Abstract
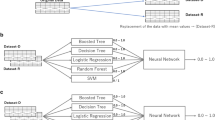
In the first trimester of pregnancy, accurately predicting the occurrence of pregnancy-induced hypertension (PIH) is important for both identifying high-risk women and adopting early intervention. In this study, we used four machine-learning models (LASSO logistic regression, random forest, backpropagation neural network, and support vector machines) to predict the occurrence of PIH in a prospective cohort. Candidate features for predicting the occurrence of middle and late PIH were acquired using a LASSO algorithm. The performance of predictive models was assessed using receiver operating characteristic analysis. Finally, a nomogram was established with the model scores, age, and nulliparity. Calibration, clinical usefulness, and internal validation were used to assess the performance of the nomogram. In the training set (2258 pregnant women), eleven candidate factors in the first trimester were significantly associated with the occurrence of PIH (P < 0.001 in the training set). Four models showed AUCs from 0.780 to 0.816 in the training set. For the validation set (939 pregnant women), AUCs varied from 0.516 to 0.795. The nomogram showed good discrimination, with an AUC of 0.847 (95% CI: 0.805–0.889) in the training set and 0.753 (95% CI: 0.653–0.853) in the validation set. Decision curve analysis suggested that the model was clinically useful. The model developed using LASSO logistic regression achieved the best performance in predicting the occurrence of PIH. The derived nomogram, which incorporates the model score and maternal risk factors, can be used to predict PIH in clinical practice.

We develop a model with good performance for clinical prediction of PIH in the first trimester.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Reddy S, Jim B. Hypertension and pregnancy: management and future risks. Adv Chronic Kidney Dis. 2019;26:137–45.
Wilkerson RG, Ogunbodede AC. Hypertensive disorders of pregnancy. Emerg Med Clin N Am. 2019;37:301–16.
Umesawa M, Kobashi G. Epidemiology of hypertensive disorders in pregnancy: prevalence, risk factors, predictors and prognosis. Hypertens Res. 2017;40:213–20.
Magee LA, Singer J, von Dadelszen P. Less-tight versus tight control of hypertension in pregnancy. N Engl J Med. 2015;372:2367–8.
Khan KS, Wojdyla D, Say L, Gulmezoglu AM, Van Look PF. WHO analysis of causes of maternal death: a systematic review. Lancet. 2006;367:1066–74.
Hutcheon JA, Lisonkova S, Joseph KS. Epidemiology of pre-eclampsia and the other hypertensive disorders of pregnancy. Best Pract Res Clin Obstet Gynaecol. 2011;25:391–403.
Zhuang C, Gao J, Liu J, Wang X, He J, Sun J, et al. Risk factors and potential protective factors of pregnancy-induced hypertension in China: a cross-sectional study. J Clin Hypertens. 2019;21:618–23.
von Dadelszen P, Magee LA. Pre-eclampsia: an update. Curr Hypertens Rep. 2014;16:454.
Magee LA, Pels A, Helewa M, Rey E, von Dadelszen P. The hypertensive disorders of pregnancy (29.3). Best Pract Res Clin Obstet Gynaecol. 2015;29:643–57.
Bergen NE, Schalekamp-Timmermans S, Roos-Hesselink J. Hypertensive disorders of pregnancy and subsequent maternal cardiovascular health. Eur J Epidemiol. 2018;33:763–71.
Roberge S, Nicolaides KH, Demers S, Villa P, Bujold E. Prevention of perinatal death and adverse perinatal outcome using low-dose aspirin: a meta-analysis. Ultrasound Obstet Gynecol. 2013;41:491–9.
Emmanuel B, Stéphanie R, Yves L. Prevention of preeclampsia and intrauterine growth restriction with aspirin started in early pregnancy: a meta-analysis. J Obstet Gynecol. 2010;116:402.
Redman CWG. Hypertension in pregnancy: the NICE guidelines. J Heart. 2011;97:1967–9.
Kenny LC, Black MA, Poston L. Early pregnancy prediction of preeclampsia in nulliparous women, combining clinical risk and biomarkers: the Screening for Pregnancy Endpoints (SCOPE) international cohort study. Hypertension. 2014;64:644–52.
Skråstad RB, Hov GG, Blaas HG, Romundstad PR, Salvesen KÅ. Risk assessment for preeclampsia in nulliparous women at 11-13 weeks gestational age: prospective evaluation of two algorithms. BJOG. 2016;122:1781–8.
Antwi E, Groenwold RH, Browne JL. Development and validation of a prediction model for gestational hypertension in a Ghanaian cohort. BMJ Open. 2017;7:e012670.
North RA, McCowan LM, Dekker GA, Poston L, Chan EH, Stewart AW, et al. Clinical risk prediction for pre-eclampsia in nulliparous women: development of model in international prospective cohort. BMJ 2011;342:d1875.
Poon LCY, Akolekar R, Lachmann R, Beta J, Nicolaides KH. Hypertensive disorders in pregnancy: screening by biophysical and biochemical markers at 11-13 weeks. Ultrasound Obstet Gynecol. 2010;35:662–70.
Odibo AO, Zhong Y, Goetzinger KR, Odibo L, Bick JL, Bower CR, et al. First-trimester placental protein 13, PAPP-A, uterine artery Doppler and maternal characteristics in the prediction of pre-eclampsia. Placenta. 2011;32:598–602.
Boldrini L, Bibault JE, Masciocchi C, Shen Y, Bittner MI. Deep Learning: A Review for the Radiation Oncologist. Front Oncol. 2019;9:977.
Breiman L. Random Forests. Machine Learning, 2001;45:5–32.
Huang MW, Chen CW, Lin WC. SVM and SVM ensembles in breast cancer prediction. PLoS ONE. 2017;12:e0161501.
Krizhevsky A, Sutskever I, Hinton G. ImageNet classification with deep convolutional neural networks. Advances in Neural Information Processing Systems (NIPS). 2012;25.1097–105.
Beam AL, Kohane IS. Translating artificial intelligence into clinical care. JAMA. 2016;316:2368–9.
Tibshirani R. Regression shrinkage and selection via the lasso: a retrospective. Journal of the Royal Statistical Society. 2011;73:267–88.
Oliveira N, Magder LS, Blitzer MG. First-trimester prediction of pre-eclampsia: external validity of algorithms in a prospectively enrolled cohort. Ultrasound Obstet Gynecol. 2014;44:279–85.
Farina A, Rapacchia G, Freni Sterrantino A, Pula G, Morano D, Rizzo N. Prospective evaluation of ultrasound and biochemical-based multivariable models for the prediction of late pre-eclampsia. Prenat Diagn. 2011;31:1147–52.
Franklin J. The elements of statistical learning: data mining, inference and prediction. Publ Am Stat Assoc 2010;99:567–567.
Baschat A, Magder L, Doyle L, Atlas R, Jenkins C, Blitzer M. Prediction of preeclampsia utilizing the first trimester screening examination. Am J Obstet Gynecol. 2014;211:514.e511–517.
Mo X, Chen X, Li H, Li J, Zeng F, Chen Y, et al. Early and accurate prediction of clinical response to methotrexate treatment in juvenile idiopathic arthritis using machine learning. Front Pharmacol. 2019;10:1155.
Deis S, Rouzier R, Kayem G. Development of a nomogram to predict occurrence of preeclampsia. Eur J Obstet Gynecol Reprod Biol. 2008;137:146–51.
Huang YQ, Liang CH, He L, Tian J, Liang CS, Chen X, et al. Development and validation of a radiomics nomogram for preoperative prediction of lymph node metastasis in colorectal cancer. J Clin Oncol. 2016;34:2157–64.
Wu S, Zheng J, Li Y, Yu H, Shi S, Xie W, et al. A radiomics nomogram for the preoperative prediction of lymph node metastasis in bladder cancer. Clin Cancer Res. 2017;23:6904–11.
Ananth CV, Keyes KM, Wapner RJ. Pre-eclampsia rates in the United States, 1980-2010: age-period-cohort analysis. BMJ. 2013;347:f6564.
Sibai BM. Diagnosis and management of gestational hypertension and preeclampsia. Obstet Gynecol. 2003;102:181–92.
Luo ZC, An N, Xu HR, Larante A, Audibert F, Fraser WD. The effects and mechanisms of primiparity on the risk of pre-eclampsia: a systematic review. Paediatr Perinat Epidemiol. 2007;21:36–45.
Gray KJ, Saxena R, Karumanchi SA. Genetic predisposition to preeclampsia is conferred by fetal DNA variants near FLT1, a gene involved in the regulation of angiogenesis. Am J Obstet Gynecol. 2018;218:211–8.
Nilsson E, Salonen Ros H, Cnattingius S, Lichtenstein P. The importance of genetic and environmental effects for pre-eclampsia and gestational hypertension: a family study. BJOG. 2004;111:200–6.
Cnattingius S, Reilly M, Pawitan Y, Lichtenstein P. Maternal and fetal genetic factors account for most of familial aggregation of preeclampsia: a population-based Swedish cohort study. Am J Med Genet A. 2004;130a:365–71.
Esplin MS, Fausett MB, Fraser A, Kerber R, Mineau G, Carrillo J, et al. Paternal and maternal components of the predisposition to preeclampsia. N Engl J Med. 2001;344:867–72.
Roten LT, Johnson MP, Forsmo S, Fitzpatrick E, Dyer TD, Brennecke SP, et al. Association between the candidate susceptibility gene ACVR2A on chromosome 2q22 and pre-eclampsia in a large Norwegian population-based study (the HUNT study). Eur J Hum Genet. 2009;17:250–7.
Johnson MP, Roten LT, Dyer TD, East CE, Forsmo S, Blangero J, et al. The ERAP2 gene is associated with preeclampsia in Australian and Norwegian populations. Hum Genet. 2009;126:655–66.
Zadora J, Singh M, Herse F, Przybyl L, Haase N, Golic M, et al. Disturbed placental imprinting in preeclampsia leads to altered expression of DLX5, a human-specific early trophoblast marker. Circulation. 2017;136:1824–39.
Crovetto F, Figueras F, Triunfo S, Crispi F, Rodriguez-Sureda V, Dominguez C, et al. First trimester screening for early and late preeclampsia based on maternal characteristics, biophysical parameters, and angiogenic factors. Prenat Diagn. 2015;35:183–91.
Sibai B, Dekker G, Kupferminc M. Pre-eclampsia. Lancet. 2005;365:785–99.
Kleinrouweler CE, Mol BW. Clinical prediction models for pre-eclampsia: time to take the next step. Ultrasound Obstet Gynecol. 2014;44:249–51.
Acknowledgements
We thank the participants involved in this study for their critical contributions. The corresponding author attests that all listed authors meet authorship criteria and that no others meeting the criteria have been omitted.
Funding
1. Grant for Key Disciplinary Project of Clinical Medicine under the High-level University Development Program, Guangdong, China (2020); 2. Innovation Team Project of Guangdong Universities, China (Natural, No.2019KCXTD003); 3. Supported by 2020 Li Ka Shing Foundation Cross-Disciplinary Research Grant (2020LKSFG19B); 4. Funding for Guangdong Medical Leading Talent, the First Affiliated Hospital, SUMC, China (2019-2022); 5. Supported by the National Natural Science Foundation of China (No. 82073659); 6. Supported by 2021 Special Fund Project for Science and Technology Innovation Strategy of Guangdong Province (2021-88-53); 7. Supported by 2022 Special Fund Project for Science and Technology Innovation Strategy of Guangdong Province (2022-124-6).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, Y., Huang, X., Wu, S. et al. Machine-learning predictive model of pregnancy-induced hypertension in the first trimester. Hypertens Res 46, 2135–2144 (2023). https://doi.org/10.1038/s41440-023-01298-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41440-023-01298-8