Summary
Plant height is one of the most important agronomic traits that directly determines plant architecture, and compact or dwarf plants can allow for increased planting density and land utilization as well as increased lodging resistance and economic yield. At least four dwarf/semidwarf genes have been identified in different melon varieties, but none of them have been cloned, and little is known about the molecular mechanisms underlying internode elongation in melon. Here, we report map-based cloning and functional characterization of the first semidwarf gene short internode (Cmsi) in melon, which encodes an ERECTA-like receptor kinase regulating internode elongation. Spatial-temporal expression analyses revealed that CmSI exhibited high expression in the vascular bundle of the main stem during internode elongation. The expression level of CmSI was positively correlated with stem length in the different melon varieties examined. Ectopic expression of CmSI in Arabidopsis and cucumber suggested CmSI as a positive regulator of internode elongation in both species. Phytohormone quantitation and transcriptome analysis showed that the auxin content and the expression levels of a number of genes involved in the auxin signaling pathway were altered in the semidwarf mutant, including several well-known auxin transporters, such as members of the ABCB family and PIN-FORMED genes. A melon polar auxin transport protein CmPIN2 was identified by protein–protein interaction assay as physically interacting with CmSI to modulate auxin signaling. Thus, CmSI functions in an auxin-dependent regulatory pathway to control internode elongation in melon. Our findings revealed that the ERECTA family gene CmSI regulates stem elongation in melon through auxin signaling, which can directly affect polar auxin transport.
Similar content being viewed by others
Introduction
The deployment of dwarf or semidwarf genes was one of the main driving forces of the Green Revolution starting in the 1960s1,2,3. The reduction in plant height can increase lodging resistance, the harvest index and yield and more efficiently utilize resources1,3. Several phytohormones, such as auxins, gibberellic acids (GAs), brassinosteroids (BRs), and strigolactones (SLs), play important roles in regulating plant height4,5,6,7,8. GAs are well known to play significant roles in regulating stem elongation, leaf differentiation and seed germination. Most GA-insensitive and GA-deficient mutants are characterized by short internodes with rough and dark green leaves2,4,9,10,11. BRs and SLs are also extensively involved in regulating plant height12,13. A number of dwarfism genes involved in BR and SL biosynthesis and signal pathways have been characterized in various plant species8,14,15.
Auxin is a shoot-to-root phytohormone, and its roles in regulating several important agricultural traits, such as plant height and shoot branching, have also been well established5,16. The TRYPTOPHAN AMINOTRANSFERASE OF ARABIDOPSIS (TAA) and YUCCA (YUC) flavin monooxygenase-like enzymes control auxin biosynthesis in two independent pathways17,18. Polar auxin transport, mediated by the efflux-facilitating PIN-FORMED (PIN) family members, the influx carrier AUX1 protein family, and a number of the p-glyco-protein ABC transporters (also called ABCB family), plays an essential role in plant height regulation by building up the auxin maxima and gradient19,20,21,22. PIDs, encoding serine/threonine protein kinases belonging to the AGCVIIIa kinase family, have been reported to catalyze the efflux of auxin from cells through apical-basal PIN polar localization and/or phosphorylation of PIN proteins23,24,25. Overexpression of ZmPIN1a in maize resulted in reduced internode length, plant height, and ear height by increasing IAA transport from shoots to roots26. Overexpression of PIN2 and PIN5a in rice enhanced auxin transport from shoots to roots, resulting in shorter plant height and larger tiller angle27,28.
The Arabidopsis ERECTA family receptor kinase genes ERECTA (ER), ERECTA-LIKE1 (ERL1), and ERECTA-LIKE2 (ERL2) all encode leucine-rich repeat receptor-like kinases, which have been shown to regulate stem elongation. The erecta mutant exhibits reduced stem and hypocotyl size, and the triple mutant ererl1erl2 is extremely dwarfed29. The mechanisms of ER-family genes in regulating the development of leaf serrations and stomata have been intensively studied30,31,32. The EPIDERMAL PATTERNING FACTOR (EPF)/EPIDERMAL PATTERNING FACTOR LIKE (EPFL) family signaling peptides can physically interact with ER family receptor kinases to affect leaf teeth and stomatal development30,31,32. However, how stem elongation is regulated at the molecular level by ER-family genes remains unclear. Previous studies have shown that overexpression of the Arabidopsis auxin synthesis gene YUCCA5 can rescue the dwarf phenotype of the ERECTA mutant er-103 (ref. 33). Increasing endogenous or exogenous auxin levels could also partially rescue short hypocotyl defects in the ererl1erl2 triple mutant34. However, how the genetic interaction between the ERECTA family and auxin signaling control stem and hypocotyl development is still poorly understood.
Melon (Cucumis melo L. 2n = 2x = 24), a member of the Cucurbitaceae family, is an economically important vegetable crop worldwide35,36. More than 32 million tons of melon was produced in 2017, and China was the largest melon producer and consumer, accounting for more than half of the total production (FAO; http://faostat.fao.org/). In China, melon is produced in either open fields (where it exhibits a creeping habit) or in protected environments (with trellis support). In recent years, the area of protected melon cultivation has been constantly increasing, accounting for 65.87% of the total cultivated areas in 2017. The semidwarf plant architecture (reduced internode length and few lateral branches) has great advantages for protected melon cultivation, with its increase in plant density and hence yield per unit land area and reduced labor cost due to less pruning. To date, four recessively inherited dwarf/semidwarf mutants, si-1, si-2, si-3, and mdw1, have been reported in melon35,37. However, only mdw1 was loosely mapped on chromosome 7 by comparative mapping with cucumber35. None of these genes have been cloned, and little is known about the molecular mechanisms of plant height regulation in melon.
Here, we report the identification, map-based cloning, and functional characterization of a novel short internode semidwarf mutant in melon. We show that CmSI encodes an ERECTA-like receptor kinase and is mainly expressed in the vascular bundle of melon stems during internode elongation, and its overexpression in cucumber and Arabidopsis can promote stem elongation. We demonstrated that CmSI physically interacted with the polar auxin transport gene CmPIN2, which established the link between ERECTA family genes and auxin signaling in regulating stem elongation.
Results
Morphological characterization of short internode mutants in melon
Compared with the wild-type (WT) inbred line TopMark, the stem length was significantly decreased in the semidwarf mutant M406. The plant height of M406 could be easily distinguished after the fifth true leaf stage (Fig. 1a–c). At full maturity, the stem length of the mutant plants was approximately half that of the WT plants (Fig. 1d). All F1 plants from the cross between TopMark and M406 showed normal plant height (Fig. 1d), suggesting the recessive nature of the mutation. There was no significant difference in the total number of internodes between the two parental lines. Therefore, the semidwarf phenotype of M406 was due to the reduced internode length (Fig. 1e, f). Furthermore, the length of lateral branches at each node was significantly longer in the WT than in the mutant, but a difference in the diameter of the main stem and lateral branches was not observed between the two parental lines (Fig. 1g, h).
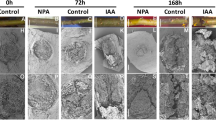
a–c Morphological characterization of two melon lines at different stages (1, 3, and 6 weeks after transplanting). d, e Plant height and stem internode numbers of two melon lines at different stages (1–10 weeks) after transplanting. The internode length (f), lateral branch length (g), and stem diameter (h) for the first ten internodes of two parental lines at 45 days after transplanting. i, j Cytological characterization of the melon normal line TopMark (i) and semidwarf line M406 (j). k Quantification of the cell size of the eighth internodes in three different TopMark and M406 plants. The cell size in the M406 plants was significantly smaller than that in TopMark plants. The data are presented as the mean ± SD. Scale bars: 10 cm (a, b), 100 μm (i, j)
Considering that the short internode may be caused by a decrease in cell number or a reduction in cell size, we examined the microscopic structure of the internodes for TopMark and M406. The cell structures of thin-sectioned eighth internodes of 40-day-old plants were examined under a light microscope (Fig. 1i, j). We found that the cells, including the parenchyma cells in particular, in the internodes of M406 were significantly smaller than those of TopMark (Fig. 1k), suggesting that the shorter internodes of the main stems of M406 were due to reduced cell size.
Fine mapping of the Cmsi locus
Among 1261 TopMark × M406 F2 plants, 330 exhibited a semidwarf phenotype, and 931 had a WT phenotype, which was consistent with the 3 (WT):1 (semidwarf) segregation ratio (Table S1), indicating that the semidwarf mutation was controlled by a single recessive gene in melon, which was named Cmsi. Initial linkage analysis with SSR markers in 92 F2 plants indicated that the Cmsi locus was located on the short arm of melon chromosome (Chr) 7 (LGVII). The two flanking markers were CmSSR17145 and CmSSR17293, which were 21.70 and 5.21 cM to the Cmsi locus, respectively. According to their physical positions on the DHL92 draft genome assembly, CmSSR17145 and CmSSR17293 were both located in the same scaffold, CM3.5_scaffold00029, with a physical distance of 1.08 Mb. Among 30 additional SSR markers from this region, four were polymorphic between M406 and TopMark. The six markers were employed to genotype 239 F2 plants. The resulting genetic map is shown in Fig. 2a. The Cmsi locus was flanked by CsSSR17251 and CsSSR17238, which were physically 127.20 kb.
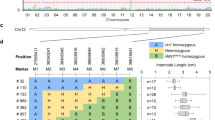
a Fine mapping of the Cmsi gene. Cmsi was delimited in a 110 kb region between markers dCAPS4 and dCAPS1. Marker dCAPS2 cosegregates with the semidwarf phenotype in F2 plants. b Genomic structure of the candidate gene MELO3C016916, which contains 27 exons and is predicted to encode an LRR receptor-like serine/threonine protein kinase ERECTA. c The single base substitution in the 1995th CDS of CmSI results in premature termination of the protein in the mutant
To identify new markers in the candidate region, TopMark and M406 were resequenced with Illumina HiSeq2500. After trimming low quality and short reads, 32,479,967 (9.73 Gb) and 32,479,967 (10.53 Gb) clean reads were obtained. Read alignment against the DHL92 sequence in the target region identified 14 polymorphic indels (>4 bp difference) between the two parental lines, 7 of which were successfully mapped with 239 F2 plants (Fig. 2a). To further confirm the mapping result, three indels and two SSR markers closely linked to Cmsi were selected for genotyping in an extended large F2 mapping population with 1,261 plants, and Cmsi was mapped between Indel 3 and Indel 14, which were 1.7 cM away from each other. The two Indel markers were used to genotype 1548 F2 plants, and 44 recombinants were identified. Four new SNP-derived dCAPS markers were developed in the candidate region between Indel 3 and Indel 14 and used to genotype these recombinants. The Cmsi locus was finally mapped in a 110 kb region defined by dCAPS1 and dCAPS4 (from 30,394,615 to 30,501,700 on CM3.5_scaffold00029) (Fig. 2a).
In the DHL92 reference genome, 14 genes were predicted in the 110 kb region (Table S2). From the resequencing data, only two SNPs were identified in the 110 kb region, with one in an intergenic region and the other in the exon of MELO3C016916 (Table S2). A dCAPS2 marker was developed based on the SNP in MELO3C016916, which showed cosegregation with the semidwarf phenotype among all 1548 F2 plants used in this study, suggesting that MELO3C016916 is a candidate gene for Cmsi (Fig. 2a).
CmSI is a homolog of ERECTA family receptor kinases
The genomic DNA sequence of MELO3C016916 in the DHL92 melon reference genome is 7128 bp, which was predicted to have 27 exons (Fig. 2b). The full length of the coding sequence (CDS) of MELO3C016916 was 2976 bp, encoding a protein with 991 amino acid residues. The single nucleotide substitution from T to G in the 25th exon introduced a premature stop codon, which led to a truncated protein in M406 (Fig. 2c). Gene prediction and functional annotation revealed that MELO3C016916 encoded an LRR receptor-like serine/threonine protein kinase ERECTA (Fig. S1a). Sequence alignment of CmSI and other members of the ERECTA protein family from Arabidopsis (AtERECTA, AtELK1, and AtELK2) and cucumber (CsERECTA) showed that CmSI shared the highly conserved leucine-rich repeat N-terminal domain (LRRNT), the LRR repeat region, and the S_TKc domain with CsERECTA (Fig. S1a). The amino acid sequence identity of CmSI to CsERECTA, AtERECTA, AtELK1, and AtELK2 was 98.99, 78.73, 62.20, and 61.09%, respectively (Fig. S1a).
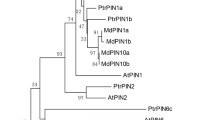
To investigate the evolutionary relationship between CmSI and other ERECTA family proteins, a neighbor-joining phylogenetic tree was developed using protein sequences from 34 species (Fig. S1b). The phylogenetic tree of the ERECTA family can be divided into two main groups: monocotyledons and dicotyledons. CmSI was clustered within the dicotyledon group, which includes known ERECTA-like protein kinases, such as AtERECTA (Arabidopsis thaliana), PtERECTA (Populus trichocarpa), and VvERECTA (Vitis vinifera) (Fig. S1b). These results indicated that CmSI may have a similar function to other ERECTA proteins in melon.
CmSI is highly expressed in the stem vascular bundle and ovary
We examined the temporal-spatial expression of CmSI in roots, stems, leaves, male flowers, and ovaries of TopMark and M406 using qRT-PCR. The expression level of CmSI was lower in all the tested organs in the mutant M406 than in TopMark (Fig. 3a). The highest expression of CmSI was detected in stems and ovaries of both lines (Fig. 3a). Furthermore, we analyzed the transcript level of CmSI in different internodes of the main stems of WT plants. The transcripts of CmSI were abundantly accumulated in the young internodes (upper part) but were rapidly reduced as the internode elongation stopped (Fig. 3b).
a qRT-PCR analysis of CmSI expression in different organs of melon. b CmSI expression in stem internodes at different developmental stages. The melon gene ACTIN was used as the internal control. Error bars indicate standard deviations of three biological replicates. R root, S stem, L leaf, MF male flower, O ovary on the day of flowering. c–f mRNA in situ hybridization of CmSI in stems at 1 week after transplanting. CmSI is highly expressed in the vascular bundle of TopMark stems (c, d) and is decreased in stems of the M406 mutant (e, f). g Negative controls hybridized in stem with the sense probe. Bars = 100 µm in c–g
We further examined the expression pattern of CmSI in young stems of both TopMark and M406 with RNA in situ hybridization. CmSI transcripts were also detected in the epidermis and the vascular bundle of TopMark stems (Fig. 3c, d) but not in the vascular bundles of M406 stems (Fig. 3e, f), which was consistent with the qRT-PCR results. These observations suggested that CmSI may be involved in regulating internode elongation in melon.
Nucleotide variation in CmSI among melon varieties
To examine the allelic diversity of the CmSI gene in natural melon populations, we examined the nucleotide variation of the CmSI locus among 200 resequenced accessions, including 36 wild melons. The 164 cultivated accessions were further classified into two subspecies: 66 subsp. melo accessions and 98 subsp. Agrestis accessions. Eight SNPs were identified within the coding sequence of CmSI, of which seven were synonymous and only one was nonsynonymous (Fig. 4a). Interestingly, the only polymorphism resulting in amino acid substitutions occurred only in the wild melon group. However, there was no obvious difference in plant height between the wild melon group and cultivated melon group, indicating that this mutation may not be in the functional domains and did not affect internode elongation. Moreover, the Cmsi mutant allele was only found in the dwarf line M406, indicating that the Cmsi allele was a spontaneous mutation that was not under selection during domestication or diversifying selection (Fig. 4a).
a Nucleotide variation at the si locus among 200 resequenced lines. The si allele was found in only M406, which suggested that the si allele was a spontaneous mutation and was not under selection during domestication or diversifying selection. b, c The expression pattern of the CmSI gene in 14 different melon germplasms and F1 and F2 of TopMark and M406. High CmSI expression levels were closely correlated with increased plant height among 14 different melon lines
The relationship between plant height and the expression level of CmSI was also analyzed in another 12 melon varieties, the F1 and F2 plants of TopMark and M406. The expression levels of CmSI were the lowest in M406 and the F2 semidwarf plants, indicating that the Cmsi special mutation caused a decreased expression level of the CmSI gene. The relationship between the CmSI expression level and plant height in different melon varieties was further demonstrated by the observed correlation of CmSI expression level and plant height (R2 = 0.6728; Fig. 4b, c). This further confirmed that CmSI functions in an expression-dependent manner in plant height regulation.
Ectopic overexpression of CmSI increases plant height in Arabidopsis and cucumber
To investigate the functional conservation of CmSI in other plants, 35S:CmSI overexpression vectors were constructed and then introduced into the Arabidopsis ERECTA mutant er105. The 35S promoter was used instead of the AtERECTA promoter because AtERECTA showed a different expression pattern from CmSI29,38. Seven independent transgenic lines were obtained, and three representative overexpression (OE) lines (35S:CmSI::er105#1, 35S:CmSI::er105#2 and 35S:CmSI::er105#3) were selected for detailed analysis. We observed that all three OE lines partially rescued the dwarf phenotype of the er105 mutant (Fig. 5a); their plant heights were 68, 160, and 134% higher in lines 1, 2, and 3 than in the er105 mutant (Fig. 5b). To further validate the function of CmSI, we also introduced the overexpressed CmSI construct into WT Arabidopsis (Col). In all six independent transgenic lines, the plant height was higher than that in the WT (Fig. 5c, d). These data confirmed the function of CmSI in promoting internode elongation and increasing plant height. These observations suggest that homologs of CmSI may perform similar functions in Arabidopsis and melon.
a, b Phenotypic comparison and statistical analysis of plant height of er105, 35S:CmSI::er105 lines of Arabidopsis. Overexpression of CmSI in er105 rescues the dwarf phenotype. c, d Phenotypic comparison and statistical analysis of plant height of col, 35S:CmSI::col lines of Arabidopsis. Overexpression of CmSI in wild-type col can also increase plant height. e, f Phenotypes and statistical analysis of 35S:CmSI transgenic cucumber plants. Overexpression of CmSI increases plant height in cucumber
Considering the difficulty of transformation technology in melon, we transformed the CmSI overexpression construct into the WT cucumber line 3546. Eight overexpression transgenic lines were generated, and the expression level of CmSI was much higher in three representative overexpression lines (35S:CmSI::3546#1, 35S:CmSI::3546#2, and 35S:CmSI::3546#3) than in the WT line (Fig. S2a). We observed that overexpression of CmSI increased the plant height in cucumber due to increased internode length (Fig. 5e, f and Fig. S2b). However, there was no significant difference in the total number of internodes and the diameter of the main stem in the control plants and that in 35S:CmSI transgenic plants (Fig. S2c, d).
Transcriptome profiling reveals important roles of auxin in internode elongation
To reveal the regulatory network of internode development, two RNA-Seq datasets were analyzed using the eighth internode of the main stem. One dataset was from the comparison of the transcriptomes of the eighth internodes of TopMark and M406 plants. The second dataset is from the comparison between transcriptomes of the D-bulk (dwarf) and N-bulk (WT) plants, which were developed by pooling dwarf and WT bulks in F2 plants (see “Experimental procedure” for details). Ten RNA-Seq libraries were subjected to high-throughput sequencing, which generated an average of 6.84 and 17.07 Gb clean data for the two parental lines and two bulks, respectively (Fig. S3a and Table S3). Using a false discovery rate (FDR) of 0.05 as the cutoff, 643 down- and 633 upregulated differentially expressed genes (DEGs) were identified in the semidwarf mutant M406 compared with the WT, respectively (Fig. S3b and Table S4). Compared with the N-bulk (WT), among 1161 DEGs, 558 and 603 were up- and downregulated, respectively, in the D-bulk (Fig. S3b and Table S5). From the two datasets, 228 common DEGs with the same expression patterns were identified, including 94 up- and 134 downregulated DEGs (Fig. S3b and Table S6).
To analyze the functions of these DEGs, functional categorization of the DEGs was carried out by MapMan and Gene Ontology (GO) term enrichment analyses. MapMan analysis revealed that a large number of these DEGs were related to “hormone metabolism”, “transport”, “development”, and “signaling”, which were the most significantly enriched between two parental lines, two bulks and common DEGs (Fig. 6a and Tables S4–6). The GO enrichment analysis of the DEGs also identified various hormone metabolism-related terms, particularly those related to auxin biosynthetic processes, which were significantly enriched in the DEGs of the two parental lines and two bulks (Fig. 6b and Tables S4–6). A number of DEGs identified in this study have been shown to be involved in the auxin signaling pathway, including several auxin biosynthesis and polar transport genes, such as PINs and ABCBs (Fig. 6c, d). Quantitative real-time PCR analysis verified that the relative expression levels of auxin biosynthesis and polar transport genes were significantly higher in the semidwarf mutant M406 than in TopMark (Fig. 6e). Moreover, we measured the auxin levels in the stems of the two parental lines. As expected, the auxin concentration was significantly reduced in the semidwarf mutant M406 (Fig. 6f), which is consistent with the promoting role of auxin in internode elongation. These results suggested that CmSI may affect auxin transport or signaling in stems.
a Functional categories for common DEGs. b GO enrichment analysis of DEGs in two bulks. c, d Heat maps of auxin-related genes that were differentially expressed in the M406 mutant (c) and dwarf bulks (d). e Quantitative RT-PCR of auxin-related genes in stems of TopMark and M406 mutants. f IAA concentration in the stems of TopMark and M406 mutant plants
CmSI interacts with CsPIN2 at the protein level
To identify the putative interacting proteins of CmSI, we conducted yeast two hybridization (Y2H) assays to detect interactions of CmSI with the differentially expressed auxin-related genes and transcriptional regulators required for plant organ development, including the efflux-facilitating PIN-FORMED2 (MELO3C004520.2), three auxin response factors (MELO3C013003.2, MELO3C005476.2, MELO3C004026.2), two boundary genes (MELO3C021578.2, MELO3C026269.2), two Gretchen Hagen 3 genes (MELO3C018166.2, MELO3C016616.2), a member of pleiotropic drug resistance ABC transporter (MELO3C009686.2), five different transcription factors (MELO3C004181.2, MELO3C010984.2, MELO3C021426.2, MELO3C026299.2, MELO3C030287.2). Among them, only CmPIN2 was found to interact with CmSI (Fig. 7a). Moreover, the interactions between CmSI and CmPIN2 were further confirmed by BiFC analysis, which showed that CmPIN2 could direct interact with CmSI rather than the mutated protein Cmsi (Fig. 7b). These results suggested that CmSI may regulate stem elongation via interacting with CmPIN2 in melon.
a The protein interaction was examined using various combinations of prey and bait vectors. The vector pPR3-N and pTSU2-App was used as a negative control, and the interaction between pTSU2-APP and pNubG-Fe65 was used as a positive control. Yeast cells were grown on SD/-Leu-Trp medium and interactions were confirmed by an SD/-Leu-Trp-His-Ade-X-a-GAL assay on medium. Dilutions (1 and 10−1) of saturated cultures were spotted onto the plates. Yeast two-hybrid assay showing that CmSI rather than Cmsi interacted with CmPIN2 by growth on SD/-Leu/-Trp/-His/-Ade/X-α-gal plates. b BiFC analysis of the physical interaction between CmSI and Cmsi (fused with the C-terminal fragment of YFP) and CmPIN2 (fused with the N-terminal fragment of YFP). INDEHISCENT (IND)-YFPC and SPATULA (SPT)-YFPN were used as positive controls. CmPIN2-YFPN and the empty YFPC were used as negative controls. Different combinations of the fused constructs were co-expressed in leaves of Nicotiana tabacum, and the cells were then visualized using confocal microscopy
Discussion
Identification and map-based cloning of a novel semidwarf mutant in melon
Plant height is an important agronomic trait affecting crop architecture, resistance to lodging, tolerance to crowding, fruit yield, and mechanical harvesting. Almost all commercial melon varieties have long main stems and many lateral branches, which requires pruning in standard cultivation practices. Pruning of these plants is very labor intensive and time consuming. Therefore, the development of semidwarf cultivars with short branches is an important objective in melon breeding, especially for production under protected environments. To date, four recessive dwarf/semidwarf mutants, si-1, si-2, si-3, and mdw1, have been identified35,37, but only mdw1 was genetically mapped on Chr7 (ref. 35). In this study, Cmsi was also mapped on Chr7 in a 110.00 kb region, and CmERECTA was identified as the candidate gene (Fig. 2). Hwang et al. (2014) found that the mdw1 locus was closely linked with the mERE gene, which is the same as CmERECTA (MELO3C016916). Therefore, it is reasonable to speculate that Cmsi and mdw1 probably belong to the same locus, but whether the two mutants have the same mutant allele is not known. The only SNP in the CmERECTA gene between the two parental lines located in the exon resulted in a premature termination codon in the semidwarf line M406. The SNP-derived dCAPS2 marker cosegregated in the F2 population. Furthermore, we examined the diversity of nucleotides and amino acids at the CmSI locus in a panel of 200 melon accessions of the global collection. The SNP allele in CmERECTA was only present in the mutant M406 among accessions from wild melon and cultivated melons. It should be noted that there were also seven other SNP variants among the 200 melon accessions. All seven SNPs were synonymous mutations in 164 cultivated melon accessions. Although there was only one nonsynonymous SNP in some wild melon accessions, the mutant allele was not associated with stem length variation (Fig. 4), suggesting that this SNP site is probably not located in the functional domain. Therefore, the cosegregated marker dCAPS2 of CmSI will be very useful in marker-assisted selection for use of this Cmsi allele in melon breeding.
CmSI encodes an ERECTA-like protein of the leucine-rich repeat (LRR) subfamily
The ERECTA gene family, ERECTA, ERL1, and ERL2, which encode leucine-rich repeat receptor-like kinases, has been found in many monocots and dicots. These genes play important roles in different developmental processes, including inflorescence architecture, hypocotyl development, seed germination, ovule development, and stomatal and trichome formation34,39,40,41. We show that CmSI encodes an ERECTA protein (Fig. 2) that shares a high level of sequence identity with the AtERECTA protein (Fig. S1). Overexpression of CmSI in Arabidopsis and cucumber increased their plant height (Fig. 5). This convincing evidence suggests that CmSI, like other ERECTA-like homologs, regulates cell development and stem internode elongation in melon. The expression of CmSI was much higher in the young internodes during internode elongation and declined rapidly when internode elongation stopped (Fig. 3b). Moreover, the CmSI transcript was more abundant in stems and ovaries than in other tissues in both WT and semidwarf plants (Fig. 3). In previous studies, the expression levels of grape VvERECTA and tomato SlERECTA were higher in young fruits than in mature fruits40,42, indicating that ERECTA-like genes play important roles in early organ development in different species. However, the temporal and spatial expression pattern of CmSI in melon was not exactly the same as that of AtERECTA in Arabidopsis. The expression of AtERECTA was higher in developing above-ground organs such as flowers and young rosettes but lower in stems and almost undetectable in roots29,38,43. In our study, the expression level of CmSI was high in roots and stems, indicating that CmSI and AtERECTA may play distinct roles in different developmental processes between the two species. Moreover, the semidwarf mutant M406 has shorter primary roots than the normal line TopMark, indicating that CmSI also affects root development in melon (Fig. S4).
In Arabidopsis, AtERECTA has partial functional redundancy with its two homologous genes ERL1 and ERL2 in hypocotyl and stem elongation, leaf serration development, and stoma and trichome formation. The triple mutant ererl1erl2 intensifies the erecta mutant phenotype, and the ererl1erl2 plant is extremely dwarfed compared with erecta29,44. Compared with the WT TopMark, the length of the main stem and lateral branches was significantly decreased in the mutant M406 (Fig. 1). However, the trichome numbers on the stems of the semidwarf mutant did not show a significant change (Fig. S5c, d). In this study, we found that the transcript of the CmERL1 gene was significantly increased in the mutant M406 (Fig. 6e and Fig. S5b), indicating that CmERL1 may play a redundant role with CmSI in melon trichome development.
CmSI regulates stem elongation through the auxin pathway
Plant height is regulated by multiple genes and associated complex regulatory networks. As a classical phytohormone, auxin plays important roles in regulating key agricultural traits associated with plant height and shoot branching16,45. Previous studies have shown that overexpression of the Arabidopsis auxin biosynthesis gene YUCCA5 can rescue the dwarf phenotype of the ERECTA mutant er-103 (ref. 33). Increasing endogenous or exogenous auxin levels could also partially rescue short hypocotyl defects of the ererl1erl2 triple mutant34. Here, transcriptomic analysis showed that “auxin biosynthetic process” and “auxin metabolic process” were the most significantly enriched GO terms for these DEGs in stem elongation (Table S4), and the functions of a number of DEGs were related to “hormone metabolism”, “transport”, and “signaling”, suggesting that auxin plays an important role in regulating internode elongation in melon. Moreover, the auxin content was decreased in the stem of M406, and the cell size in the stem of semidwarf plants was also smaller than that of TopMark. This evidence indicated that CmSI may regulate stem elongation by affecting the auxin content in melon stems. From the Y2H assay, we found that CmSI directly interacted with the efflux-facilitating PIN-FORMED protein CmPIN2. The homologs of CmPIN2 in rice, maize, and many other species have been shown to positively regulate auxin transport during organ formation23,24,25. Overexpression of OsPIN2 in rice can enhance auxin transport from stems to roots and thus decrease plant height27. ZmPIN1a overexpression also increased IAA transport from shoots to roots and reduced plant height, ear height, and internode length in maize26. More importantly, CmSI rather than Cmsi could interact with CmPIN2, indicating that CmSI may regulate stem development by interacting with CmPIN2, and the single nucleotide mutation of Cmsi in the semidwarf mutant M406 disrupted the interaction between CmSI and CmPIN2. Taken together, our data elucidated a novel link between the ERECTA protein and auxin polar transport in melon.
Experimental procedures
Plant material and growth conditions
The short internode mutant line M406 displays shorter internodes and fewer lateral branches than the normal line. To confirm the inheritance mode and fine mapping of the Cmsi gene, M406 was crossed with the WT muskmelon inbred line TopMark to generate a large F2 mapping population. The χ2-test for goodness-of-fit was used to test for deviation of the observed data from the theoretically expected segregation for semidwarf phenotype data in F2 plants. All plant materials were grown in greenhouses at the Maozhuang Research Station of Henan Agricultural University (Zhengzhou, China). The plant height (stem length) of each plant was qualitatively recorded as either normal or mutant at 30 and 60 days after transplanting. In addition, another 12 melon inbred lines from different regions and a monoecious cucumber (Cucumis sativus L.) inbred line 3546 (WT) used in this study were also cultivated in a greenhouse under normal conditions. The Arabidopsis mutant er105 (Col background) and Col WT were used for functional complementary verification. Arabidopsis seeds were germinated on Murashige–Skoog (MS) medium containing 0.2% Phytagar and 1% sucrose. The seeds were kept at 4 °C for 3 days and then moved to 22 °C under a 16 h light/8 h dark light regime. Seedlings were transferred to soil 7–10 days after germination.
Unexpanded young leaves from test plants were collected into 1.5 mL microcentrifuge tubes, lyophilized in a freeze dryer, and ground into fine powder. Genomic DNA was extracted using the CTAB method34.
Microscopic examination of internodes
To compare the microscopic structure of internodes in WT and mutant plants, the eighth internodes of the stems of 40-day-old plants of TopMark and M406 were fixed, rinsed, postfixed, washed, dehydrated, and embedded. Semithin sections were prepared with Formvar-coated gold grids and observed with an Olympus-BX53 light microscope as previously described46.
Molecular marker analysis and fine mapping
For initial mapping of Cmsi, a linkage map was developed using 92 TopMark × M406 F2 plants. Linkage analysis placed the Cmsi locus on the short arm of melon chromosome 7 (LGVII) flanked with CmSSR17145 and CmSSR17293, which were used as the starting point for fine mapping. In the target region, we first explored SSR markers47. Additional SNPs and Indels were identified through bioinformatics analysis of resequencing reads of TopMark and M406. The resequencing of TopMark and M406 was carried out following the standard Illumina protocol, and the library was used for paired-end sequencing on the Illumina HiSeq2500 analyzer. After removing short reads and low-quality reads, the clean reads from the two parental lines were used for mapping to the melon reference genome DHL92 (https://melonomics.net/)48 using BWA software49. SNPs and small Indels detected from the alignments were called using Samtools, and output was given in pileup format50. The SNPs and small Indels between two parental lines were detected using the GATK software tool package51, and reliable SNPs and small Indels were noted and predicted using SnpEff software52. Only those Indels with ≥3 bp differences were selected for primer design with Primer3 software (http://primer3.ut.ee/), and dCAPS markers were developed for SNP genotyping by dCAPS Finder 2.0 (ref. 53).
Polymorphic SSR and Indel markers were used for chromosome walking and fine mapping in a large population containing 1261 F2 plants. Finally, additional dCAPS makers were used for genotyping those F2 plants to identify recombinants identified from 1548 F2 plants. The PCR amplification of molecular markers and subsequent gel electrophoresis were performed as previously described47. Linkage analysis of the Cmsi locus with molecular markers was performed with the Kosambi mapping function using JoinMap 3.0.
Sequence alignments and phylogenetic analysis
The coding sequence (CDS) of CmSI was amplified by PCR from stem cDNA using gene-specific primers (Supplementary Table S7). The amino acid sequences of the related ERECTA-like proteins from Arabidopsis, cucumber, and other species were obtained by BLAST searches of the National Center for Biotechnology Information nucleotide database (http://www.ncbi.nlm.nih.gov/nucleotide/). A multiple sequence alignment of CmSI and the related ERECTA-like proteins was carried out as previously described54. A phylogenetic tree was developed using the neighbor-joining (NJ) method55 in the MEGA5 software package.
To examine the allelic diversity of the CmSI gene in natural melon populations, the clean reads of 200 resequenced melon accessions were aligned to the reference sequence of the CmSI gene (DHL92 draft genome) for SNP calling as previously described56. Furthermore, the plant height of five plants from each of the 200 melon accessions was recorded as normal height or dwarf height at 30 and 60 days in the field in 2017 and 2018.
Spatial and temporal expression analysis by quantitative real-time PCR
Total RNA of the stems, roots, leaves, male flowers, ovaries, and internodes was extracted using the Quick RNA isolation kit (Huayueyang, China) and reverse transcribed to first strand cDNA using the PrimeScript First Strand cDNA Synthesis Kit (Takara). SYBR® Premix Ex Taq from TaKaRa was used for qPCR with the Applied Biosystems StepOne™ Real-Time PCR System. The melon ACTIN gene was used as the internal control57 in all qPCR reactions. All experiments were performed with three biological and three technical replicates. The CmSI-specific qPCR primers are listed in Supplementary Table S7.
RNA in situ hybridization
Tender stems of WT and mutant plants were fixed, dehydrated, dewaxed, embedded, sectioned, and hybridized with digoxigenin-labeled probes as previously described58. Digoxigenin-labeled sense and antisense RNA probes were obtained using T7 and SP6 RNA polymerases (Roche). The primer pairs used are listed in Supplementary Table S7.
Ectopic expression of CmSI in Arabidopsis and cucumber
The full-length coding regions of CmSI were cloned without the stop codon and inserted into the SuperpCAMBIA1300 vector59 between the SpeI and SmaI sites. The CmSI-SuperpCAMBIA1300-overexpressing vector was transformed into er105 mutant and Col (WT) plants using the floral dip method60. The transgenic Arabidopsis plants were screened on Murashige and Skoog (MS) medium with 25 mg l–1 hygromycin.
The CmSI-SuperpCAMBIA1300 fusion vector was transformed into cucumber line 3546 (WT) using a cotyledon transformation method as previously described61. In brief, the Agrobacterium strain AGL1 was transformed with the CmSI-SuperpCAMBIA1300 fusion vector and used to transform embryonic callus of cucumber by cocultivation. MS medium supplemented with 10 mg/l hygromycin was used to select transformants. The positive transgenic plants were verified by PCR using specific primers. The primers are listed in Supplementary Table S7. At least three representative transgenic lines and three plants in each line were used for further analysis.
Transcriptome analysis
To investigate the regulatory network of the CmSI gene, we used a strategy combining the RNA-seq of two parental lines and bulked-segregant analysis (BSR-seq) of the F2 population. For RNA-Seq, two bulks, the dwarf bulk (D-bulk) and the normal bulk (N-bulk), were constructed by pooling 20 dwarf and 20 WT F2 plants, respectively.). Total RNA of the main stems of the two bulks and the eighth nodes of both parents were extracted and used for strand-specific RNA-Seq library construction and next-generation sequencing on an Illumina HiSeqTM 4000 platform. Three biological and two technical replications were sequenced for the parental lines and bulks, respectively.
After removing the adapters and low-quality reads, the clean data were used for alignment to the melon reference genome DHL92 using TopHat v2.1.1 (ref. 62). The read numbers of annotated genes were counted by the HTSeq program (v0.9.1)63. The number of transcripts per million reads (TPM) for each gene was calculated based on the length of the gene and mapped read counts. DEGs between two parental lines and two bulks were identified using the DEGSeq R package (1.12.0). A corrected P value of 0.05 was set as the threshold for DEG selection. GO terms for these DEGs were determined using InterProScan program64,65. Then, GO functional enrichment analysis was performed to identify DEGs with significantly enriched GO terms. The GO analysis was carried out with AgriGO with an FDR ≤ 0.05 to obtain the GO annotations based on the biological process, molecular function, and cellular component categories66. The functional categorization of these DEGs was classified using MapMan.
Auxin quantitation
For measurement of the auxin content, fresh stems (50 mg) of 30-day-old plants were frozen in liquid nitrogen. Quantification of endogenous auxin levels was performed using high-performance liquid chromatography electrospray ionization tandem mass spectrometry (HPLC-ESI-MS/MS)67.
Yeast two-hybrid assay
For the yeast two-hybrid assay, we cloned the CDS without an N-terminal cleavable signal sequence, and the stop codon of CmSI and Cmsi fused them into the pBT3-SUC vector. The full-length CDSs of CmPIN2 were cloned and fused with the pPR3-N vector. The combination of pTSU2-APP and pNubG-Fe65 was used as a positive control, and the combination of pTSU2-APP and pPR3-N was used as a positive control. All recombinant constructs were separately transformed into the yeast strain NMY51. According to the DUAL membrane starter kit user manual, the transformed yeast cells were grown on synthetic defined (SD) plates lacking tryptophan and histidine (SD/−Trp−His) and lacking tryptophan, histidine, and adenine (SD/−Trp−His−Ade) with α-gal.
Accession numbers
GenBank accession numbers of ERECTA-LIKE protein sequences used in this study included Arabidopsis ERECTA (AT2G26330), ERECTA-LIKE1 (AY244745), ERECTA-LIKE2 (AY244746), and cucumber ERECTA-LIKE (EST241733).
Data availability
The resequencing data and transcriptome sequencing data of two parental lines and two bulks are available from the NCBI Short Read Archive (SRA PRJNA608205).
References
Ferrero-Serrano, A., Cantos, C. & Assmann, S. M. The role of dwarfing traits in historical and modern agriculture with a focus on rice. Cold Spring Harb. Perspect. Biol. 11, https://doi.org/10.1101/cshperspect.a034645 (2019).
Ishiyama, K. et al. Green revolution: a mutant gibberellin-synthesis gene in rice. Nature 416, 701–702 (2002).
Wang, B., Smith, S. M. & Li, J. Genetic regulation of shoot architecture. Annu. Rev. Plant Biol. 69, 437–468 (2018).
Ayano, M. et al. Gibberellin biosynthesis and signal transduction is essential for internode elongation in deepwater rice. Plant Cell Environ. 37, 2313–2324 (2014).
Multani, D. S. et al. Loss of an MDR transporter in compact stalks of maize br2 and sorghum dw3 mutants. Science 302, 81–84 (2003).
Nomura, T. et al. Brassinosteroid deficiency due to truncated steroid 5alpha-reductase causes dwarfism in the lk mutant of pea. Plant Physiol. 135, 2220–2229 (2004).
Pearce, S. et al. Molecular characterization of Rht-1 dwarfing genes in hexaploid wheat. Plant Physiol. 157, 1820–1831 (2011).
Tamiru, M. et al. A cytochrome P450, OsDSS1, is involved in growth and drought stress responses in rice (Oryza sativa L.). Plant Mol. Biol. 88, 85–99 (2015).
Fleet, C. M. et al. Overexpression of AtCPS and AtKS in Arabidopsis confers increased ent-kaurene production but no increase in bioactive gibberellins. Plant Physiol. 132, 830–839 (2003).
Hedden, P. & Phillips, A. L. Gibberellin metabolism: new insights revealed by the genes. Trends Plant Sci. 5, 523–530 (2000).
Hedden, P. & Thomas, S. G. Gibberellin biosynthesis and its regulation. Biochem. J. 444, 11–25 (2012).
de Saint Germain, A. et al. Strigolactones stimulate internode elongation independently of gibberellins. Plant Physiol. 163, 1012–1025 (2013).
Wang, Z. Y., Bai, M. Y., Oh, E. & Zhu, J. Y. Brassinosteroid signaling network and regulation of photomorphogenesis. Annu. Rev. Genet. 46, 701–724 (2012).
Hou, S. et al. A mutant in the CsDET2 gene leads to a systemic brassinosteriod deficiency and super compact phenotype in cucumber (Cucumis sativus L.). Theor. Appl. Genet. Theor. Angew. Genetik 130, 1693–1703 (2017).
Wang, H. et al. The cytochrome P450 gene CsCYP85A1 is a putative candidate for Super Compact-1 (Scp-1) plant architecture mutation in cucumber (Cucumis sativus L.). Front. Plant Sci. 8, 266 (2017).
Palme, K., Dovzhenko, A. & Ditengou, F. A. Auxin transport and gravitational research: perspectives. Protoplasma 229, 175–181 (2006).
Cheng, Y., Dai, X. & Zhao, Y. Auxin biosynthesis by the YUCCA flavin monooxygenases controls the formation of floral organs and vascular tissues in Arabidopsis. Genes Dev. 20, 1790–1799 (2006).
Mashiguchi, K. et al. The main auxin biosynthesis pathway in Arabidopsis. Proc. Natl Acad. Sci. USA 108, 18512–18517 (2011).
Bennett, S. R., Alvarez, J., Bossinger, G. & Smyth, D. R. Morphogenesis in pinoid mutants of Arabidopsis thaliana. Plant J. 8, 505–520 (1995).
Blakeslee, J. J., Peer, W. A. & Murphy, A. S. Auxin transport. Curr. Opin. Plant Biol. 8, 494–500 (2005).
Friml, J. Auxin transport—shaping the plant. Curr. Opin. Plant Biol. 6, 7–12 (2003).
Geisler, M. & Murphy, A. S. The ABC of auxin transport: the role of p-glycoproteins in plant development. FEBS Lett. 580, 1094–1102 (2006).
Lee, S. H. & Cho, H. T. PINOID positively regulates auxin efflux in Arabidopsis root hair cells and tobacco cells. Plant Cell 18, 1604–1616 (2006).
Michniewicz, M. et al. Antagonistic regulation of PIN phosphorylation by PP2A and PINOID directs auxin flux. Cell 130, 1044–1056 (2007).
Zourelidou, M. et al. Auxin efflux by PIN-FORMED proteins is activated by two different protein kinases, D6 PROTEIN KINASE and PINOID. Elife 3, e02860 (2014).
Li, Z. et al. Enhancing auxin accumulation in maize root tips improves root growth and dwarfs plant height. Plant Biotechnol. J. 16, 86–99 (2018).
Chen, Y., Fan, X., Song, W., Zhang, Y. & Xu, G. Over-expression of OsPIN2 leads to increased tiller numbers, angle and shorter plant height through suppression of OsLAZY1. Plant Biotechnol. J. 10, 139–149 (2012).
Lu, G. et al. OsPIN5b modulates rice (Oryza sativa) plant architecture and yield by changing auxin homeostasis, transport and distribution. Plant J. 83, 913–925 (2015).
Shpak, E. D., Berthiaume, C. T., Hill, E. J. & Torii, K. U. Synergistic interaction of three ERECTA-family receptor-like kinases controls Arabidopsis organ growth and flower development by promoting cell proliferation. Development 131, 1491–1501 (2004).
Jewaria, P. K. et al. Differential effects of the peptides Stomagen, EPF1 and EPF2 on activation of MAP kinase MPK6 and the SPCH protein level. Plant Cell Physiol. 54, 1253–1262 (2013).
Lin, G. et al. A receptor-like protein acts as a specificity switch for the regulation of stomatal development. Genes Dev. 31, 927–938 (2017).
Meng, L., Buchanan, B. B., Feldman, L. J. & Luan, S. CLE-like (CLEL) peptides control the pattern of root growth and lateral root development in Arabidopsis. Proc. Natl Acad. Sci. USA 109, 1760–1765 (2012).
Woodward, C. et al. Interaction of auxin and ERECTA in elaborating Arabidopsis inflorescence architecture revealed by the activation tagging of a new member of the YUCCA family putative flavin monooxygenases. Plant Physiol. 139, 192–203 (2005).
Qu, X., Zhao, Z. & Tian, Z. ERECTA regulates cell elongation by activating auxin biosynthesis in Arabidopsis thaliana. Front. Plant Sci. 8, 1688 (2017).
Hwang, J. et al. Fine genetic mapping of a locus controlling short internode length in melon (Cucumis melo L.). Mol. Breed. 34, 949–961 (2014).
Kerje, T. & Grum, M. The origin of melon, Cucumis melo: a review of the literature[C]//VII Eucarpia Meeting on Cucurbit Genetics and Breeding 510. 37–44 (2000).
Paris, H. S., Nerson, H. & Karchi, Z. Genelics of internode length in melons. J. Heredity 75, 403–406 (1984).
Torii, K. U. et al. The Arabidopsis ERECTA gene encodes a putative receptor protein kinase with extracellular leucine-rich repeats. Plant Cell 8, 735–746 (1996).
Chen, M. K., Wilson, R. L., Palme, K., Ditengou, F. A. & Shpak, E. D. ERECTA family genes regulate auxin transport in the shoot apical meristem and forming leaf primordia. Plant Physiol. 162, 1978–1991 (2013).
Liu, M., Li, W., Min, Z., Cheng, X. & Fang, Y. Identification and expression analysis of ERECTA family genes in grape (Vitis vinifera L.). Genes Genomics 41, 723–735 (2019).
Nanda, A. K., El Habti, A., Hocart, C. H. & Masle, J. ERECTA receptor-kinases play a key role in the appropriate timing of seed germination under changing salinity. J. Exp. Bot. 70, 6417–6435 (2019).
Villagarcia, H., Morin, A.-C., Shpak, E. D. & Khodakovskaya, M. V. Modification of tomato growth by expression of truncated ERECTA protein from Arabidopsis thaliana. J. Exp. Bot. 63, 6493–6504 (2012).
Shpak, E. D. Diverse roles of ERECTA family genes in plant development. J. Integr. Pant Biol. 55, 1238–1250 (2013).
Pillitteri, L. J., Bemis, S. M., Shpak, E. D. & Torii, K. U. Haploinsufficiency after successive loss of signaling reveals a role for ERECTA-family genes in Arabidopsis ovule development. Development 134, 3099–3109 (2007).
Mattsson, J., Ckurshumova, W. & Berleth, T. Auxin signaling in Arabidopsis leaf vascular development. Plant Physiol. 131, 1327–1339 (2003).
Yang, S. et al. A CsTu-TS1 regulatory module promotes fruit tubercule formation in cucumber. Plant Biotechnol. J. 17, 289–301 (2019).
Zhu, H. et al. Development of genome-wide SSR markers in melon with their cross-species transferability analysis and utilization in genetic diversity study. Molecular Breeding 36, https://doi.org/10.1007/s11032-016-0579-3 (2016).
Garcia-Mas, J. et al. The genome of melon (Cucumis melo L.). Proc. Natl Acad. Sci. 109, 11872–11877 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
DeLuca, D. S. et al. RNA-SeQC: RNA-seq metrics for quality control and process optimization. Bioinformatics 28, 1530–1532 (2012).
Chen, K. et al. BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nat. Methods 6, 677 (2009).
Neff, M. M., Neff, J. D., Chory, J. & Pepper, A. E. dCAPS, a simple technique for the genetic analysis of single nucleotide polymorphisms: experimental applications in Arabidopsis thaliana genetics. Plant J. 14, 387–392 (1998).
Yang, S. et al. A CsMYB6-CsTRY module regulates fruit trichome initiation in cucumber. J. Exp. Bot. 69, 1887–1902 (2018).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Yang, L. et al. LITTLELEAF (LL) encodes a WD40 repeat domain‐containing protein associated with organ size variation in cucumber. Plant J. 95, 834–847 (2018).
Zhu, H. et al. GLABROUS (CmGL) encodes a HD-ZIP IV transcription factor playing roles in multicellular trichome initiation in melon. Theor. Appl. Genet. 131, 569–579 (2018).
Zhang, Y. et al. A GAMYB homologue CsGAMYB1 regulates sex expression of cucumber via an ethylene-independent pathway. J. Exp. Bot. 65, 3201–3213 (2014).
Chen, C. et al. The WD-repeat protein CsTTG1 regulates fruit wart formation through interaction with the homeodomain-leucine zipper I protein Mict. Plant Physiol. 171, 1156–1168 (2016).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium‐mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Wang, H. et al. Antisense suppression of cucumber (Cucumis sativus L.) sucrose synthase 3 (CsSUS3) reduces hypoxic stress tolerance. Plant Cell Environ. 37, 795–810 (2014).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq-a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676 (2005).
Quevillon, E. et al. InterProScan: protein domains identifier. Nucleic Acids Res. 33, W116–W120 (2005).
Du, Z., Zhou, X., Ling, Y., Zhang, Z. & Su, Z. agriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res. 38, W64–W70 (2010).
Wu, L. et al. An ethylene-induced regulatory module delays flower senescence by regulating cytokinin content. Plant Physiol. 173, 853–862 (2017).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (31872133), the Project for Scientific and Technological Activities of Overseas Students of Henan Province, the Zhongyuan Youth Talent Support Program (ZYQR201912161), and the Program for Science & Technology Innovation Talents in Universities of Henan Province (20HASTIT035).
Author information
Authors and Affiliations
Contributions
S.Y., K.Z., H.Z., and X.Z. performed phenotyping in F2 plants and fine mapping. L.Y., S.Y., H.Z., W.Y., N.X., Y.W., and J.H. contributed to data processing and analysis. S.Y. and D.L. contributed to the microscopic analysis. L.Y., S.Y., and Y.W. wrote the manuscript. All authors reviewed and approved this manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, S., Zhang, K., Zhu, H. et al. Melon short internode (CmSi) encodes an ERECTA-like receptor kinase regulating stem elongation through auxin signaling. Hortic Res 7, 202 (2020). https://doi.org/10.1038/s41438-020-00426-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-020-00426-6
This article is cited by
-
PmLBD3 links auxin and brassinosteroid signalling pathways on dwarfism in Prunus mume
BMC Biology (2024)
-
Identification and characterization of CsERECTA, a major gene controlling stem elongation through regulating GA biosynthesis in cucumber
Theoretical and Applied Genetics (2024)
-
Natural allelic variation in the EamA-like transporter, CmSN, is associated with fruit skin netting in melon
Theoretical and Applied Genetics (2023)
-
Mutation in the GA3ox gene governs short-internode characteristic in a korean cucumber inbred line
Horticulture, Environment, and Biotechnology (2023)
-
An LTR retrotransposon insertion inside CsERECTA for an LRR receptor-like serine/threonine-protein kinase results in compact (cp) plant architecture in cucumber
Theoretical and Applied Genetics (2023)