Abstract
Poly(ADP-ribose) (PAR), a non-canonical nucleic acid, is essential for DNA/RNA metabolism and protein condensation, and its dysregulation is linked to cancer and neurodegeneration. However, key structural insights into PAR’s functions remain largely uncharacterized, hindered by the challenges in synthesizing and characterizing PAR, which are attributed to its length heterogeneity. A central issue is how PAR, comprised solely of ADP-ribose units, attains specificity in its binding and condensing proteins based on chain length. Here, we integrate molecular dynamics simulations with small-angle X-ray scattering to analyze PAR structures. We identify diverse structural ensembles of PAR that fall into distinct subclasses and reveal distinct compaction of two different lengths of PAR upon the addition of small amounts of Mg2+ ions. Unlike PAR15, PAR22 forms ADP-ribose bundles via local intramolecular coil-to-globule transitions. Understanding these length-dependent structural changes could be central to deciphering the specific biological functions of PAR.
Similar content being viewed by others
Introduction
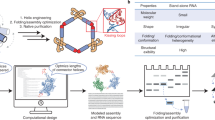
Poly(ADP-ribose) (PAR) is a non-canonical nucleic acid produced in cells as a post-translational modification by ADP-ribosyltransferases commonly known as PARPs (Fig. 1A)1,2. Functionally, PAR may serve as a signal mediator, conjugating to proteins involved in restoring homeostasis after stress such as DNA damage and inflammation (Fig. 1B)3,4,5,6,7. PAR may also serve as a scaffold for protein complex or biomolecular condensate formation, where its negative charge allows for multivalent noncovalent interactions with proteins8,9.
A ADP-ribosylation is heterogenous, presented as monomeric, linear or branched polymer forms as poly(ADP-ribose) or PAR. B PAR mediates various cellular processes at distinct subcellular locations. “Animal cell photo” by Tomáš Kebert & umimeto.org, used under CC BY-SA 4.0 and was cropped from the original image. C Pulldown of PAR-binding proteins in nuclear lysate coupled with mass spectrometry analysis indicates a preference of proteome to the length of PAR (data reproduced from Dasovich et al. with permission)12. D Biochemical studies10 showing that the strength of PAR-proteins interaction is influenced by the chain length (n). ‘−’ indicates no binding, ‘+’ indicates weak binding and ‘+++’ indicates strong binding (<1 µM dissociation constant). E PAR modulates the formation and dynamics of biomolecular condensates linked to human disease. In vitro microscopic studies showing that PAR chain length influences material properties of FUS condensates, both at 1 µM; scale bar, 5 µm. These images were adopted from Rhine et al. with permission16.
The binding affinity of PAR to proteins depends on their chain length on a global scale, where its specificity could be even down to a single ADP-ribose difference (Fig. 1C, D)10,11,12,13,14. In certain cases, such as with oncoprotein DEK, a threshold PAR length is necessary for appreciable binding to occur15. Moreover, 4-mer PAR does not induce the condensation of FUS (a key protein associated with cancers and neurodegenerative diseases), whereas 8- and 16-mer do, and 32-mer drives its aggregation (Fig. 1E)16. So, how does a simple homopolymer achieve such length-specific protein interactions? We posited that PAR may adopt distinct structures based on its polymer length.
Despite its functional importance, the 3D structure of PAR has not been extensively investigated, hampered by the challenges in synthesizing and characterizing PAR due to its heterogeneity in length. Previous circular dichroism analyses of mixed chain populations suggested the possibility of secondary structure formation contingent on cations17. However, atomic structures beyond dimeric ADP-ribose have not been identified by experiments18,19,20. Nuclear magnetic resonance studies did not identify any well-defined structures within populations characterized by mixed chain lengths21. Molecular dynamics (MD) simulations reported multiglobular conformations in a 25-mer, but not in a 5-mer22. This phenomenon occurs despite the rigidity of dihedral bonds at individual ribose-ribose linkages, where configurational entropy becomes predominant in longer polymers. Importantly, the simulations faced limitations in effectively sampling configurations attributed to their constrained duration (50 ns) and imperfections in the molecular force field describing cation-phosphate interactions23. Notably, interpreting the dynamic structural ensembles of such macromolecules is complex, underscoring the need for advanced analysis and visualization methods. Although PAR’s functionality is influenced by its chain length, it remains uncertain whether a single structural model can capture the length-dependent diversity in biological PAR activity. A detailed analysis of the structural features of PAR at different lengths could elucidate its length-dependent conformations and potentially clarify its selective interactions with binding partners.
In mammalian cells under normal and mildly stressed conditions, PAR oligomers exhibit a size distribution spanning from as few as 2 units to approximately 20 units24,25,26. Likewise, bacterial PARPs generate PAR within a similar range of lengths27,28.
Here, we integrate MD simulations with small-angle X-ray scattering (SAXS) measurements to determine structural ensembles of two physiologically relevant PAR lengths, PAR15 and PAR22, under various ionic conditions, yielding atomic-level snapshots of PAR structures substantiated by experimental data. This approach further distinguishes itself from earlier MD analyses by incorporating multiple microsecond-long atomistic simulations, enhanced with state-of-the-art corrections to nucleic acid force field parameters29. This advance enables us to accurately map the equilibrium ensembles of two PAR polymers. We further refine these conformational ensembles using SAXS data, thereby accurately assigning statistical weights to the conformations observed in the MD simulations. In addition, we employ graph theory as a systematic approach to categorize and visualize the PAR ensembles—a technique that could be broadly applied to the study of other disordered macromolecules. Based on the SAXS-guided MD analyses, we define structural order parameters to categorize backbone conformations across the structural ensembles of PAR15 and PAR22. Our results indicate that both backbone tortuosity and base stacking contribute to PAR compaction in unique ways for both lengths. By decomposing the structural ensembles into easily visualizable components using 3D class averaging in real space, we identified widespread bundles of ADP-ribose units across the PAR22 ensemble, but not PAR15, in the presence of Mg2+. This detailed structural description offers a possible explanation for the heterogeneity in structural conformations and binding behaviors exhibited by PARs of different lengths.
Results
PAR compaction and structural dynamics are influenced by cationic environment
To build a comprehensive understanding of PAR’s conformational behavior in different environments, we performed explicit solvent MD simulations of PAR22 for a range of electrolyte conditions. A typical simulation began with the PAR polymer in a fully extended state, submerged in a 26 × 23 × 9 nm3 volume of electrolyte solution, a system of about 500,000 atoms (Fig. 2A). Upon energy minimization, each system was simulated for over 300 ns in the absence of any restraints using a CHARMM-compatible model of the PAR polymer (see “Methods” for details). Within the first 50 ns of equilibration, the polymer transitioned to a more compact state (Fig. 2B). The remaining 250 ns of each free equilibration simulation were characterized by computing the end-to-end distance (REE) and the radius of gyration (Rg) of the PAR polymer.
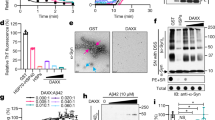
A Initial configuration of a typical simulation system. One 22-mer PAR polymer is placed in electrolyte solution (semitransparent molecular surface) containing 50 mM of NaCl. B End-to-end distance (REE, in red) and radius of gyration (Rg, in blue) of a PAR molecule as a function of time simulated in 50 mM NaCl electrolyte. C, E, G Average equilibrium end-to-end distance (circles) and radius of gyration (squares) of the 22-unit PAR polymer at various ion conditions. Dashed lines connect the points to guide the eye. Each data point represents a 250-ns trajectory average after exclusion of the first 50 ns in each simulation where the molecule started in an extended state. Error bars denote S.D. from the average value. D, F, H Representative snapshots of PAR conformation at the end of a 300 ns equilibration performed at the specified ion concentration conditions. The O3′, C3′, C4′, C5′, and O5′ atoms of PAR are shown in green, whereas all other atoms are in blue. Na+ (yellow), Cl− (green) and Mg2+ (pink) ions located within 6 Å of PAR are shown as spheres. The ends of the PAR chains are depicted in red.
Prior research, including circular dichroism analyses of mixed-chain length PAR molecules and our recent single-molecule Förster resonance energy transfer (smFRET) measurements, revealed that PAR compaction is sensitive to cations17,30. These findings align with established knowledge that nucleic acid structures are sensitive to their cationic environment31,32. In our MD simulations, we observed that both REE and Rg steadily decrease as the concentration of Na+ increases (Fig. 2C). Visualization of typical conformations indicates a transition: PAR adopts an extended, linear-like conformation at low salt concentrations and changes to a more compact, globular conformation at high salt concentrations (Fig. 2D). In the compact state, Na+ ions screened the backbone charge of PAR22, facilitating proximity between ADP-ribose units (Fig. 2D).
We extended our simulations to examine the impact of Mg2+ on PAR conformation and found an extreme sensitivity to this cation (Fig. 2E). In this set of simulations, we kept the total charge of cations equal in magnitude to the charge of PAR22, while varying the fraction of the charge neutralized by Mg2+ from 0 to 100% (Fig. 2E, red points). Remarkably, increasing the fraction of charge neutralization by Mg2+ from 25 to 50% led to a two-fold reduction in REE. Further analysis revealed that Mg2+ ions do not compact PAR in a homogenous manner; instead, they induce the formation of highly compacted globules separated by extended polymer chains (Fig. 2F). This compaction was visualized over time across different systems using heatmaps that depict the number of neighboring ADP-ribose units within a 10 Å radius of each PAR residue (Supplementary Fig. 1). Counting the net charge within this radius over time further revealed the interplay between spatial localization and electrolyte charge compensation (Supplementary Fig. 2). Interestingly, the globular conformations observed at a neutralizing concentration of Mg2+ (100%, 7 mM) closely resemble those at much higher MgCl2 concentrations (50 mM). These data indicate that PAR22 is optimized for compaction even at Mg2+ concentrations much closer to physiological levels. This high sensitivity of PAR to Mg2+, including the formation of locally compacted domains (Supplementary Movies 1 and 2), suggests that Mg2+ may play a structural role. Beyond simply screening electrostatic charges, Mg2+ may trigger a transition of PAR from an elongated polymer to a condensed globule.
To further quantify the effect of adding Mg2+, we conducted simulations with a fixed NaCl concentration of 50 mM while varying the MgCl2 concentration (Fig. 2G). These conditions align with those previously examined through smFRET experiments30. Our simulation showed that, in the presence of Mg2+, PAR transitioned from extended to compacted states and exhibited conformations where both states coexist within the same molecule (Fig. 2H, Supplementary Figs. S1, S2). When compared to simulations in pure NaCl solvent (Fig. 2C, D), the onset of compact conformations occurred at significantly lower MgCl2 concentrations. Specifically, a complete globular collapse occurred at 21 mM of MgCl2, in contrast to 2 M NaCl. Furthermore, over 39% of total compaction was achieved at a MgCl2 concentration as low as 3 mM. These observations are consistent with previous studies on single-stranded RNA, where only 5 mM MgCl2 was required to induce the same Rg change as 600 mM NaCl31.
Taken together, our MD simulations show that PAR is structurally dynamic, adopting a range of conformations depending on the ionic environment. In the absence of Mg2+, PAR adopts extended conformations at physiological Na+ concentrations. However, even small amounts of Mg2+ can trigger local compaction of the PAR polymer—a structural transition that we further explored in the remainder of this study.
SAXS reveals distinct compaction for PAR15 and PAR22 with Mg2+
Having surveyed a broad range of ionic conditions in the MD simulations of PAR, we focused on specific conditions for experimentally identifying various structural parameters of PAR using SAXS. We examined PAR15 and PAR22, both of which are found in normal and cancer cells, with the shorter one being more abundant24,25,26. The SAXS experiments were conducted in a 100 mM NaCl environment to approach physiological conditions, and we assessed the impact of adding 1 mM MgCl2 on PAR compaction.
SAXS provides us with the overall size of the PAR structural ensembles, represented by the Rg values (Fig. 3A, B)33. In a 100 mM NaCl solution, PAR15 had an Rg of 24.4 ± 1.2 Å, while PAR22 had an Rg of 32.6 ± 0.5 Å (Fig. 3B). Adding 1 mM Mg2+ led to a 1.3 Å, or 5.3%, reduction in Rg for PAR15 and a larger 6.2 Å, or 19.0%, reduction for PAR22, indicating a greater compacting effect on the longer PAR22 molecule.
A SAXS profiles of PAR15 and PAR22 in 100 mM NaCl with (red) and without (blue) the addition of 1 mM MgCl2. Data are plotted in dimensionless Kratky axes, normalizing out size differences and emphasizing changes in shape and disorder in the mid-angle scattering regime. Experimental scattering is shown in light-colored points, and solid lines show molecular form factor (MFF) model fits to the data, extracting the Flory scaling parameter ν (Supplementary Fig. 3)34. Error bars are derived from experimental error and rebinning. B SAXS-derived Rg values for PAR15 and PAR22 in the conditions assayed. Error bars show errors in the linear Guinier fits used to extract Rg. (Supplementary Fig. 4) C Radius of gyration of PAR15 and PAR22 polymers in MD simulations carried out at 100 mM NaCl, with and without 1 mM MgCl2. The histograms next to the timeseries plots illustrate the distribution of the Rg values. D Average simulated radius of gyration of PAR15 and PAR22 determined as a weighted mean ± square root of the weighted variance of the two Gaussian fit to the histograms. SAXS source data are provided as a Source Data file.
We also calculated the Flory (ν) parameters for both PAR lengths at each salt concentration to understand their interaction with its surrounding solvent (Fig. 3A, Supplementary Fig. S3)34. Kratky plots were used for clearer data visualization, emphasizing the mid-angle scattering regime where the overall shape and degree of disorder of the molecular ensemble can be discerned35.
When Mg2+ was absent, fits to SAXS profiles for both PAR15 and PAR22 yielded a ν value of 0.60 ± 0.02 and 0.60 ± 0.01, respectively. This ν value is expected for a self-avoiding random walk, suggesting similar polymer properties for both PAR lengths in a 100 mM Na+ environment. The ratio of Rg values between PAR22 and PAR15 (32.6 Å / 24.4 Å = 1.34) is consistent with the expected behavior for molecules with ν = 0.6, according to the classical scaling law, Rg α lengthν. This agreement provides additional confidence in this polymer description of PAR.
When Mg2+ was introduced, PAR22 (ν = 0.55 ± 0.01) underwent a significant decrease in ν while PAR15 (ν = 0.59 ± 0.02) did not. These changes point to a stronger partial compaction of PAR22 (Fig. 3A, Supplementary Fig. S3). Overall, these experimental results indicate that the longer PAR22 undergoes a more significant conformational change when exposed to Mg2+ compared to its shorter counterpart, PAR15 (Fig. 3B).
MD-SAXS reveals Mg2+ increases tortuosity and base stacking more in PAR22 than PAR15
To examine specific conformations in PAR responsible for the observed differences in size and shape, we integrated SAXS data with additional MD simulations. Specifically, we simulated the two PAR22 systems, both containing 100 mM NaCl electrolyte and differing by the presence of 1 mM MgCl2, for 1 µs each. In addition, we built and simulated, also for 1 µs each, two complementary PAR15 systems. All four systems contained about 160,000 atoms and initially occupied a volume of 12 × 12 × 12 nm3. During these simulations, we noticed that the PAR conformations transitioned between extended and compact states (Fig. 3C). Because of this bimodal behavior, and since the duration of simulation duration was comparable to the lifetime of each state, we were unable to determine the average Rg values for direct comparison (i.e., a large error margin when averaging Rg values across the entire simulations; Fig. 3D). As obtaining a microsecond duration trajectory of a 160,000 atom-system is, at the time of writing, at the practical limit of the MD methodology, we employed an ensemble optimization method (EOM) to refine the full pool of MD structures using SAXS data36,37. The computed scattering profiles from the refined ensembles closely matched the experimentally measured SAXS profiles in molecular shapes and overall Rg values (Figs. 4A, B, Supplementary Figs. S5, 6).
As an example, the case for PAR15 in 100 mM NaCl is shown. The same plots for PAR22 are shown in Supplementary Figs. 5, 6. A The pool of structures from the entire MD simulation is shown in orange, and the subset ensemble that agrees with the SAXS data in blue. Structures are parameterized in {Rg,REE} space. 1D histograms are weighted by the prevalence of each structure in the final ensembles. B Final agreement of the structural ensemble determined by EOM (blue) to the SAXS data (gray), compared to initial agreement of the structural ensemble of all MD conformers (orange). Gray error bars represent experimental errors. Residuals are shown in the bottom plot. C Ensembles of EOM-determined PAR structures, with and without Mg2+ for PAR15 and PAR22. Arrows show differential shifts to more compact states with the addition of Mg2+. D Tortuosities and E Fraction of adenine bases that are stacked in each structural ensemble of PAR, calculated as a weighted mean across the ensemble. For the tortuosity box-and-whisker plot in (D), the center mark is the medium and the box edges are the 25th and 75th percentiles; points outside the whisker edges are outliers (>2.7 S.D. from the mean). To gauge differences between groups, a two-sample two-sided t-test assuming unequal variances was performed, and ‘*’ denotes p < 0.05. Exact p-values: PAR15 100 mM NaCl vs PAR15 100 mM NaCl + 1 mM MgCl2: p = 0.1862, PAR22 100 mM NaCl vs PAR22 100 mM NaCl + 1 mM MgCl2: p = 5.969E-7, PAR15 100 mM NaCl vs PAR22 100 mM NaCl: p = 0.0027, PAR15 100 mM NaCl + 1 mM MgCl2 vs PAR22 100 mM NaCl + 1 mM MgCl2: p = 7.707E-11. For the base stacking plot in E, error bars show standard error. Number of structures in each refined ensemble: PAR15, 100 mM NaCl: 225; PAR15, 100 mM NaCl + 1 mM MgCl2: 563; PAR22, 100 mM NaCl: 75; PAR22, 100 mM NaCl + 1 mM MgCl2: 78. Note that, while the PAR15 pools have more unique structures, the weights of each structure are higher in the PAR22 pools (weights sum to 1000 in all pools).
Next, we analyzed how PAR conformation and compaction change with Mg2+ for both PAR15 and PAR22 (Fig. 4C). The refined pools displayed a relatively uniform distribution of structures around an average, with no pronounced bimodality, as one would expect for a macromolecular ensemble (Supplementary Fig. 5). Importantly, the final fits between the EOM structural ensembles and SAXS data consistently fell within the experimental error margin, with ensemble Rg values in agreement across both methods (Supplementary Fig. 6). Supplementary Fig. 5 also lists the number of unique structures identified in each refined pool.
With refined ensembles now available for all conditions (PAR15 and PAR22, both with and without Mg2+), we analyzed the included structures. We computed the tortuosity index of each structure in the identified ensemble to gauge their backbone conformations (Fig. 4D; see “Methods” for calculation method). Tortuosity quantifies how “twisted” the polymer is compared to a straight line connecting its endpoints. Without Mg2+, PAR22 has a significantly greater mean tortuosity across its structural ensemble than PAR15, indicating a more twisted backbone (Fig. 4D), despite the identical ν values. Interestingly, introducing Mg2+ significantly increased the tortuosity of PAR22, but not PAR15 (Fig. 4D).
We also examined the role of π-π stacking in driving PAR chain compaction (Fig. 4E; see “Methods” for calculation method). Such interactions are particularly common among adenine bases and are known to induce intra-chain helicity in adenine-rich RNA sequences31. In the presence of 1 mM MgCl2, PAR15 underwent a 24% increase in base stacking events, whereas PAR22 exhibited a 103% increase (Fig. 4E). This greater frequency of π-π interactions between adenine bases could contribute to the greater compaction of PAR22 compared to PAR15.
PAR22 displays ADP-ribose bundles
To further analyze the structural ensembles revealed by EOM, we parameterized the structures according to their {Rg,REE} values and performed hierarchical clustering38. Using this approach, we found that the ensembles are highly heterogeneous: Rg and REE values covered 30 Å and 100 Å ranges, respectively. Through hierarchical clustering, we grouped structures into clusters with similar size, ranging from highly extended to highly compact (Supplementary Fig. 7).
To elucidate unique conformational features within these clusters, we computed heatmaps of pairwise distances between bases in EOM-selected structure ensembles (Supplementary Fig. 8). These heatmaps revealed how ADP-ribose bases are connected along the PAR chains. In these maps, off-diagonal regions with shorter distance implies the crowding of distal bases. For PAR15, the heatmap revealed proximity mainly along the diagonal of the heatmap. This trend progressed monotonically toward the corners, suggesting that the molecule predominantly adopts relatively featureless extended conformations, irrespective of the presence or absence of MgCl2 (Supplementary Fig. 8A). In contrast, PAR22’s heatmap showed significant connections, or close contacts, between bases that are close together (blue regions, slightly off-diagonal). One such region appeared in the 100 mM NaCl map (upper left corner), while two were evident when 1 mM MgCl2 was added (upper left and lower right corners, Supplementary Fig. 8B). These off-diagonal regions indicate a local bundle of non-adjacent bases at the end(s) of the molecule, corroborating findings of local compaction initially identified in our MD simulations (Fig. 2).
We next considered the role of π-π stacking in these distinct ensembles. For PAR15, regions of proximity (i.e., low inter-base distance) correlated somewhat with where base stacking occurs, mainly along the diagonals (Supplementary Fig. 8A). Yet, for PAR22, these off-diagonal low-distance regions were not enriched with base stacks (Supplementary Fig. 8B). Most stacking events occurred between adjacent bases along the chain. Thus, while PAR22 has more base stacking with 1 mM Mg2+ than PAR15 in general (Fig. 4E), the observed ADP-ribose bundles appear unrelated to base stacking. Rather, local chain compaction due to the proximity of the PAR phosphate backbone to an Mg2+ ion likely triggers the intra-chain coil-to-globule transitions that lead to these bundles. This transition is evident in individual frames of the MD simulations, where the compaction correlates to some extent with proximity of Mg ions to the backbone (Supplementary Fig. 9, Supplementary Movie 2). Taken together, our analyses on the heterogeneous structural ensembles confirm the presence of ADP-ribose bundle formation unique to PAR22.
Distinct backbone conformations for PAR15 and PAR22
Hierarchical clustering partitioned the structural ensembles into groups based on the overall size of each structure; however, deriving a more concise description of the PAR backbone conformations was challenging due to a variety of poorly related structures populating in any size subgroup (Supplementary Fig. 7). Inspired by 2D classification of structures in single particle cryo-electron microscopy39, and graph theory, we grouped the ensembles into unique, interrelated conformational subclasses (Box 1). By applying 3D spatial alignment into network graphs and performing spectral clustering, we captured all conformations present in the ensembles, with no graph outliers (Supplementary Fig. 10). The low spatial variation between the constituent structures of these subclasses (Fig. 5 and Supplementary Fig. 11) supports that our algorithm effectively identified sensible classes, justifying the subsequent averaging to depict a single representative conformation in each class.
A–D PAR15 and PAR22 in 100 mM NaCl, both without and with the presence of 1 mM MgCl2, is shown. Top plots show the graphs of PAR structures in each ensemble, color-coded by the clusters identified by K-means. The box colors around each identified subclass match their locations in the graphs. Wireframe models show the mean PAR backbone conformation in each case—each dot represents the mean position of each pair of phosphorus atoms across the entire subclass, colored by the degree of spatial variance present across that class. Red squares denote the 1″ ends of the backbones and red triangles denote the 2′ ends. The fraction of each structural subclass within the entire ensemble is shown adjacent to the respective averaged backbone conformer models. E, F Proposed model of a critical length for coil-to-globule transitions in PAR in the presence of MgCl2, linking to previously observed differences in binding and condensing certain proteins16. Above a certain length, potentially between 15 and 22 subunits, PAR forms ADP-ribose bundles that impose super-anion functionality, accumulating negative charge and giving PAR a disproportionate amount of electrostatic potential. In longer PAR chains, these bundles may periodically appear along the chain, similar to the beads-on-a-string model of classical polymer theory.
Without MgCl2 in 100 mM NaCl, the subclasses identified for PAR15 largely displayed similar conformations, with the backbone predominantly bent slightly into an inverted U shape (Fig. 5A). PAR22 exhibited similar U-shaped bends, but with additional variations: 21.3% of its conformations were more extended (Fig. 5B, green) and 16.0% more twisted (turquoise), likely contributing to the observed increase in tortuosity (Fig. 4E). Notably, bundles of ADP-ribose units were observed in PAR22 at the 1″ ends of each subclass (Fig. 5B), visually confirming our pairwise distance measurements between bases (Supplementary Fig. 8).
The introduction of 1 mM MgCl2 accentuated the conformational differences between PAR15 and PAR22. The structural ensembles at 100 mM NaCl of both PARs were generally less connected, exhibiting greater distances between pairs of structures (Supplementary Fig. 10A, B). However, the presence of Mg2+ led to greater similarity among the structures within the ensemble, as evidenced by the increased number of structures demonstrating low root mean square deviations in pairwise comparisons (Supplementary Fig. 10C, D). Specifically, in PAR22, the occurrence of ADP-ribose bundles was now noted in all five of the identified subclasses, spanning those with extended and more compact conformations (Fig. 5D). This observation is consistent with the heatmap indicating an increase in the number of regions that have short pairwise distance between bases (Supplementary Fig. 8). The consistent low spatial variance (<20 Å) in these bundle regions across all subclasses further indicates the systematic presence of bundles throughout the structural ensemble (Fig. 5D). Each bundle contained ∼8 ADP-ribose units at each end, interconnected by 6 additional units.
In contrast, such bundles were not present systematically enough in the PAR15 ensemble to be coherently observed with 1 mM MgCl2. These conformations closely resembled PAR15 in 100 mM NaCl alone, exhibiting similar backbone bending. A small fraction (4.6%) of the structures collapsed into a globule (Fig. 5C, yellow), akin to the most compact conformation observed in our initial MD simulations (Fig. 2). This subclass of collapsed globules may account for the slight 5.3% decrease in Rg in PAR15 as observed through SAXS (Fig. 3B). These data imply that only a small subset of molecules could undergo relatively featureless collapse with the small amount of Mg2+ present, leaving the rest of the ensemble relatively uninfluenced. On the other hand, the widespread bundling of the ADP-ribose units in PAR22 could explain the larger 19.0% decrease in its Rg (Fig. 3B). The difference in ADP-ribose bundle appearance alludes to a model of PAR’s length-dependent function (Fig. 5E, F).
PAR has less helicity and base stacking than poly-adenosine RNA
Our characterization of PAR and its distinct structural features prompted us to draw a comparison with poly-adenosine RNA. Though composed of the same ribose, phosphate, and adenine base building blocks, poly-A RNA and PAR have vastly different cellular functions. The former largely acts as a termination signal and binding motif, while the latter functions as a flexible binding scaffold. To delve into the structural difference between these two nucleic acids likely tied to their divergent functions, we compared a 15-mer of ADP-ribose (PAR15) to a 30-mer of AMP (rA30) RNA. Both molecules were measured with SAXS in identical solutions containing 100 mM NaCl. Because ADP-ribose (in PAR) contains twice the number of phosphate and ribose groups as AMP (in RNA), these two macromolecules have comparable length and overall charge. Their Rg values further affirmed their similarity (Fig. 6A). Importantly, we also chose PAR15 for comparison due to its lack of ADP-ribose bundles (Fig. 5), a feature not known to be present in poly-A RNA.
A SAXS-derived Rg of PAR15 vs rA30. Error bars represent errors in the Guinier fits. B Mean fraction of adenine bases that are stacked in the PAR15 and rA30 structural ensembles. C Ensemble-averaged orientation correlation functions of PAR15, compared to that of rA30. D Mean correlation lengths of PAR15 vs rA30, computed across the structural ensembles. E Four structures from the conformational ensemble of PAR15 that are most highly selected by EOM. F Four representative rA30 structures; accessible via SASDFB9 in the Small Angle Scattering Biological Data Bank31. Throughout this figure, blue refers to PAR15, and green refers to rA30. For (B–D), Error bars represent the variance in the datasets and are derived from analysis of N = 20 poly-A RNA structures (constituting the pool of structures from SASDFB9) and N = 225 PAR15 structures (constituting the pool of structures in the current study).
The orientational correlation function (OCF) can describe the orientational alignment of local regions of a polymer chain as a function of the distance between its monomers (|i-j|). Peaks in OCF signify high periodic orientational directionality, while a featureless exponential decay represents random chain orientations40. Previous structural characterization of rA30 has revealed its well-ordered helix form, attributed to the propensity of adenine bases to undergo π-π interactions. The helix formation is driven by an extensive base stacking network, with 85.2 ± 2.5% of the bases adopting a parallel stacked configuration (Fig. 6B)31. The OCF of rA30 displayed strong oscillatory behavior with peaks spaced by the periodicity of an A-form helix (Fig. 6C). In contrast, PAR15 exhibited less orientational correlation along the chain, with the exponential decay of its OCF more closely resembled that of a random coil at |i-j| > 4 (Fig. 6C)40. This difference suggests that the bases in PAR are more randomly arranged than in RNA, supported by the mean PAR correlation length (6.1 ± 0.3 Å) being only a third of that of poly-A RNA (18.7 ± 0.2 Å) (Fig. 6D). The correlation length, which is greater when repeating backbone orientations are present, reflects the lower degree of order for PAR. Moreover, less than 14.4 ± 0.9% of the bases in PAR15 were stacked (Fig. 6B), preventing π-π interactions from stabilizing an ordered helical conformation, as is observed in rA30. The additional phosphate group and ribose sugar between each adenine base in PAR, compared to poly-A, may place the bases too far apart for extensive π-π base stacking interactions. Such differences in helicity and base stacking were evidently observed in individual sample conformers (Fig. 6E, F).
Discussion
The building blocks of PAR are configured uniquely compared to other nucleic acids, potentially contributing to its distinct functional properties. Previously, we have shown that PAR possesses a larger persistence length than RNA, thereby being stiffer—a characteristic attributed to the different distribution of the phosphates in PAR relative to RNA30. Both PAR and RNA assume more compact states with increasing salt concentration—a compaction that occurs with 100x fewer divalent than monovalent ions30. Building on this polymeric characterization, we set out to explore the 3D structure of PAR. Our studies reveal that the structure of PAR markedly diverges from that of poly-A RNA. In contrast to RNA, PAR possesses a shorter correlation length, exhibits less π-π stacking between bases, and adopts a less helical structure (Fig. 6). These findings are aligned with previous studies showing that PAR, when exposed to only monovalent ions at room temperature, lacks a well-defined secondary structure17,18,19,20,21. While the two delocalized rings in adenine make them especially prone to undergo a π-π interaction, the enhanced electrostatic repulsion in PAR, relative to RNA, could also prevent bases from achieving the close proximity needed to form such an interaction. This distinction may align with PAR’s enhanced ability, compared to RNA, to bind positive ligands, both in 1:1 interactions and large-scale condensates3,11,12,16.
PAR has historically presented challenges for structural characterization. The few published studies on PAR structure suggested that it lacks a well-defined structure but may have some subtle structural features17,21,22. However, a common limitation among these studies is their reliance on data derived from mixtures of PAR lengths, potentially masking signals from individual lengths. In this study, we have synthesized homogenous, single-length PARs for structural investigation. We have integrated structures generated through PAR15 and PAR22 MD simulations with experimental data offered by SAXS. With this approach, we can ascertain and analyze conceivable heterogeneous ensembles of structures, aiming to comprehend not just how PAR interacts with the surrounding ions but also the impact of length on its structural heterogeneity.
SAXS data revealed that PAR22 compacts more than PAR15 with Mg2+ (Fig. 3). Analysis of order parameters in the SAXS/MD-derived structural ensembles further elucidates differences in Mg2+-induced PAR compaction between 15- and 22-mer. The additional compaction observed in the latter is driven by a more tortuous backbone (Fig. 4D) and increased stacking among adenine bases (Fig. 4E). These phenomena are mediated by Mg2+-induced transient contacts with the PAR phosphate backbone, causing local distortion and leading to the more compact configuration observed in the MD trajectories (Supplementary Fig. 9, Supplementary Movies 1 and 2).
By examining subclasses of the full structural ensembles, we characterized the presence of ADP-ribose bundles unique to PAR22 (Fig. 5). This phenomenon aligns with a prior MD study that observed multiglobular behavior for a 25-mer, based on the dihedral constraints of the molecule22. However, the previous study, using improper force fields without corrections for ions, resulted in more compact PAR structures than realistically expected. Our 1 µs-simulations demonstrate that such compact states can be long-lasting (50–100 ns), oscillating transiently between compact and extended states (Supplementary Fig. 9). This behavior, especially under conditions with divalent cations or at higher monovalent cation concentrations, was not observed in the previous study due to its methodological limitations. Specifically, the heightened complexity inherent in simulations featuring divalent cations, like Mg2+ ions, was not addressed, underscoring the significance of our study in revealing the dynamic nature of PAR structures.
Our approach, combining experimental data with MD simulation, reveals that these ADP-ribose bundles are seeded by the presence of Mg2+ (Supplementary Fig. 9), and these features are less pronounced in PAR15. Once formed, the structural stability of these bundles does not seem to rely on adenine base stacking or direct association with divalent Mg2+ ions alone. Instead, Na+ ions within the bundle appear to play a stabilizing role (Supplementary Movie 2). We speculate that a minimum number of ADP-ribose units are required to form a stable bundle—a critical length not attained by PAR15—possibly due to insufficient length between bundles to adequately separate congregated negative charges. Ensemble-level pairwise base distance heatmaps support the observation that globular bundles of ADP-ribose units are diffusely present across the entire ensemble in PAR22, but not in PAR15 (Supplementary Fig. 8). It is conceivable that in longer lengths of PAR (>50 mer), multiglobular ADP-ribose bundles could periodically appear along the chain, consistent with the beads-on-a-string models of classical polymer theory41,42.
Based on these findings, we propose a model that addresses unresolved questions concerning PAR’s diverse interactions with its binding partners. The model suggests the formation of globular ADP-ribose bundles beyond a crucial length, which falls between 15 and 22 units. These bundles appear to form along the chain through an intramolecular coil-to-globule phase transition induced by divalent Mg2+ cations dynamically coming into close contact (Supplementary Fig. 9), wherein PAR of sufficient length can have part of its chain as a collapsed globule and the rest in a more extended state. In addition to this effect of Mg2+, an elevated Na+ presence around PAR is observed (Fig. 2D, F, H, Supplementary Movies 1–4). These phenomena were characterized through our MD simulations, details that would have been overlooked by simply observing the mean field behavior of the system. At physiological cationic conditions, we observe that PAR undergoes this transition rapidly on the sub-microsecond timescale. We speculate that these bundles could be of variable size, thereby granting PAR more conformational variety in molecular recognition of binding partners (Fig. 5E, F)17. Additionally, these bundles localize the negative charge within the molecule, forming super-anion beads-on-a-string in a length-dependent manner which could afford enhanced condensate formation (Fig. 5E, F)16. Follow-up studies are warranted to test these hypotheses.
Our focus has been on characterizing linear PAR—a choice driven by the current capability in the field to synthesize sufficient, pure quantities of this single defined-length molecule, essential for the interpretation of SAXS experiments. Exploring branched PAR, another physiological structural form, is not technically feasible at this juncture. Moreover, synthesizing and simulating specific lengths of long PAR—potentially reaching up to 200-mer in cells—poses notable challenges. Under physiological conditions, PAR is primarily covalently conjugated to proteins, rather than existing as freely diffusing molecules. Given that proteins are conjugated to the 1″ ends of PAR, we postulate that ADP-ribose bundles might form in the middle of a chain, possibly exhibiting a periodicity of 15–22 units, rather than exclusively at the ends. Future systematic studies on PAR of different lengths, structures, and conjugations to proteins are essential for a more comprehensive understanding of the structure and function of PAR.
In this study, we characterized the structure of PAR15 and PAR22 by performing MD in conjunction with SAXS and carrying out detailed analyses on the resulting conformational ensembles. We showed that PAR rapidly compacts with increasing ionic strength and that this compaction occurs differently in the two different PAR lengths. The structural ensembles of PAR were found to be highly heterogeneous. By breaking down this heterogeneity through biophysical parameters and real space class averaging, we characterized the conformational mechanism for this difference in compaction, identifying globular bundles of ADP-ribose unique to PAR22. We speculate that this structural feature may enable PAR to bind ligands specifically, forming part of the PAR code43.
Methods
PAR sample preparation
Monodisperse samples of PAR15 and PAR22 were produced in three steps.
Step 1 (bulk PAR preparation): PAR was produced as a multi-length mixture (bulk PAR) via an enzymatic synthesis. Briefly, 250 nM recombinant PARP1 (full length)44, 5 µM PARP1 (379–1014)45, 50 mM HEPES pH 7.0, 10 mM MgCl2, 0.1% (v/v) NP-40, 1 mM DTT, 1 mM NAD+ and 0.3 µM oligonucleotide duplex GGAATTCC in 32 mL reaction volume in a 50 mL DNA Lo-Bind tube were incubated at 37 °C for 1 h. The reaction was then quenched with 8 mL of 50% (w/v) ice-cold TCA and incubated at 4 °C for 15 min. The sample was centrifuged at 24,000×g for 10 min, and the insoluble material was washed with 30 mL of ice-cold 75% ethanol before being dried at 37 °C for 5 min. The PAR was enzymatically released by resuspension of pellet in a neutral solution of 10 mL of 2 mg/mL proteinase K, 400 mM hydroxylamine pH 7.0, 0.5 % (w/v) SDS, 10 mM EDTA pH 8.0, 100 mM MES pH 6.0 and incubated at 37 °C at 1000 rpm for 2 h. The PAR was purified by ethanol precipitation by the addition of 25 mL ice-cold 75% ethanol and incubation at −80 °C for 1 h. The PAR was centrifuged at 24,000×g for 30 min at 4 °C and washed with 25 mL of ice-cold 75% ethanol. The PAR was dried at 37 °C for 5 min before resuspension in 1 mL of 10 mM Tris pH 7.0, 1 mM EDTA. The concentration of bulk PAR was estimated by absorbance at 258 nm with \({{{\rm{\varepsilon }}}}_{{ADPr}}=13500\) M−1cm−1.
Step 2 (1″ alkyne- or biotin modification): A 2 mL reaction containing 15 mM bulk PAR, 100 mM potassium acetate pH 4.6, 100 mM aniline, and 15 mM alkyne-(Broadpharm # BP-23164) or biotin-PEG-oxyamine (Broadpharm # BP-22179) was incubated at 21 °C for 16 h with shaking (1400 rpm).
Step 3 (PAR fractionation): 2 mL of the filtered labeled PAR was injected in DNApac-PA100 (22 × 250 mM) using mobile phase A (25 mM Tris buffer pH 9) and mobile phase B (25 mM Tris buffer pH 9 + 1 M NaCl) and fractionated at 5 mL/min by the following gradient program: 0–6 min 0%, 6–10 min 30%, 10–60 min 40%, 60–78 min 50%, 78–108 min 56%, 108–112 min, 100%, 112–114 min 100%. The defined-length PAR was concentrated and buffer-exchanged with water using a 15 mL centrifugal filter with a 3000 MWCO. The concentration of PARn was measured by absorbance at 258 nm following this formula [PARn] (M) \(=\frac{{{\rm{A}}}258}{{{\rm{\varepsilon}}} _{{PAR}}},\,{{{\rm{\varepsilon }}}}_{{PAR}}=({n\times \varepsilon }_{{ADPr}})\).
The purity of PAR15 and PAR22 are ~85%, with impurities mostly within ±1 ADP-ribose unit, as judged based on gels stained by SYBR gold. Additionally, the PAR was analyzed by electrospray ionization mass spectrometry for molecular weight confirmation (Supplementary Fig. 12). In the molecules used for SAXS experiments, the first ADP-ribose is either attached with biotin (PAR22) or alkyne (PAR15) at the 1″ termini. However, it should be noted that such different attachment did not result in SAXS measurement or the resultant Rg values obtained, with or without Mg2+ (Supplementary Fig. 12).
Prior to solution scattering experiments, PAR samples were concentrated and buffer-exchanged into 100 mM NaCl, 20 mM Tris-HCl (pH 7.4) using 3k MWCO Amicon microcentrifuge spin columns (Millipore Sigma, St. Louis MO, USA), using six centrifugation steps of 14,000×g for 15 min, maintaining 4 °C temperature. Samples were then annealed by heating to 90 °C for 5 min and snap cooling at 4 °C for 20 min, then stored on ice until SAXS data collection. For samples requiring Mg2+, the final concentration was spiked to 1 mM MgCl2 by mixing in a 1 M MgCl2 stock solution minutes prior to SAXS measurements.
Molecular dynamics (MD) simulations
All MD simulations were performed using NAMD2.1446, the CHARMM36 parameter set for protein and DNA47, TIP3P water model48, and a custom hexahydrate model for magnesium ions along with the CUFIX corrections to ion–nucleic acid interactions23. Multiple time stepping was used: local interactions were computed every 2 fs, whereas long-range interactions were computed every 6 fs49. All short-range nonbonded interactions were cut off starting at 1 nm and completely cut off by 1.2 nm. Long-range electrostatic interactions were evaluated using the particle-mesh Ewald method computed over a 0.11 nm spaced grid50. SETTLE and RATTLE82 algorithms were applied to constrain covalent bonds to hydrogen in water and in non-water molecules, respectively51,52. The temperature was maintained at 300 K using a Langevin thermostat with a damping constant of 0.5 ps−1 unless specified otherwise. Constant pressure simulations employed a Nose-Hoover Langevin piston with a period and decay of 200 and 50 fs, respectively53. Energy minimization was carried out using the conjugate gradients method. Atomic coordinates were recorded every 9.6 ps unless specified otherwise. An example MD timeseries plot is shown in Fig. 2B. Visualization and analysis were performed using VMD and MDanalysis54,55,56.
CHARMM-compatible force field parameters for PAR were obtained by combining existing parameters for chemically similar moieties. Specifically, a custom patch was written to define a PAR residue (PAR), whereby the oxygen atom on the terminal phosphate group of an ADP molecule (atom O5D) was connected to the ribose sugar (atom C5D) using analogy from NADP. Similarly, a custom patch (BND) connected the ribose (atom O1D) with ADP (C2′). With these two patches, a PAR molecule of an arbitrary number of monomers could be defined. Separate patches were defined for the terminal atoms, a hydrogen on the O1D atom of the ribose (1TER) and an OH group on the C2′ atom of ADP (2TER). The topology and parameter files for a PAR residue are deposited at https://github.com/TongGeorgeWang/polyADPribose-Structural-Analysis/tree/main/MolecularDynamics.
Small-angle X-ray scattering (SAXS)
Solution SAXS experiments on PAR15 and PAR22 were performed at the ID7A1 BioSAXS beamline at the Cornell High Energy Synchrotron Source and the 16-ID Life Science X-ray Scattering beamline at the National Synchrotron Light Source II of Brookhaven National Laboratory57,58. Radial integration and data reduction were performed in BioXTAS RAW to obtain 1D scattering profiles, plotting scattering intensities (I, in arbitrary scattering units) as a function of momentum transfer (q, in Å−1)59. PAR was assayed at 60 µM, 40 µM, and 20 µM concentrations to observe for interparticle effects at low q, and these were corrected when present by linearly extrapolating each point at q < 0.05 Å−1 to the zero-concentration limit. Radii of gyration were obtained through Guinier analysis40, approximating the scattering profile at (q*Rg) < 1.3 as a Gaussian function:
Guinier analyses are shown in Supplementary Fig. 4. Data were plotted in the size-normalized Kratky format to emphasize changes in overall macromolecular shape. Beyond the Guinier regime, the SAXS data were fit to the function:
Where MFF is the molecular form factor model, whose parameters are Rg and ν, the Flory scaling parameter. The MFF is an empirical function that was derived by Riback et al. from computing the theoretical SAXS profiles of many disordered macromolecular ensembles with expected values of ν, based on the classical polymer scaling law R|i–j| ∝ |i – j|ν for intra-chain distances indexed by i and j34,60. It offers a way to quantify scattering changes in the region (q*Rg) > 1.3 by analyzing the continuum of ν∈[0.33,0.6], where the lower limit corresponds to a more compact globule and the upper limit corresponds to a disordered self-avoiding random walk34,60. MFF fits were performed using the SAXSonIDPs webserver34. More details on SAXS data acquisition are provided in Supplementary Table 1. Scattering data are deposited on the Small Angle Scattering Biological Data Bank (SASBDB) with the identifiers SASDSJ5, SASDSK5, SASDSL5, SASDSM5.
Determining PAR structural ensembles
To provide a diverse pool of structures for PAR, we sampled the MD trajectories in 1 nanosecond snapshots, obtaining a set of 1000–1200 individual structures over a broad, continuous conformational range. Supplementary Fig. 5 shows the structural ensembles parameterized in Cartesian space by their Rg and REE. CRYSOL v2.0 was used to compute the theoretical scattering profile of all structures in the SAXS regime, with a maximum order of harmonics of 15, Fibonacci grid order of 18, 0.3 Å−1 maximum scattering angle, and 61 calculated data points61.
The raw MD simulations were seen to approximate well the overall size of the conformational ensembles but not their overall shape. This is evidenced by the discrepancies between the summed theoretical scattering of all structures in the pool and the experimental scattering beyond the low q regime (q > 0.03 Å−1) (Supplementary Fig. 6). To provide an accurate depiction of the solution-state ensemble of structures that the molecules sample, the information content of the SAXS data was leveraged through EOM v2.1 (ATSAS, EMBL Hamburg, Germany)36,37. Briefly, subsets of the pool of structures were sampled, and the summated theoretical SAXS profile of the subset of structures was computed and fit to the experimental SAXS profile—the process was iterated until agreement with the experimental data was reached. A more detailed description of EOM is provided in the Supplementary Discussion. EOM was run over 1000 generations with 50 ensembles, 20 curves and 10 mutations per ensemble, over 100 iterations. This was found to yield convergence of the fit to the experimental SAXS data, with χ2 values of 0.1–0.4 and the SAXS profiles of the final pool post-EOM falling within error of the experimental measurement across the entire q range (Supplementary Fig. 6). Introducing more iterations into the algorithm beyond that executed was not found to improve convergence of the χ2 meaningfully, except for PAR22 in 100 mM NaCl where 10000 iterations were employed. Moreover, to assess the repeatability of EOM results across different runs, we showed that the selected structural pool is comparable within errors across five independent runs (Supplementary Fig. 13). The final ensembles were assessed to fall along a continuous distribution in their Rg, REE, and maximal dimensions, with no implausible bimodalities (Supplementary Fig. 5).
Structural order parameters to characterize PAR structures
To further describe the structural features of the determined PAR conformer ensembles and to compare them to the previously characterized behavior of poly-adenine RNA, we computed the tortuosities, number of base stacking events, and correlation lengths. These parameters were computed as weighted averages in each structural ensemble, with weights given by how many times each conformer was selected by the EOM algorithm. The coordinates of the average position of each pair of Phosphorus atoms and the Oxygen atom in between each pair of ribose sugars were sampled along the PAR chain to break it into even segments. Tortuosity indices (T) were computed as62:
where we take the sum of all n−1 bond vector angles ϴi along the sampled chain, n as the total number of ADP-ribose monomers, and divide by the end-to-end distance L. T was computed for all structures in the PAR ensembles and comparison of tortuosities were performed through a two-sample, two-sided t-test assuming unequal variances, with a p-value threshold of 0.05 considered significantly different.
The orientation correlation function (OCF) was employed to visualize structural features across the selected PAR conformer ensembles40. The OCF was calculated as:
Where r is the normalized bond vector between each pair of sampled coordinates. OCFs were plotted as a function of the distance between linkages |i-j|, for all {i,j} pairs. To get a metric of approximate polymer stiffness and how structured the chain is, correlation lengths (lOCF) were computed by summing across the OCFs:
Where b is the length of bonds between sampled coordinates. OCF curves were truncated to |i-j| = 20 for visualization—past this, greater OCF error made their comparison difficult.
Lastly, to determine the prevalence of π-π interactions in stabilizing the PAR chains, the number of base stacking events was computed for each structure. Adenine bases were indexed by sampling the coordinates of base atoms and fit using a plane. Pairs of base planes were considered stacked when the normal vectors to each plane were both <5 Å apart and approximately collinear, with <45° separation. These parameters were optimized such that they reproduced results from previously characterized structures of poly-A RNA that were derived from dinucleotide libraries with base stacking information built in.
All structural descriptors and order parameters were computed using in-house software written in MATLAB R2021a (Mathworks, Natick, MA, USA).
3D classification and averaging of disordered PAR ensembles
To break the multitudinous structural ensembles into a small subset of conformations, spectral analysis was performed to identify 4–5 unique conformational subclasses representative of the entire population. This allowed for visualization of the structural pool in real space in a practical manner. First, PAR structural ensembles were rotationally aligned using PyMOL v2.4.0 (Schrodinger, LLC), and the resulting root mean square deviation (RMSD) between each pair of aligned structures was input into an Nstructures × Nstructures matrix. Within this matrix, an RMSD threshold was set between 10 and 15 Å, wherein a pair of structures was declared to be connected if the RMSD between them was below threshold. In this way, an adjacency matrix was constructed, which was input into the spectral clustering algorithm of Ng, Jordan, and Weiss to map the structures to graph space wherein pairs of structures (nodes) are connected if they fall below the RMSD threshold63. Unique classes in the graph were then identified through K-means clustering in MATLAB, where it was found that 4–5 clusters sensibly categorized the node set. Upon classifying the PAR ensembles into classes, the structures were coarse-grained by sampling the average positions of each pair of phosphorus atoms along the backbone, further aligned by RMSD minimization, and averaged to determine the characteristic 3D chain conformation of each subclass. Supplementary Fig. 11 shows all structures overlayed with the average structure in each class, to showcase the validity of the method. This method, which we call Class Averaging via Spectral Analysis of Totally Disordered Macromolecules (CASA ToDiMo), is applicable to macromolecules beyond PAR (e.g., RNA, proteins). The algorithm is described in Box 1 and is available at https://github.com/TongGeorgeWang/CASA-ToDiMo.
Data analysis
All simulation systems were constructed and visualized using VMD 1.9.4a43. Custom code, either using Tcl 8.0 or Python 3.7.3, was used to analyze the MD trajectories. For reading the trajectory (DCD) files, either VMD (for Tcl) or MDAnalysis 1.1.1 (for Python) was used. Structural ensemble analysis, using MD trajectories as input, was performed using EOM v2.1 in ATSAS. Custom scripts written in MATLAB R2021 were used for the visualization of EOM results, hierarchical clustering, computing structural order parameters, and class averaging via spectral clustering. PyMOL v2.5.5 was used to visualize structures and for structural alignment during class averaging. SAXS data reduction and processing were done in BioXTAS RAW.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The solution scattering data of PAR along with EOM fits are available in the Small Angle Scattering Biological Data Bank (SASBDB, https://www.sasbdb.org/) with the identifiers SASDSJ5, SASDSK5, SASDSL5, SASDSM5. Source data, as well as the topology, parameter files, and system preparation scripts for the simulation, are available at https://github.com/TongGeorgeWang/polyADPribose-Structural-Analysis. Source data are provided with this paper.
Code availability
Computer scripts used for data analysis are available at https://github.com/TongGeorgeWang/polyADPribose-Structural-Analysis and https://github.com/TongGeorgeWang/CASA-ToDiMo.
References
Dasovich, M. & Leung, A. K. L. PARPs and ADP-ribosylation: deciphering the complexity with molecular tools. Mol. Cell 83, 1552–1572 (2023).
Lüscher, B. et al. ADP-ribosyltransferases, an update on function and nomenclature. FEBS J. 289, 7399–7410 (2022).
Gibson, B. A. & Kraus, W. L. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat. Rev. Mol. Cell Biol. 13, 411–424 (2012).
Schreiber, V., Dantzer, F., Ame, J.-C. & de Murcia, G. Poly(ADP-ribose): novel functions for an old molecule. Nat. Rev. Mol. Cell Biol. 7, 517–528 (2006).
Jin, X., Cao, X., Liu, S. & Liu, B. Functional roles of poly(ADP-ribose) in stress granule formation and dynamics. Front. Cell Dev. Biol. 9, 671780 (2021).
Leung, A. K. L. et al. Poly(ADP-ribose) regulates stress responses and microRNA activity in the cytoplasm. Mol. Cell 42, 489–499 (2011).
Fehr, A. R. et al. The impact of PARPs and ADP-ribosylation on inflammation and host-pathogen interactions. Genes Dev. 34, 341–359 (2020).
Alemasova, E. E. & Lavrik, O. I. A sePARate phase? Poly(ADP-ribose) versus RNA in the organization of biomolecular condensates. Nucleic Acids Res. 50, 10817–10838 (2022).
Rhine, K., Odeh, H. M., Shorter, J. & Myong, S. Regulation of biomolecular condensates by poly(ADP-ribose). Chem. Rev. 123, 9065–9093 (2023).
Reber, J. M. & Mangerich, A. Why structure and chain length matter: on the biological significance underlying the structural heterogeneity of poly(ADP-ribose). Nucleic Acids Res. 49, 8432–8448 (2021).
Löffler, T. et al. Influence of chain length and branching on poly(ADP-ribose)-protein interactions. Nucleic Acids Res. 51, 536–552 (2023).
Dasovich, M. et al. Identifying poly(ADP-ribose)-binding proteins with photoaffinity-based proteomics. J. Am. Chem. Soc. 143, 3037–3042 (2021).
Fahrer, J., Kranaster, R., Altmeyer, M., Marx, A. & Bürkle, A. Quantitative analysis of the binding affinity of poly(ADP-ribose) to specific binding proteins as a function of chain length. Nucleic Acids Res. 35, e143 (2007).
Kliza, K. W. et al. Reading ADP-ribosylation signaling using chemical biology and interaction proteomics. Mol. Cell 81, 4552–4567.e8 (2021).
Fahrer, J. et al. High-affinity interaction of poly(ADP-ribose) and the human DEK oncoprotein depends upon chain length. Biochemistry 49, 7119–7130 (2010).
Rhine, K. et al. Poly(ADP-ribose) drives condensation of FUS via a transient interaction. Mol. Cell 82, 969–985.e11 (2022).
Minaga, T. & Kun, E. Spectral analysis of the conformation of polyadenosine diphosphoribose. Evidence indicating secondary structure. J. Biol. Chem. 258, 725–730 (1983).
Barkauskaite, E. et al. Visualization of poly(ADP-ribose) bound to PARG reveals inherent balance between exo- and endo-glycohydrolase activities. Nat. Commun. 4, 2164 (2013).
Lambrecht, M. J. et al. Synthesis of dimeric ADP-ribose and its structure with human poly(ADP-ribose) glycohydrolase. J. Am. Chem. Soc. 137, 3558–3564 (2015).
Thirawatananond, P. et al. Structural analyses of NudT16-ADP-ribose complexes direct rational design of mutants with improved processing of poly(ADP-ribosyl)ated proteins. Sci. Rep. 9, 5940 (2019).
Schultheisz, H. L., Szymczyna, B. R. & Williamson, J. R. Enzymatic synthesis and structural characterization of 13C, 15N-poly(ADP-ribose). J. Am. Chem. Soc. 131, 14571–14578 (2009).
D’Annessa, I., Coletta, A. & Desideri, A. Geometrical constraints limiting the poly(ADP-ribose) conformation investigated by molecular dynamics simulation. Biopolymers 101, 78–86 (2014).
Yoo, J. & Aksimentiev, A. Improved parametrization of Li+, Na+, K+, and Mg2+ ions for all-atom molecular dynamics simulations of nucleic acid systems. J. Phys. Chem. Lett. 3, 45–50 (2012).
Kiehlbauch, C. C., Aboul-Ela, N., Jacobson, E. L., Ringer, D. P. & Jacobson, M. K. High resolution fractionation and characterization of ADP-ribose polymers. Anal. Biochem. 208, 26–34 (1993).
Kleczkowska, H. E., Malanga, M., Szumiel, I. & Althaus, F. R. Poly ADP-ribosylation in two L5178Y murine lymphoma sublines differentially sensitive to DNA-damaging agents. Int. J. Radiat. Biol. 78, 527–534 (2002).
Malanga, M. & Althaus, F. R. Poly(ADP-ribose) molecules formed during DNA repair in vivo. J. Biol. Chem. 269, 17691–17696 (1994).
Ando, Y. et al. ELTA: enzymatic labeling of terminal ADP-ribose. Mol. Cell 73, 845–856.e5 (2019).
Slade, D. et al. The structure and catalytic mechanism of a poly(ADP-ribose) glycohydrolase. Nature 477, 616–620 (2011).
Yoo, J. & Aksimentiev, A. New tricks for old dogs: improving the accuracy of biomolecular force fields by pair-specific corrections to non-bonded interactions. Phys. Chem. Chem. Phys. 20, 8432–8449 (2018).
Badiee, M. et al. Switch-like compaction of poly(ADP-ribose) upon cation binding. Proc. Natl Acad. Sci. USA 120, e2215068120 (2023).
Plumridge, A., Andresen, K. & Pollack, L. Visualizing disordered single-stranded RNA: connecting sequence, structure, and electrostatics. J. Am. Chem. Soc. 142, 109–119 (2020).
Chen, H. et al. Ionic strength-dependent persistence lengths of single-stranded RNA and DNA. Proc. Natl Acad. Sci. USA 109, 799–804 (2012).
Skou, S., Gillilan, R. E. & Ando, N. Synchrotron-based small-angle X-ray scattering of proteins in solution. Nat. Protoc. 9, 1727–1739 (2014).
Riback, J. A. et al. Innovative scattering analysis shows that hydrophobic disordered proteins are expanded in water. Science 358, 238–241 (2017).
Rambo, R. P. & Tainer, J. A. Characterizing flexible and intrinsically unstructured biological macromolecules by SAS using the Porod-Debye law. Biopolymers 95, 559–571 (2011).
Tria, G., Mertens, H. D. T., Kachala, M. & Svergun, D. I. Advanced ensemble modelling of flexible macromolecules using X-ray solution scattering. IUCrJ 2, 207–217 (2015).
Bernadó, P., Mylonas, E., Petoukhov, M. V., Blackledge, M. & Svergun, D. I. Structural characterization of flexible proteins using small-angle X-ray scattering. J. Am. Chem. Soc. 129, 5656–5664 (2007).
Baul, U., Chakraborty, D., Mugnai, M. L., Straub, J. E. & Thirumalai, D. Sequence effects on size, shape, and structural heterogeneity in intrinsically disordered proteins. J. Phys. Chem. B 123, 3462–3474 (2019).
Zhao, Z. & Singer, A. Rotationally invariant image representation for viewing direction classification in cryo-EM. J. Struct. Biol. 186, 153–166 (2014).
Ullner, M. & Woodward, C. E. Orientational correlation function and persistence lengths of flexible polyelectrolytes. Macromolecules 35, 1437–1445 (2002).
Schoon, T. G. F. Microstructure in solid polymers. Br. Polym. J. 2, 86–91 (1970).
Reches, M., Snyder, P. W. & Whitesides, G. M. Folding of electrostatically charged beads-on-a-string as an experimental realization of a theoretical model in polymer science. Proc. Natl Acad. Sci. USA 106, 17644–17649 (2009).
Leung, A. K. L. Poly(ADP-ribose): a dynamic trigger for biomolecular condensate formation. Trends Cell Biol. 30, 370–383 (2020).
Langelier, M.-F., Steffen, J. D., Riccio, A. A., McCauley, M. & Pascal, J. M. Purification of DNA damage-dependent PARPs from E. coli for structural and biochemical analysis. Methods Mol. Biol. 1608, 431–444 (2017).
Tan, E. S., Krukenberg, K. A. & Mitchison, T. J. Large-scale preparation and characterization of poly(ADP-ribose) and defined length polymers. Anal. Biochem. 428, 126–136 (2012).
Phillips, J. C. et al. Scalable molecular dynamics on CPU and GPU architectures with NAMD. J. Chem. Phys. 153, 044130 (2020).
Huang, J. et al. CHAR MM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Batcho, P. F., Case, D. A. & Schlick, T. Optimized particle-mesh Ewald/multiple-time step integration for molecular dynamics simulations. J. Chem. Phys. 115, 4003–4018 (2001).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Miyamoto, S. & Kollman, P. A. Settle: an analytical version of the SHAKE and RATTLE algorithm for rigid water models. J. Comput. Chem. 13, 952–962 (1992).
Andersen, H. C. Rattle: a “velocity” version of the shake algorithm for molecular dynamics calculations. J. Comput. Phys. 52, 24–34 (1983).
Feller, S. E., Zhang, Y., Pastor, R. W. & Brooks, B. R. Constant pressure molecular dynamics simulation: the Langevin piston method. J. Chem. Phys. 103, 4613–4621 (1995).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–8–27–8 (1996).
Gowers, R. et al. MDAnalysis: a Python package for the rapid analysis of molecular dynamics simulations. In Proc. 15th Python in Science Conference (SciPy, 2016).
Michaud-Agrawal, N., Denning, E. J., Woolf, T. B. & Beckstein, O. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 32, 2319–2327 (2011).
Yang, L. et al. Solution scattering at the Life Science X-ray Scattering (LiX) beamline. J. Synchrotron Radiat. 27, 804–812 (2020).
Acerbo, A. S., Cook, M. J. & Gillilan, R. E. Upgrade of MacCHESS facility for X-ray scattering of biological macromolecules in solution. J. Synchrotron Radiat. 22, 180–186 (2015).
Hopkins, J. B., Gillilan, R. E. & Skou, S. BioXTAS RAW: improvements to a free open-source program for small-angle X-ray scattering data reduction and analysis. J. Appl. Crystallogr. 50, 1545–1553 (2017).
Martin, E. W., Hopkins, J. B. & Mittag, T. Small-angle X-ray scattering experiments of monodisperse intrinsically disordered protein samples close to the solubility limit. Methods Enzymol. 646, 185–222 (2021).
Svergun, D., Barberato, C. & Koch, M. H. CRYSOL—a program to evaluate X-ray solution scattering of biological macromolecules from atomic coordinates. J. Appl. Crystallogr. 28, 768–773 (1995).
Bullitt, E., Gerig, G., Pizer, S. M., Lin, W. & Aylward, S. R. Measuring tortuosity of the intracerebral vasculature from MRA images. IEEE Trans. Med. Imaging 22, 1163–1171 (2003).
Ng, A., Jordan, M. I. & Weiss, Y. On spectral clustering: analysis and an algorithm. Adv. Neural Inf. Process. Syst. 849, 856 (2001).
Acknowledgements
We thank Drs. Shirish Chodankar, Richard Gillilan, Qingqiu Huang, and Suzette Pabit, as well as Qingyue Hu, Scout Fronhofer, and Sarah Uttormark, for their invaluable support with SAXS data collection. This work is based on research conducted at the Center for High-Energy X-ray Sciences (CHEXS), which is supported by the National Science Foundation (BIO, ENG and MPS Directorates) under award DMR-1829070, and the Macromolecular Diffraction at CHESS (MacCHESS) facility, which is supported by award 1-P30-GM124166-01A1 from the National Institutes of Health (NIH) and by New York State’s Empire State Development Corporation (NYSTAR). The LiX beamline is part of the Center for BioMolecular Structure (CBMS), which is primarily supported by S10 OD012331, P30GM133893, and by the DOE Office of Biological and Environmental Research KP1605010. As part of NSLS-II, a national user facility at Brookhaven National Laboratory, work performed at the CBMS is supported in part by the U.S. Department of Energy under contract number DE-SC0012704. The supercomputer time was provided by ACCESS allocation MCA05S028 and Leadership Resource Allocation MCB20012 on Frontera at the Texas Advanced Computing Center. Frontera is made possible by the National Science Foundation award OAC-1818253. This work was also supported by the NIH grants R01-GM137015 (A.A.), T32-CA009110 (M.B.), R01-GM104135 (A.K.L.L.), T32-GM080189 (M.R.M.), NSF GRFP (M.R.M.), R35-GM122514 (L.P.) as well as Viven Thomas Scholars Initiative (M.R.M.), and T.W. acknowledges the support of the Natural Sciences and Engineering Research Council of Canada (NSERC). Cette recherche a été financée par le Conseil de recherches en sciences naturelles et en génie du Canada (CRSNG).
Author information
Authors and Affiliations
Contributions
Conceptualization: A.K.L.L., L.P., A.A. Formal analysis: T.W., K.C. Investigation: T.W., K.C. Validation: T.W., M.R.M. Resources: M.B., M.R.M. Writing—Original Draft: A.K.L.L., L.P., T.W. Writing—Review & Editing: A.K.L.L., L.P., A.A., T.W., K.C., M.B., M.R.M. Supervision: A.K.L.L., L.P., A.A. Funding Acquisition: A.K.L.L., L.P., A.A.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Thomas Weikl and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, T., Coshic, K., Badiee, M. et al. Cation-induced intramolecular coil-to-globule transition in poly(ADP-ribose). Nat Commun 15, 7901 (2024). https://doi.org/10.1038/s41467-024-51972-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-51972-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.