observations made in in vivo NMR analysis of the putamen metabolome in PD patients41. We also detected increased levels of creatine in PM, such measurements in PD patients are contradictory42,43.
The metabolome of SM and PM day 11 neural precursors resulting from three separate differentiations (1–3) were analyzed by HILIC positive mode (a–c) and HILIC negative mode (d–f) mass-spectrometry. a. d Principal component analysis (PCA plots) illustrates distinct metabolomic profiles for PM vs SM. b, e Hierarchical clustering maps provide global comparisons for PM vs SM differences of the top 500 significantly different individual metabolite levels (rows) clustered by SM and PM cell lines and differentiations (columns). Metabolites are colored according to relative feature abundance across all samples ranging from low (green) to high (red). c, f Volcano plots illustrate the total number of detected metabolites (gray) and significantly increased (red) or decreased (blue) metabolites in PM vs SM (p < 0.05). A total of 2725 (HILIC pos) and 1,770 (HILIC neg) compounds were detected of which 542 (HILIC pos) and 194 (HILIC neg) are significantly different between PM vs SM. Representative members of the identified pathways (Fig. 7) altered between the two brothers are highlighted. Some of the most significantly changed molecules cannot not be annotated, because there are too many isomers as possible candidate metabolites.
Metabolites present at statistically significantly different levels (p < 0.05) between PM and SM developing dopamine neuron cultures were classified according to their biochemical properties. For the hydrophilic interaction liquid chromatography positive/negative ion mode (HILIC pos/neg) analysis (method #2), the largest percentage of metabolites belonged to the lipid and lipid-like molecules (34%), with the second-largest group (24%) identifying as amino acids, peptides and analogues (Fig. 7a). The same overall distribution was observed for the reverse-phase liquid chromatography (RPLC) pos/HILIC pos analysis (method #1) although, not surprisingly, the lipid and lipid-like molecules were present more prominently (44%), and the amino acids, peptides and analogues contributed to a lesser degree (17%) (Fig. 7b).
a, b Metabolites significantly different between PM and SM were classified according to their biochemical properties and c, d pathway analysis was performed. The top 9 significant response pathways are shown for each data set with a total of 8 pathways being significantly different between SM and PM (p ≤ 0.1 is indicated by the dashed line).
Pathway analysis
Pathway analysis was performed separately for the metabolomic and RNA-Seq data (Figs. 7, 8). The metabolite pathways identified to be different between the two brothers were qualitatively similar between the two chromatography methods (method #1 and #2) but ranked slightly differently. Thus, sphingolipid metabolism was identified as the most significantly different pathway for the HILIC pos/neg analysis (method #2), but did not reach statistical significance for the RPLC pos/HILIC pos analysis (method #1) (Fig. 7c, d). The biosynthesis of aromatic amino acids (phenylalanine, tyrosine and tryptophan) was identified as the most significantly different pathways by RPLC pos/ HILIC pos, but did not reach statistical significance in the HILIC pos/HILIC neg analysis (Fig. 7c, d). RPLC pos/HILIC pos analysis identified glutathione metabolism as significantly different between SM and PM (Fig. 7d). GSH metabolism was also identified in the RNA-seq based GO analysis for Biological Processes and Molecular Functions for day 11 and day 25 cultures (Fig. 8a–f). GO analysis of the RNA-seq data further identified cell adhesion and dendrite self-avoidance, processes which play important roles in neuronal development (Fig. 8a–f). In addition, several pathways and processes relating to immune function were identified that were driven by several HLA genes (Fig. 8a–f). The HLA gene products most likely play a role in neural development independent of their function in cellular immunity (see discussion).
a, b STRING networks with differentially expressed genes on day 11 (a) and day 25 (b). Only those genes connected to a node are shown. c, d Statistically significantly different GO Molecular Functions and e, f GO Biological Processes (FDR < 0.05) determined from the differentially expressed genes for day 11 (c, e) and day 25 (d, f) of differentiation are plotted. Pathways relating to GSH metabolism are indicated in green, neural development in red and all others in blue. The GO Biological Processes labelled in blue involve MHC class II gene expression, genes which have been demonstrated to play a role in neural development independent of their function in cell immunity.
Discussion
The remarkable difference in the clinical presentation with respect to age of onset and severity of symptoms between the brothers SM and PM who carry the same compound heterozygous PRKN mutations and have lived in the same geographical area for the majority of their lives, led to the hypothesis of the presence of additional genetic elements that could potentially play a role in the differential onset and progression of PD in these two patients.
There is increasing evidence that there is a developmental component to PD pathogenesis21,22,23,24,25. Proper brain development requires finely tuned interactions among hundreds of genes and a dysregulation in these networks can lead to neurodevelopmental disorders44. Indeed, many PD-associated genes have been implicated in neuronal developmental processes45,46. We identified a number of differentially expressed genes between SM and PM developing dopamine neurons that are either directly implicated in dopamine neuron development, neural development in general, or are candidate genes for neurodevelopmental functions.
The most disparate gene expression between SM and PM developing dopamine neurons was observed for a group of cell adhesion proteins CNTN6, CNTN4 and CHL1, that were expressed at levels orders of magnitude lower in PM than SM neurons at both stages of differentiation (day 11 and day 25) suggesting persistent differential expression during development (Tables 3 and 4). Intriguingly, CHL1 was previously shown to be differentially regulated in the substantia nigra of brains from PD patients compared to control brains47. Another study using living tissue from frontal lobe biopsies of patients and controls demonstrated a significant difference in the expression of CNTN648. Thus, two of these three genes have been independently verified to be differentially expressed in PD patients. CNTN6, CNTN4 and CHL1 encode neuronal membrane immunoglobulin-like proteins that play a role in axon guidance and outgrowth, orientation of dendrites and synaptogenesis in the developing nervous system49,50,51,52 and are expressed in the developing human embryonic substantia nigra pars compacta (https://www.brainspan.org)51,52,53. Mesencephalic dopamine neurons extend their axons from the substantia nigra into the dorsal striatum where they form extensive ramified axonal branches54,55. Thus, a defect in axon pathfinding or synapse formation could potentially have a significant impact on mesencephalic-striatal circuit formation and connectivity. Variants of each of these three cell adhesion molecules in isolation have been implicated in several different neurodevelopmental disorders49,53,56. We observed a concomitant and rather large difference in the expression of all three of these genes between SM and PM dopamine neuron cultures, an observation that supports our hypothesis of a potential difference in the development of the dopaminergic system between the two brothers.
Differential expression of the gene EN2 in the prefrontal cortex of patients with PD compared to controls has been reported57 and the analysis of human haplotype implicates variations in the engrailed 2 gene (EN2) in the development for early-onset PD58. The development of mesencephalic dopamine neurons is tightly orchestrated by the spatiotemporal expression of highly conserved transcription factors such as engrailed 1 and 2, which are involved in early lineage specification and axon guidance23,59,60,61,62,63. We observed EN2 expression levels in day 11, but not day 25 cells to be 20-fold higher in PM than SM cells. Dysregulation of EN1/2 expression in either direction has been shown to interfere with the development of mesencephalic dopamine neurons61,64,65,66. However, since the differential EN2 expression was only observed on day 11 but not day 25 of differentiation, we cannot exclude the possibility that this higher expression reflects a slightly different time frame of development between SM and PM lines, rather than a significant overall developmental difference.
Human HLA genes are ubiquitously expressed and their developmentally regulated expression throughout the brain including the ventral mesencephalon suggests an immune system unrelated role for these proteins in early neuronal differentiation67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83. Our observation that several of the HLA genes were differentially expressed between in SM and PM (Table 4) so early in dopamine neuron differentiation raises the possibility of potential differences in midbrain development between PM and SM. Enhanced expression of HLA genes has been shown to lead to aberrations in neurodevelopment74. In this context, it is interesting to point out that the majority of the HLA genes differentially expressed on day 11 (4 out of 5) and on day 25 (3 out of 4) were expressed at higher levels in PM than SM cultures. Importantly, three human HLA II genes (HLA-DQB1, HLA-DRB1, and HLA-DRB5) that we observed to be differentially expressed are located within the chromosome 6p21.3 major histocompatibility complex class II cluster and were previously reported to be a top GWAS hit for late-onset PD84,85,86,87,88,89. Moreover, evidence points to the possibility of multiple PD risk-associated variants in this region including both coding and noncoding, regulatory alterations89,90.
We identified variants at a total of 29,732 positions, and of those, 34 were located on 14 established PD-relevant genes several of which carried multiple variants (Table 2). We identified 8 variants that were unique to PM and 3 for SM. Although the large majority of these 34 variants on their own are classified as benign or tolerated, it is not clear how the combination of multiple variants within one gene affects protein levels and/or function. The significance of the presence of multiple variants in these PD-relevant genes for the clinical discordance between PM and SM is difficult to deduce and would require protein expression- and functional studies.
Very striking was the differential expression of GSTM1 and GSTT1, which we demonstrate to be the result of SM being homozygous null for both genes; thus, although the RNA samples analyzed represent a snapshot during early nigral development, the fact that the lack of expression of these two genes are due to null mutations implies that these two enzymes are permanently lacking in SM. Both genes encode glutathione S-transferases, enzymes involved in the protection against xenobiotics and oxidative stress, cellular processes that have been correlated with PD pathogenesis91,92,93,94,95,96. These large-scale deletions are relatively common in white, non-Hispanic populations (~50% for GSTM1 and ~20% for GSTT1)97 but studies on their role in PD pathogenesis are conflicting98,99,100,101,102.
Another interesting variant representing a potential modifier of the discordant clinical presentation is DBH, a gene that encodes the enzyme dopamine beta-hydroxylase (DBH) that converts dopamine to norepinephrine. In this study, a rare splicing variant (rs74853476) was identified in PM that may disrupt the splice donor site adjacent to exon 1. Risk assessments of DBH variants for PD are conflicting103,104,105,106,107,108,109,110. To our knowledge the DBH variant (rs74853476) we observed in PM has not been assessed for its PD risk potential.
Our metabolomics pathway analysis identified three biochemical processes differentially regulated between SM and PM: 1) sphingolipid and glycerophospholipid metabolism, 2) the metabolism of several amino acids including phenylalanine, tyrosine, tryptophan, arginine and proline together with aminoacyl-tRNA biosynthesis and 3) GSH metabolism which was also identified in the RNA-seq-based GO analysis. A decreased GSH/GSSG ratio is an indicator of oxidative stress, a mechanism believed to play a role in PD pathogenesis. We observed a 3-fold lower GSH/GSSG ratio for PM as compared to SM floor plate cells. Very similar observations have been reported for day 25 PRKN-KO hiPSC-derived mesencephalic dopamine neurons111, as well as for PD patient tissue112. The absence of functional parkin causes abnormal mitochondrial morphology and function resulting in increased oxidative stress36,113. Thus, an additional depletion of GSH, the most abundant antioxidant in the CNS, could therefore add additional stress on dopaminergic neurons of patient PM versus SM. Importantly in the context of this study, aberrant GSH metabolism has not only been associated with PD91,96,102 but also serves as an indicator of developmental toxicity114. The metabolism of aromatic amino acids has been shown to be different between PD patients and control subjects in most metabolite profiling studies, the directionality of these differences however, are not always concordant43. Alterations in mitochondrial aminoacyl-tRNA biosynthesis resulting from mutations in mitochondrial aminoacyl-tRNA synthetases cause a variety of pathogenic CNS phenotypes including infantile-onset parkinsonism115,116,117. In addition, a protein complex interacting with aminoacyl-tRNA synthetase, a substrate of parkin, accumulates in parkin-inactivated PD animal models and causes dopamine neuron degeneration118,119. The metabolism of lipids, including glycerophospholipids and sphingolipids, has been reported to be altered in PD patients120,121 and importantly, has been found to play a central role in all mechanisms presently believed to underly PD pathogenesis121,122,123,124,125,126,127. Sphingolipids and glycerophospholipids are major constituents of the plasma membrane and not surprisingly, are important for neural maintenance and development, playing roles in cell-cell recognition, cell adhesion, signal transduction and synapse formation122,128,129,130,131,132,133.
The authors acknowledge that the differences in gene variants, gene transcription and metabolism between SM and PM reported here are not functionally validated in this current study and, thus, do not provide causal evidence for the clinical divergence between the brothers. However, our observations contribute to a basis of PD associated genetic, transcriptional and metabolomic phenotypes that builds the foundation for the establishment of causal relationships between genetic and biochemical cellular phenotypes and clinical observations. Indeed, our and other groups’ reports of PD patient dopamine neurons genetic and biochemical phenotypes which have also been described in PD patients suggests that these human in vitro approaches can serve as models to establish causal relationships between genetic differences and PD pathogenesis.
In summary, we assessed genetic differences that could potentially play a role in the drastically different clinical presentations of two brothers carrying the same compound heterozygous PRKN mutations. Combining exon sequencing, transcriptomics and metabolomics we found differences in gene expression (CNTN4, CNTN6, CHL1, HLA genes) and metabolism (sphingo- and glycerophospholipids) relevant to the development of mesencephalic dopamine neurons. We further observed differences in cellular processes underlying dopamine homeostasis (DBH, tyrosine biosynthesis) and protection against insults from oxidative stress and xenobiotics (GSTM1 and GSTT1, GSH metabolism), cellular functions that have been shown to play a role in PD pathogenesis. Future studies should be aimed at examining cellular consequences of these genetic differences on the development, function and vulnerability of these human dopamine neurons.
Methods
Patient Information
We have previously published a detailed description of the patients PM and SM20. In brief, PM and SM are brothers, aged 6 years apart who share the same compound heterozygous disease-causing mutations in PRKN20. PM displayed significant postural and gait impairment in his mid 20 s and was diagnosed with PD in his mid 30 s. At age 42 he started with bilateral subthalamic nucleus deep brain stimulation and at age 50 obtained additional bilateral deep brain stimulation in the globus pallidus interna. Since age 53 the patient is functionally wheelchair-bound. His six-year younger brother SM developed exercise-induced foot dystonia which was resolved by selegiline. Examined at age 47 he did not meet UK Brain Bank Criteria for PD35.
Derivation, validation and differentiation of hiPSCs
hiPSC lines were derived from dermal fibroblasts from two human subjects, PM and SM, who underwent a dermal biopsy at age 46 and 40, respectively, after providing the appropriate patient consent/assent under the guidelines of an approved Internal Review Board (IRB) protocol at Vanderbilt University (#080369). Four independent hiPSC clones from each patient were used in this study (PM1, PM12, PM17, PM18 and SM3, SM5, SM14 and SM15). Three of the hiPSC lines (PM1, SM3, SM5) were generated by transducing the fibroblasts with a lentivirus as previously described38. The other five lines (PM12, PM17, PM18, SM14, SM15) were reprogrammed by electroporation with CXLE plasmid vectors (Addgene) using the Neon Transfection System (Life Technologies) following published methods39,134. Karyotype analyses were performed for all hiPSC lines using standard protocols with at least 20 metaphase spreads per cell line (Genetics Associates). The lack of plasmid integration into the genomic DNA was demonstrated by RT-qPCR. The pluripotency of all hiPSC lines was validated by Pluritest37 and/or by analyzing the expression of pluripotency markers by immunofluorescence and RT-qPCR. In addition, the capacity of the hiPSC lines to differentiate into cell types belonging to the three germ layers was assessed36,38,135.
Differentiation of the hiPSCs to a mesencephalic dopaminergic lineage was performed as previously described39,40,136. In brief, hiPSCs were differentiated in a first stage into floor plate cells (mesencephalic neural precursors) (days 0–11) via dual SMAD inhibition combined with ventral midbrain patterning. In a second stage, these floor plate cells were further differentiated (days 11–25) into early postmitotic mesencephalic dopamine neurons. The RNA-seq data were derived from two independent dopamine neuron differentiations that were performed with two different sets of SM and PM hiPSC lines each; thus, we analyzed a total of four SM and four PM lines (Supplementary Fig. 3). The metabolomics data which corroborate some of the RNA-seq data were obtained from an additional two independent differentiations. Thus, the data presented in this manuscript is based on four independent dopamine neuron differentiations.
Immunofluorescence
For immunofluorescence analysis, dopamine neurons were plated into 96-well plates (Greiner Bio-One) and immunofluorescence was performed as previously described39. Briefly, the cells were fixed in PBS containing 4% paraformaldehyde (Electron Microscopy Sciences) for 30 min at room temperature, permeabilized with 0.2% Triton X-100 for 20 min at room temperature and then incubated in PBS containing 5% normal donkey serum (Jackson ImmunoResearch) and 0.05% Triton X-100 for 2 h at room temperature or overnight at 4 °C. The following primary antibodies were used: mouse anti-β3-tubulin (Thermo Scientific; MA1-19187, 1:500), rabbit anti-tyrosine hydroxylase (TH) (Pel-Freez; P40101, 1:500) and sheep anti-TH (Pel-Freez; P60101, 1:250). Secondary antibodies conjugated to DyLight 488 (1:800), and DyLight 549 (1:800), both from Jackson ImmunoResearch. Images were obtained with a Zeiss ObserverZ1 microscope and AxioVs40 software (version 4.7.2). For high content imaging images were acquired using a Molecular Device’s ImageXpress Micro XL system and MetaXpress software available at the Vanderbilt Highthroughput Screening Core Facility. Quantification of β3-tubulin- and TH-positive cells was performed in two cultures for each cell line and at least 9000 cells per culture.
DNA extraction and wholeexome sequencing
Whole exome sequencing (WES) was performed on the PM12 and SM14 lines. Briefly, 3 μg of DNA was extracted from hiPSCs using the DNeasy Blood & Tissue Kit (Qiagen) and exonic DNA regions were selected using the Agilent SureSelect Human All Exon 60 Mb v6 kit according to the manufacturer’s instructions. The DNA was sequenced on the Illumina HiSeq 2500 with 100 base pair (bp) paired-end reads. Data were processed according to Genome Analysis Tool-kit (GATK) best practices with variants required to be called in both brothers, to have a GQ score of 99 or above, and were annotated using SeattleSeq137. All variants were compared to a list of 21 established PD genes3. Variants were then prioritized for those that were unique to either brother, “likely gene-disrupting” (LGD), and predicted to result in stop-gain (nonsense), stop-loss, frameshift, or splicing mutations.
Copy number variant (CNV) analysis
DNA isolated from the PM12 and SM14 clones for WES was also used for genotyping with the Illumina 1M-Single array. Samples had average call rates >98%. CNVs were identified using the cnvPartition v.3.2.0 algorithm implemented in Illumina GenomeStudio software. Select genes and regions of interest were also manually evaluated using the pile up data generated from the whole exome sequencing. Manual visualization of WES coverage was also used to identify large, homozygous deletions within a subset of genes that were found to be significantly different in the transcriptomic analysis.
RNA extraction and transcriptomic analysis
Total RNA was extracted from day 11 floor plate cells and day 25 early post-mitotic dopaminergic neurons differentiated from all eight hiPSC lines using the Qiagen RNeasy Mini Kit according to the manufacturer’s protocol. Cell lysates were homogenized using QIAshredder spin columns (Qiagen) and genomic DNA was removed by an on-column DNase treatment (Qiagen). The concentration and quality of RNA samples was assessed using a Bioanalyzer RNA kit (Agilent Technologies) and RNA integrity numbers for all RNA samples ranged between 8.3 to 10. cDNA libraries were generated with the Illumina TruSeq Stranded kit with a poly-A selection. Paired-end 100 base pair read sequencing was performed on indexed samples and run on the HiSeq 2500, yielding a minimum of 25 million reads/sample. Alignment of RNA-seq reads against the human genome (GRCh38) was performed using STAR Aligner and Htseq-count was used to count the number of overlapping reads with genes. Differential expression analysis was conducted using edgeR with a false discovery rate (FDR) cut-off of 0.05 to correct for the multiple comparison testing138.
Quantitative RT-PCR
RT-qPCR and transcriptomic analyses were performed on the same RNA samples. 3 µg of total RNA were used for each sample for reverse transcription (RT) into cDNA using iScript Reverse Transcription Supermix for RT-qPCR (Bio-Rad) according to the manufacturer’s instructions. RT-qPCR reactions were performed in triplicates at 1.5 ng/µl cDNA using SsoAdvanced Universal SYBR Green Supermix (Bio-Rad) at a final volume of 10 µl on the Bio-Rad CFX384 Touch™ Real-Time System in 384-well standard PCR plates. Melting curves for each reaction were visually inspected via Bio-Rad CFX Maestro™ 1.1 Software and the acceptable range of Ct values for each RT-qPCR triplicates was implemented assuming 100% primer efficiency as described139. Relative quantification of genes of interest was conducted using actin as a reference gene and fold change was calculated using the 2^-∆∆Ct method, setting the arithmetic mean ∆Ct value of SM/PM as calibrator. Collectively, for genes with RT-qPCR triplicates showing no amplification signal or a mean Ct value of ≥ 36, a mean Ct value of 36 was imputed for statistical analyses as previously described139. Two-tailed Student’s t-tests were performed to determine statistical significance for each gene with p value < 0.05 as the cut-off for significance and not assuming consistent standard deviation between groups. For the analysis of dopaminergic lineage markers, ordinary two-way ANOVA followed by Sidak’s multiple comparison post-hoc test was conducted using ∆Ct values. In all cases, data were presented as mean ± SEM. Correlation analyses between RNA-seq and RT-qPCR results were performed via Pearson’s correlation analysis.
Metabolomics
Cell harvest and metabolite extraction
Optima grade LC-MS solvents for the mass spectrometry analyses were obtained from Thermo Fisher Scientific (Fair Lawn, NJ). Two separate metabolomics analyses using slightly different methods were performed with different SM and PM hiPSC lines.
For method #1 (RPLC positive ion mode /HILIC positive ion mode) day 11 neural dopaminergic precursor cells (floor plate cells of one well of a 6 well plate) were harvested into 500 µl of ice-cold methanol, flash frozen and then stored at −80 °C. To extract the metabolites, the 500 µl methanol cell suspensions were thawed and 100 µl of H2O added. Then the samples were frozen on dry ice for 3 min, defrosted in ice over a 10 min period, and sonicated with 10 pulses using a probe sonicator at 30% power. The freeze-thaw-sonication sequence was repeated three times. The proteins were precipitated by placing the lysates at −80 °C overnight and then pelleted by centrifugation at 25,000 x g for 15 min. Cleared supernatants containing the metabolites were placed in clean Eppendorf tubes, dried in a vacuum concentrator and stored frozen at −80 °C. For RPLC positive ion mode mass spectrometry analysis the dried extracts were reconstituted in 60 μl of RPLC buffer (acetonitrile/water with 0.1% formic acid, 2:98, v/v). Samples were vortexed rigorously to solubilize the metabolites, cleared by centrifugation for 5 min at 15,000 rpm, and the supernatants were injected twice (5 μl / injection) randomly. Quality control samples were prepared by combining equal volumes (10 μl) of each sample. After RPLC mass-spectrometry, the remaining samples were dried down in vacuo, and the metabolites reconstituted in 40 μl of HILIC buffer (acetonitrile/water, 90:10, v/v) and 5 µl of each sample was injected twice in random sequence to perform HILIC positive ion mode mass-spectrometry.
In method #2 (HILIC positive ion mode/negative ion mode) the cells were washed three times with 2.5 ml of an ammonium formate buffer (50 mM, pH 6.8), scraped into the same buffer, centrifugated at 200 x for 5 min, the cell pellets flash frozen in liquid nitrogen and stored at −80 °C. To extract the metabolites, cell pellets were lysed in 200 µl ice-cold lysis buffer (1:1:2, Acetonitrile:MeOH:Ammonium Bicarbonate 0.1 M, pH 8.0, LC-MS grade) and sonicated once as described above. The protein concentration was determined (BCA assay, Thermo Fisher Scientific) and adjusted to 1 mg/ml. Isotopically labeled standard molecules, Phenylalanine-D8 and Biotin-D2 were added to the 200 µl cell lysates, the protein precipitated by addition of 800 µl of ice-cold methanol and stored at −80 °C overnight. Upon thawing, the precipitated proteins were pelleted by centrifugation at 9300 x g for 10 min, the supernatants transferred into two clean Eppendorf tubes, dried down in vacuo and stored at −80 °C. To perform HILIC-positive ion mode and HILIC-negative ion mode mass spectrometry, each sample was reconstituted in 60 μl of HILIC reconstitution buffer (acetonitrile/water, 90:10, v/v) and 5 µl of each sample was injected once for positive ion mode and 8 µl for negative ion mode. During the final reconstitution, isotopically labeled standard molecules, Tryptophan-D3, Carnitine-D9, Valine-D8, and Inosine-4N15, were added to each sample, and quality control sample was prepared by pooling equal volumes from each individual sample.
Mass spectrometry and data acquisition
UPLC-IM-MS and data-independent acquisition (MSE) were performed on a Waters Synapt G2 HDMS (Milford, MA, USA) mass spectrometer equipped with a Waters nanoACQUITY UPLC system and autosampler (Milford, MA, USA). Metabolites were separated on a reverse phase 1 mm × 100 mm HSS T3 C18 column packed with 1.8-μm particles (Waters, Milford, MA, USA) held at 45 °C. Liquid chromatography was performed using a 30-min gradient at a flow rate of 75 μl min−1 using solvent A (0.1% formic acid in H2O) and solvent B (0.1% formic acid in acetonitrile). A 1 min wash period (99% solvent A) was performed prior to any gradient changes. After 1 min, solvent B increased to 60% over 10 min and up to 99% over another 10 min. The column was re-equilibrated to 99% solvent A for 5 min after each run. IM-MSE analyses were run in resolution mode, with a capillary voltage of 2.75 kV, source temperature at 100 °C, sample cone voltage at 30 V, extraction cone voltage at 5 V, source gas flow of 400 ml min−1, desolvation gas temperature of 325 °C, He cell flow of 180 ml min−1, and an IM gas flow of 90 ml min−1. The data were acquired in positive ion mode from 50 to 2000 Da with a 1 s scan time; Leucine enkephalin was used as the lock mass (m/z 556.2771 in ES + mode) at a concentration of 2 ng ml−1 infused at a flow rate of 7 μl min−1. All analytes were analyzed using MSE with an energy ramp from 10 to 40 eV.
High resolution (HR) MS and data-dependent acquisition analyses were performed on a high-resolution Q-Exactive HF hybrid quadrupole-Orbitrap mass spectrometer (Thermo Fisher Scientific, Bremen, Germany) equipped with a Vanquish UHPLC binary system and autosampler (Thermo Fisher Scientific, Germany).
For HILIC analysis both positive and negative ion mode, metabolite extracts were separated on a SeQuant ZIC-HILIC 3.5-μm, 2.1 mm × 100 mm column (Millipore Corporation, Darmstadt, Germany) held at 40 °C. Liquid chromatography was performed at a 200 μl min−1 using solvent A (5 mM Ammonium formate in 90% water, 10% acetonitrile) and solvent B (5 mM Ammonium formate in 90% acetonitrile, 10% water) with the following gradient: 95% B for 2 min, 95–40% B over 16 min, 40% B held 2 min, and 40–95% B over 15 min, 95% B held 10 min (gradient length 45 min).
Full MS analyses were acquired over a mass range of m/z 70–1050 using electrospray ionization both positive and negative ion mode. Full mass scan was used at a resolution of 120,000 with a scan rate of 3.5 Hz. The automatic gain control (AGC) target was set at 1 × 106 ions, and maximum ion injection time was at 100 ms. Source ionization parameters were optimized with the spray voltage at 3.0 kV, and other parameters were as follows: transfer temperature at 280 °C; S-Lens RF level at 40; heater temperature at 325 °C; Sheath gas at 40, Aux gas at 10, and sweep gas flow at 1. Tandem mass spectra were acquired using a data-dependent scanning mode in which one full MS scan (m/z 70–1050) was followed by 2, 4 or 6 MS/MS scans. MS/MS scans were acquired in profile mode using an isolation width of 1.3 m/z, stepped collision energy (NCE 20, 40), and a dynamic exclusion of 4 s. MS/MS spectra were collected at a resolution of 15,000, with an automatic gain control (AGC) target set at 2 × 105 ions, and maximum ion injection time of 100 ms. The retention times and peak areas of the isotopically labeled standards were used to assess data quality.
Metabolite data processing and analysis
The acquired UPLC-IM-MSE raw data and LC-HR MS/MS raw data were imported, processed, normalized and reviewed using Progenesis QI v.2.1 (Nonlinear Dynamics, Newcastle, UK). All MS and MS/MS sample runs for one particular analysis (RPLC or HILIC, positive or negative ion mode) were aligned against a quality control (pooled) reference run, and peak picking was performed on individual aligned runs to create an aggregate data set. Unique ions (retention time and m/z pairs) were grouped using both de-adduction and de-isotoping to generate unique “features” (retention time and m/z pairs) representative of unannotated metabolites. Data were normalized to all features using Progenesis QI. Compounds with <30% coefficient of variance (%CV) were retained for further analysis P values were calculated by Progenesis QI using variance stabilized measurements achieved through log normalization, and metabolites with a p-value ≤ 0.1 (method #1) and p value ≤ 0.05 (method #2) calculated by a one-way analysis of variance (ANOVA) test were considered significant. Metabolomic data has been filtered to remove known mass spectrometry contaminants140.
Tentative and putative annotations were determined within Progenesis QI software using accurate mass measurements (<5 ppm error), isotope distribution similarity, and fragmentation spectrum matching database searches against Human Metabolome Database (HMDB)141, METLIN142 the National Institute of Standards and Technology (NIST) database143 and an in-house library. Annotations from both RPLC and HILIC analyses were performed for all significant compounds (p value ≤ 0.1 or ≤ 0.05). Annotated metabolites were further analyzed by pathway overrepresentation analysis using MetaboAnalyst 4.0144. In these experiments, the level system for metabolite identification confidence was utilized. Briefly, many annotations were considered to be tentative (level 3, L3) when a top candidate cannot be prioritized145, but they still represent families of molecules representative for the data acquired. The annotations considered putative (level 2, L2) and validated (level 1, L1) are for molecules with a fragmentation spectrum matching one of the databases or a standard molecule from the in-house library. All metabolite measurements and their annotations are uploaded (Supplementary Table 4).
Pathway analysis
Significantly differentially expressed genes were evaluated in the STRING v11 database, a program that correlates the direct and indirect associations found between proteins146. GO 2021 pathways were determined using the publicly available Enrichr database147,148.
Data availability
Raw RNA sequencing data and processed gene counts that support the findings of this study are available through the Gene Expression Omnibus at GSE184694. Metabolomics data are available at the NIH Common Fund’s National Metabolomics Data Repository (NMDR) Web site, the Metabolomics Workbench, https://www.metabolomicsworkbench.org where it has been assigned Study ID (ST001957).
References
Kalia, L. V. & Lang, A. E. Parkinson’s disease. Lancet (Lond., Engl.) 386, 896–912 (2015).
Nalls, M. A. et al. Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet Neurol. 18, 1091–1102 (2019).
Blauwendraat, C., Nalls, M. A. & Singleton, A. B. The genetic architecture of Parkinson’s disease. Lancet Neurol. 19, 170–178 (2020).
Bandres-Ciga, S., Diez-Fairen, M., Kim, J. J. & Singleton, A. B. Genetics of Parkinson’s disease: An introspection of its journey towards precision medicine. Neurobiol. Dis. 137, 104782 (2020).
Puschmann, A. New Genes Causing Hereditary Parkinson’s Disease or Parkinsonism. Curr. Neurol. Neurosci. Rep. 17, 66 (2017).
Toffoli, M., Vieira, S. R. L. & Schapira, A. H. V. Genetic causes of PD: A pathway to disease modification. Neuropharmacology. 170, 108022 (2020).
Corti, O., Lesage, S. & Brice, A. What genetics tells us about the causes and mechanisms of Parkinson’s disease. Physiol. Rev. 91, 1161–1218 (2011).
Doherty, K. M. et al. Parkin disease: a clinicopathologic entity? JAMA Neurol. 70, 571–579 (2013).
Kitada, T. et al. Mutations in the parkin gene cause autosomal recessive juvenile parkinsonism. Nature 392, 605–608 (1998).
Lücking, C. B. et al. Association between early-onset Parkinson’s disease and mutations in the parkin gene. N. Engl. J. Med. 342, 1560–1567 (2000).
Lunati, A., Lesage, S. & Brice, A. The genetic landscape of Parkinson’s disease. Rev. Neurol. (Paris) 174, 628–643 (2018).
Periquet, M. et al. Origin of the mutations in the parkin gene in Europe: exon rearrangements are independent recurrent events, whereas point mutations may result from Founder effects. Am. J. Hum. Genet 68, 617–626 (2001).
Abbas, N. et al. A wide variety of mutations in the parkin gene are responsible for autosomal recessive parkinsonism in Europe. French Parkinson’s Disease Genetics Study Group and the European Consortium on Genetic Susceptibility in Parkinson’s Disease. Hum. Mol. Genet 8, 567–574 (1999).
Nichols, W. C. et al. Linkage stratification and mutation analysis at the Parkin locus identifies mutation positive Parkinson’s disease families. J. Med. Genet 39, 489–492 (2002).
Puschmann, A. Monogenic Parkinson’s disease and parkinsonism: clinical phenotypes and frequencies of known mutations. Parkinsonism Relat. Disord. 19, 407–415 (2013).
Periquet, M. et al. Parkin mutations are frequent in patients with isolated early-onset parkinsonism. Brain 126, 1271–1278 (2003).
Klein, C. et al. Parkin deletions in a family with adult-onset, tremor-dominant parkinsonism: expanding the phenotype. Ann. Neurol. 48, 65–71 (2000).
Khan, N. L. et al. Parkin disease: a phenotypic study of a large case series. Brain 126, 1279–1292 (2003).
Deng, H. et al. Heterogeneous phenotype in a family with compound heterozygous parkin gene mutations. Arch. Neurol. 63, 273–277 (2006).
Isaacs, D., Claassen, D., Bowman, A. B. & Hedera, P. Phenotypic Discordance in Siblings with Identical Compound Heterozygous PARK2 Mutations. Brain Sci. 7, 71 (2017).
Schwamborn, J. C. Is Parkinson’s Disease a Neurodevelopmental Disorder and Will Brain Organoids Help Us to Understand It? Stem Cells Dev. 27, 968–975 (2018).
Le, W., Conneely, O. M., He, Y., Jankovic, J. & Appel, S. H. Reduced Nurr1 expression increases the vulnerability of mesencephalic dopamine neurons to MPTP-induced injury. J. Neurochem 73, 2218–2221 (1999).
von Linstow, C. U., DeLano-Taylor, M., Kordower, J. H. & Brundin, P. Does Developmental Variability in the Number of Midbrain Dopamine Neurons Affect Individual Risk for Sporadic Parkinson’s Disease? J. Parkinsons Dis. 10, 405–411 (2020).
Barlow, B. K., Cory-Slechta, D. A., Richfield, E. K. & Thiruchelvam, M. The gestational environment and Parkinson’s disease: evidence for neurodevelopmental origins of a neurodegenerative disorder. Reprod. Toxicol. 23, 457–470 (2007).
Huntley, G. W. & Benson, D. L. Origins of Parkinson’s Disease in Brain Development: Insights From Early and Persistent Effects of LRRK2-G2019S on Striatal Circuits. Front Neurosci. 14, 265 (2020).
Candelario, K. M. et al. Exosome/microvesicle content is altered in leucine-rich repeat kinase 2 mutant induced pluripotent stem cell-derived neural cells. J. Comp. Neurol. 528, 1203–1215 (2020).
Huisman, E., Uylings, H. B. & Hoogland, P. V. A 100% increase of dopaminergic cells in the olfactory bulb may explain hyposmia in Parkinson’s disease. Mov. Disord. 19, 687–692 (2004).
Hoogland, P. V., van den Berg, R. & Huisman, E. Misrouted olfactory fibres and ectopic olfactory glomeruli in normal humans and in Parkinson and Alzheimer patients. Neuropathol. Appl Neurobiol. 29, 303–311 (2003).
Garcia-Reitboeck, P. et al. Endogenous alpha-synuclein influences the number of dopaminergic neurons in mouse substantia nigra. Exp. Neurol. 248, 541–545 (2013).
d’Amora, M. et al. Expression of PINK1 in the brain, eye and ear of mouse during embryonic development. J. Chem. Neuroanat. 41, 73–85 (2011).
MacLeod, D. et al. The familial Parkinsonism gene LRRK2 regulates neurite process morphology. Neuron 52, 587–593 (2006).
Anichtchik, O. et al. Loss of PINK1 function affects development and results in neurodegeneration in zebrafish. J. Neurosci. 28, 8199–8207 (2008).
Liu, G. H. et al. Progressive degeneration of human neural stem cells caused by pathogenic LRRK2. Nature 491, 603–607 (2012).
Walter, J. et al. Neural Stem Cells of Parkinson’s Disease Patients Exhibit Aberrant Mitochondrial Morphology and Functionality. Stem Cell Rep. 12, 878–889 (2019).
Jankovic, J. Parkinson’s disease: clinical features and diagnosis. J. Neurol. Neurosurg. Psychiatry 79, 368–376 (2008).
Aboud, A. A. et al. PARK2 patient neuroprogenitors show increased mitochondrial sensitivity to copper. Neurobiol. Dis. 73, 204–212 (2015).
Muller, F. J. et al. A bioinformatic assay for pluripotency in human cells. Nat. Methods 8, 315–317 (2011).
Neely, M. D. et al. DMH1, a highly selective small molecule BMP inhibitor promotes neurogenesis of hiPSCs: comparison of PAX6 and SOX1 expression during neural induction. ACS Chem. Neurosci. 3, 482–491 (2012). (PMC888888).
Neely, M. D., Davison, C. A., Aschner, M. & Bowman, A. B. From the Cover: Manganese and Rotenone-Induced Oxidative Stress Signatures Differ in iPSC-Derived Human Dopamine Neurons. Toxicol. Sci. 159, 366–379 (2017).
Kriks, S. et al. Dopamine neurons derived from human ES cells efficiently engraft in animal models of Parkinson’s disease. Nature 480, 547–551 (2011).
Toczylowska, B., Zieminska, E., Michałowska, M., Chalimoniuk, M. & Fiszer, U. Changes in the metabolic profiles of the serum and putamen in Parkinson’s disease patients - In vitro and in vivo NMR spectroscopy studies. Brain Res 1748, 147118 (2020).
Clarke, C. E., Lowry, M. & Horsman, A. Unchanged basal ganglia N-acetylaspartate and glutamate in idiopathic Parkinson’s disease measured by proton magnetic resonance spectroscopy. Mov. Disord. 12, 297–301 (1997).
Havelund, J. F., Heegaard, N. H. H., Færgeman, N. J. K. & Gramsbergen, J. B. Biomarker Research in Parkinson’s Disease Using Metabolite Profiling. Metabolites 7, 42 (2017).
Li, M. et al. Integrative functional genomic analysis of human brain development and neuropsychiatric risks. Science 362, eaat7615 (2018).
Le Grand, J. N., Gonzalez-Cano, L., Pavlou, M. A. & Schwamborn, J. C. Neural stem cells in Parkinson’s disease: a role for neurogenesis defects in onset and progression. Cell Mol. Life Sci. 72, 773–797 (2015).
Lin, L. et al. Molecular Features Underlying Neurodegeneration Identified through In Vitro Modeling of Genetically Diverse Parkinson’s Disease Patients. Cell Rep. 15, 2411–2426 (2016).
Kelly, J., Moyeed, R., Carroll, C., Albani, D. & Li, X. Gene expression meta-analysis of Parkinson’s disease and its relationship with Alzheimer’s disease. Mol. Brain 12, 16 (2019).
Benoit, S. M. et al. Expanding the search for genetic biomarkers of Parkinson’s disease into the living brain. Neurobiol. Dis. 140, 104872 (2020).
Oguro-Ando, A., Zuko, A., Kleijer, K. T. E. & Burbach, J. P. H. A current view on contactin-4, -5, and -6: Implications in neurodevelopmental disorders. Mol. Cell Neurosci. 81, 72–83 (2017).
Alsanie, W. F., Penna, V., Schachner, M., Thompson, L. H. & Parish, C. L. Homophilic binding of the neural cell adhesion molecule CHL1 regulates development of ventral midbrain dopaminergic pathways. Sci. Rep. 7, 9368 (2017).
Bye, C. R., Jönsson, M. E., Björklund, A., Parish, C. L. & Thompson, L. H. Transcriptome analysis reveals transmembrane targets on transplantable midbrain dopamine progenitors. Proc. Natl Acad. Sci. USA 112, E1946–E1955 (2015).
Bye, C. R., Rytova, V., Alsanie, W. F., Parish, C. L. & Thompson, L. H. Axonal Growth of Midbrain Dopamine Neurons is Modulated by the Cell Adhesion Molecule ALCAM Through Trans-Heterophilic Interactions with L1cam, Chl1, and Semaphorins. J. Neurosci. 39, 6656–6667 (2019).
Sytnyk, V., Leshchyns’ka, I. & Schachner, M. Neural Cell Adhesion Molecules of the Immunoglobulin Superfamily Regulate Synapse Formation, Maintenance, and Function. Trends Neurosci. 40, 295–308 (2017).
Voorn, P., Kalsbeek, A., Jorritsma-Byham, B. & Groenewegen, H. J. The pre- and postnatal development of the dopaminergic cell groups in the ventral mesencephalon and the dopaminergic innervation of the striatum of the rat. Neuroscience 25, 857–887 (1988).
Van den Heuvel, D. M. & Pasterkamp, R. J. Getting connected in the dopamine system. Prog. Neurobiol. 85, 75–93 (2008).
Hu, J. et al. CNTN6 copy number variations in 14 patients: a possible candidate gene for neurodevelopmental and neuropsychiatric disorders. J. Neurodev. Disord. 7, 26 (2015).
Dumitriu, A. et al. Integrative analyses of proteomics and RNA transcriptomics implicate mitochondrial processes, protein folding pathways and GWAS loci in Parkinson disease. BMC Med. Genomics 9, 5 (2016).
Rissling, I., Strauch, K., Höft, C., Oertel, W. H. & Möller, J. C. Haplotype Analysis of the Engrailed-2 Gene in Young-Onset Parkinson’s Disease. Neurodegenerative Dis. 6, 102–105 (2009).
Fuchs, J. et al. Engrailed signaling in axon guidance and neuron survival. Eur. J. Neurosci. 35, 1837–1845 (2012).
Abeliovich, A. & Hammond, R. Midbrain dopamine neuron differentiation: factors and fates. Dev. Biol. 304, 447–454 (2007).
Alavian, K. N., Scholz, C. & Simon, H. H. Transcriptional regulation of mesencephalic dopaminergic neurons: the full circle of life and death. Mov. Disord. 23, 319–328 (2008).
Blaess, S. & Ang, S. L. Genetic control of midbrain dopaminergic neuron development. Wiley Interdiscip. Rev. Dev. Biol. 4, 113–134 (2015).
Arenas, E., Denham, M. & Villaescusa, J. C. How to make a midbrain dopaminergic neuron. Development 142, 1918–1936 (2015).
Simon, H. H., Saueressig, H., Wurst, W., Goulding, M. D. & O’Leary, D. D. Fate of midbrain dopaminergic neurons controlled by the engrailed genes. J. Neurosci. 21, 3126–3134 (2001).
Genestine, M. et al. Engrailed-2 (En2) deletion produces multiple neurodevelopmental defects in monoamine systems, forebrain structures and neurogenesis and behavior. Hum. Mol. Genet 24, 5805–5827 (2015).
Ceballos-Picot, I. et al. Hypoxanthine-guanine phosphoribosyl transferase regulates early developmental programming of dopamine neurons: implications for Lesch-Nyhan disease pathogenesis. Hum. Mol. Genet 18, 2317–2327 (2009).
Chacon, M. A. & Boulanger, L. M. MHC class I protein is expressed by neurons and neural progenitors in mid-gestation mouse brain. Mol. Cell Neurosci. 52, 117–127 (2013).
Shatz, C. J. MHC class I: an unexpected role in neuronal plasticity. Neuron 64, 40–45 (2009).
Zhang, A. et al. Developmental expression and localization of MHC class I molecules in the human central nervous system. Exp. Brain Res. 233, 2733–2743 (2015).
Zohar, O. et al. Cutting edge: MHC class I-Ly49 interaction regulates neuronal function. J. Immunol. 180, 6447–6451 (2008).
Corriveau, R. A., Huh, G. S. & Shatz, C. J. Regulation of class I MHC gene expression in the developing and mature CNS by neural activity. Neuron 21, 505–520 (1998).
Bilousova, T. et al. Major histocompatibility complex class I molecules modulate embryonic neuritogenesis and neuronal polarization. J. Neuroimmunol. 247, 1–8 (2012).
Escande-Beillard, N. et al. Neurons preferentially respond to self-MHC class I allele products regardless of peptide presented. J. Immunol. 184, 816–823 (2010).
Wu, Z. P. et al. Enhanced neuronal expression of major histocompatibility complex class I leads to aberrations in neurodevelopment and neurorepair. J. Neuroimmunol. 232, 8–16 (2011).
Goddard, C. A., Butts, D. A. & Shatz, C. J. Regulation of CNS synapses by neuronal MHC class I. Proc. Natl Acad. Sci. USA 104, 6828–6833 (2007).
Lv, D. et al. The similar expression pattern of MHC class I molecules in human and mouse cerebellar cortex. Neurochem Res. 39, 180–186 (2014).
Elmer, B. M. & McAllister, A. K. Major histocompatibility complex class I proteins in brain development and plasticity. Trends Neurosci. 35, 660–670 (2012).
Huh, G. S. et al. Functional requirement for class I MHC in CNS development and plasticity. Science 290, 2155–2159 (2000).
Ishii, T., Hirota, J. & Mombaerts, P. Combinatorial coexpression of neural and immune multigene families in mouse vomeronasal sensory neurons. Curr. Biol. 13, 394–400 (2003).
Loconto, J. et al. Functional expression of murine V2R pheromone receptors involves selective association with the M10 and M1 families of MHC class Ib molecules. Cell 112, 607–618 (2003).
Glynn, M. W. et al. MHCI negatively regulates synapse density during the establishment of cortical connections. Nat. Neurosci. 14, 442–451 (2011).
Vagaska, B. et al. MHC-class-II are expressed in a subpopulation of human neural stem cells in vitro in an IFNγ-independent fashion and during development. Sci. Rep. 6, 24251 (2016).
Laguna Goya, R., Busch, R., Mathur, R., Coles, A. J. & Barker, R. A. Human fetal neural precursor cells can up-regulate MHC class I and class II expression and elicit CD4 and CD8 T cell proliferation. Neurobiol. Dis. 41, 407–414 (2011).
Lampe, J. B. et al. HLA typing and Parkinson’s disease. Eur. Neurol. 50, 64–68 (2003).
Hamza, T. H. et al. Common genetic variation in the HLA region is associated with late-onset sporadic Parkinson’s disease. Nat. Genet 42, 781–785 (2010).
Ahmed, I. et al. Association between Parkinson’s disease and the HLA-DRB1 locus. Mov. Disord. 27, 1104–1110 (2012).
Nalls, M. A. et al. Imputation of sequence variants for identification of genetic risks for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet 377, 641–649 (2011).
Nalls, M. A. et al. Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease. Nat. Genet 46, 989–993 (2014).
Wissemann, W. T. et al. Association of Parkinson disease with structural and regulatory variants in the HLA region. Am. J. Hum. Genet 93, 984–993 (2013).
Hill-Burns, E. M., Factor, S. A., Zabetian, C. P., Thomson, G. & Payami, H. Evidence for more than one Parkinson’s disease-associated variant within the HLA region. PLoS One 6, e27109 (2011).
Spina, M. B. & Cohen, G. Dopamine turnover and glutathione oxidation: implications for Parkinson disease. Proc. Natl Acad. Sci. USA 86, 1398–1400 (1989).
Götz, M. E., Freyberger, A. & Riederer, P. Oxidative stress: a role in the pathogenesis of Parkinson’s disease. J. Neural Transm. Suppl. 29, 241–249 (1990).
Mazzetti, A. P., Fiorile, M. C., Primavera, A. & Lo Bello, M. Glutathione transferases and neurodegenerative diseases. Neurochem Int 82, 10–18 (2015).
Jenner, P. Oxidative stress as a cause of Parkinson’s disease. Acta Neurol. Scand. Suppl. 136, 6–15 (1991).
Allocati, N., Masulli, M., Di Ilio, C. & Federici, L. Glutathione transferases: substrates, inihibitors and pro-drugs in cancer and neurodegenerative diseases. Oncogenesis 7, 8 (2018).
Johnson, W. M., Wilson-Delfosse, A. L. & Mieyal, J. J. Dysregulation of glutathione homeostasis in neurodegenerative diseases. Nutrients 4, 1399–1440 (2012).
Garte, S. et al. Metabolic gene polymorphism frequencies in control populations. Cancer Epidemiol. Biomark. Prev. 10, 1239–1248 (2001).
Goldman, S. M. et al. Genetic modification of the association of paraquat and Parkinson’s disease. Mov. Disord. 27, 1652–1658 (2012).
Rahbar, A. et al. Glutathione S-transferase polymorphism in Parkinson’s disease. J. Neural Transm. (Vienna) 107, 331–334 (2000).
Pinhel, M. A. et al. Nullity of GSTT1/GSTM1 related to pesticides is associated with Parkinson’s disease. Arq. Neuropsiquiatr. 71, 527–532 (2013).
De Palma, G. et al. A case-control study of Parkinson’s disease and tobacco use: gene-tobacco interactions. Mov. Disord. 25, 912–919 (2010).
Smeyne, M. & Smeyne, R. J. Glutathione metabolism and Parkinson’s disease. Free Radic. Biol. Med. 62, 13–25 (2013).
DeStefano, A. L. et al. Genome-wide scan for Parkinson’s disease: the GenePD Study. Neurology 57, 1124–1126 (2001).
Punia, S. et al. Role of polymorphisms in dopamine synthesis and metabolism genes and association of DBH haplotypes with Parkinson’s disease among North Indians. Pharmacogenet Genomics 20, 435–441 (2010).
Healy, D. G. et al. A functional polymorphism regulating dopamine beta-hydroxylase influences against Parkinson’s disease. Ann. Neurol. 55, 443–446 (2004).
Zabetian, C. P. et al. A quantitative-trait analysis of human plasma-dopamine beta-hydroxylase activity: evidence for a major functional polymorphism at the DBH locus. Am. J. Hum. Genet 68, 515–522 (2001).
Shao, P., Yu, Y. X. & Bao, J. X. Association of Dopamine Beta-Hydroxylase (DBH) Polymorphisms with Susceptibility to Parkinson’s Disease. Med Sci. Monit. 22, 1617–1622 (2016).
Chun, L. S. et al. DBH -1021C->T does not modify risk or age at onset in Parkinson’s disease. Ann. Neurol. 62, 99–101 (2007).
Ross, O. A. et al. Dopamine beta-hydroxylase -1021C>T association and Parkinson’s disease. Parkinsonism Relat. Disord. 14, 544–547 (2008).
Ghosh, A. et al. Dopamine β Hydroxylase (DBH) is a potential modifier gene associated with Parkinson’s disease in Eastern India. Neurosci. Lett. 706, 75–80 (2019).
Okarmus, J. et al. Identification of bioactive metabolites in human iPSC-derived dopaminergic neurons with PARK2 mutation: Altered mitochondrial and energy metabolism. Stem Cell Rep. 16, 1510–1526 (2021).
Ostrakhovitch, E. A. et al. Analysis of circulating metabolites to differentiate Parkinson’s disease and essential tremor. Neurosci. Lett. 769, 136428 (2022).
Okarmus, J. et al. Lysosomal perturbations in human dopaminergic neurons derived from induced pluripotent stem cells with PARK2 mutation. Sci. Rep. 10, 10278 (2020).
Palmer, J. A. et al. Establishment and assessment of a new human embryonic stem cell-based biomarker assay for developmental toxicity screening. Birth Defects Res B Dev. Reprod. Toxicol. 98, 343–363 (2013).
Burke, E. A. et al. Biallelic mutations in mitochondrial tryptophanyl-tRNA synthetase cause Levodopa-responsive infantile-onset Parkinsonism. Clin. Genet 93, 712–718 (2018).
Boczonadi, V., Jennings, M. J. & Horvath, R. The role of tRNA synthetases in neurological and neuromuscular disorders. FEBS Lett. 592, 703–717 (2018).
Sissler, M., González-Serrano, L. E. & Westhof, E. Recent Advances in Mitochondrial Aminoacyl-tRNA Synthetases and Disease. Trends Mol. Med. 23, 693–708 (2017).
Lee, Y. et al. Parthanatos mediates AIMP2-activated age-dependent dopaminergic neuronal loss. Nat. Neurosci. 16, 1392–1400 (2013).
Ko, H. S. et al. Phosphorylation by the c-Abl protein tyrosine kinase inhibits parkin’s ubiquitination and protective function. Proc. Natl Acad. Sci. USA 107, 16691–16696 (2010).
Hu, L. et al. Integrated Metabolomics and Proteomics Analysis Reveals Plasma Lipid Metabolic Disturbance in Patients With Parkinson’s Disease. Front Mol. Neurosci. 13, 80 (2020).
Fernández-Irigoyen, J., Cartas-Cejudo, P., Iruarrizaga-Lejarreta, M. & Santamaría, E. Alteration in the Cerebrospinal Fluid Lipidome in Parkinson’s Disease: A Post-Mortem Pilot Study. Biomedicines 9, 491 (2021).
Quinville, B. M., Deschenes, N. M., Ryckman, A. E. & Walia, J. S. A Comprehensive Review: Sphingolipid Metabolism and Implications of Disruption in Sphingolipid Homeostasis. Int. J. Mol. Sci. 22, 793 (2021).
Ayub, M., Jin, H. K. & Bae, J. S. Novelty of Sphingolipids in the Central Nervous System Physiology and Disease: Focusing on the Sphingolipid Hypothesis of Neuroinflammation and Neurodegeneration. Int. J. Mol. Sci. 22, 353 (2021).
Avisar, H. et al. Lipidomics Prediction of Parkinson’s Disease Severity: A Machine-Learning. Anal. J. Parkinsons Dis. 11, 1141–1155 (2021).
Dong, M. X., Wei, Y. D. & Hu, L. Lipid metabolic dysregulation is involved in Parkinson’s disease dementia. Metab. Brain Dis. 36, 463–470 (2021).
Klemann, C. et al. Integrated molecular landscape of Parkinson’s disease. NPJ Parkinsons Dis. 3, 14 (2017).
Xicoy, H. et al. Shared genetic etiology between Parkinson’s disease and blood levels of specific lipids. NPJ Parkinsons Dis. 7, 23 (2021).
Yu, R. K., Tsai, Y. T. & Ariga, T. Functional roles of gangliosides in neurodevelopment: an overview of recent advances. Neurochem Res. 37, 1230–1244 (2012).
Palmano, K., Rowan, A., Guillermo, R., Guan, J. & McJarrow, P. The role of gangliosides in neurodevelopment. Nutrients 7, 3891–3913 (2015).
Lukacs, M., Blizzard, L. E. & Stottmann, R. W. CNS glycosylphosphatidylinositol deficiency results in delayed white matter development, ataxia and premature death in a novel mouse model. Hum. Mol. Genet 29, 1205–1217 (2020).
Lukacs, M., Roberts, T., Chatuverdi, P. & Stottmann, R. W. Glycosylphosphatidylinositol biosynthesis and remodeling are required for neural tube closure, heart development, and cranial neural crest cell survival. Elife 8, e45248 (2019).
Buddington, R. K. et al. A Phosphatidylserine Source of Docosahexanoic Acid Improves Neurodevelopment and Survival of Preterm Pigs. Nutrients 10, 637 (2018).
Fuller, M. & Futerman, A. H. The brain lipidome in neurodegenerative lysosomal storage disorders. Biochem Biophys. Res Commun. 504, 623–628 (2018).
Okita, K. et al. A more efficient method to generate integration-free human iPS cells. Nat. Methods 8, 409–412 (2011).
Aboud, A. A. et al. Genetic risk for Parkinson’s disease correlates with alterations in neuronal manganese sensitivity between two human subjects. Neurotoxicology 33, 1443–1449 (2012).
Kumar, K. K. et al. Cellular manganese content is developmentally regulated in human dopaminergic neurons. Sci. Rep. 4, 6801 (2014).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20, 1297–1303 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinforma. (Oxf., Engl.) 26, 139–140 (2010).
de Ronde, M. W. J. et al. Practical data handling pipeline improves performance of qPCR-based circulating miRNA measurements. Rna 23, 811–821 (2017).
Weber, R. J., Li, E., Bruty, J., He, S. & Viant, M. R. MaConDa: a publicly accessible mass spectrometry contaminants database. Bioinforma. (Oxf., Engl.) 28, 2856–2857 (2012).
Wishart, D. S. et al. HMDB 3.0-The Human Metabolome Database in 2013. Nucleic Acids Res 41, D801–D807 (2013).
Smith, C. A. et al. METLIN: a metabolite mass spectral database. Ther. Drug Monit. 27, 747–751 (2005).
Jablonski, F., Salvat, F., Powell, C. J. & Lee, A. Y. in NIST Standard Reference Database Number 64 (ed National Institute of Standards and Technology) (2016).
Chong, J., Wishart, D. S. & Xia, J. Using MetaboAnalyst 4.0 for Comprehensive and Integrative Metabolomics Data Analysis. Curr. Protoc. Bioinforma. 68, e86 (2019).
Schrimpe-Rutledge, A. C., Codreanu, S. G., Sherrod, S. D. & McLean, J. A. Untargeted Metabolomics Strategies-Challenges and Emerging Directions. J. Am. Soc. Mass Spectrom. 27, 1897–1905 (2016).
Szklarczyk, D. et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–d613 (2019).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinforma. 14, 128 (2013).
Xie, Z. et al. Gene Set Knowledge Discovery with Enrichr. Curr. Protoc. 1, e90 (2021).
Acknowledgements
This study was supported by R01 ES010563 (A.B.B.), by the John P. Hussman Institute for Human Genomics (H.N.C) and the Center for Innovative Technology (CIT) at Vanderbilt University (J.A.M) and R03NS125243 (S.D.S). This study was also partially supported by CTSA award No. UL1 TR002243 from the National Center for Advancing Translational Sciences (subaward to M.D.N.). Its contents are solely the responsibility of the authors and do not necessarily represent official views of the National Center for Advancing Translational Sciences or the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
H.N.C., A.B.B., and M.D.N. conceived, designed, and analyzed the experiments. M.D.N. and K.C.E. generated the hiPSC lines and derived them into dopaminergic neurons. H.N.C. and A.J.G. performed the whole exome sequencing and RNA-sequencing, analysis, and interpretation. A.J.G. carried out the CNV analysis. H.K. and L.M.P. performed the RT-qPCR validation and analysis. S.G.C., S.D.S. and J.A.M. executed the metabolomics experiments and data analysis. H.N.C. and S.G.C. performed pathway analysis. D.M.D. assisted in data analysis and interpretation for the sequencing data. P.H. and K.C.E. ascertained the SM and PM brothers. H.N.C. and M.D.N. wrote the manuscript and H.K., S.G.C., and A.B.B. edited the manuscript. All authors have reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cukier, H.N., Kim, H., Griswold, A.J. et al. Genomic, transcriptomic, and metabolomic profiles of hiPSC-derived dopamine neurons from clinically discordant brothers with identical PRKN deletions. npj Parkinsons Dis. 8, 84 (2022). https://doi.org/10.1038/s41531-022-00346-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41531-022-00346-3