Abstract
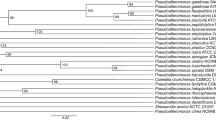
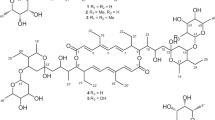
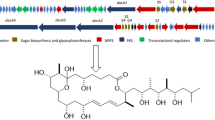
Two laxaphycin type-B cyclic dodecapeptides, laxaphycins B5 and B6, were obtained from UIC 10484, a freshwater cf. Phormidium sp. Analysis using the 16S rRNA sequence found UIC 10484 to clade with UIC 10045, a known laxaphycin type-A and -B producer, and MS/MS analysis revealed the presence of two novel laxaphycin type-B compounds. The structures of the metabolites were elucidated using 2D NMR and MS/MS. The absolute configurations of the amino acids were determined by advanced Marfey’s analysis. Both metabolites were evaluated against the same three cancer cell lines. The IC50 of both laxaphycins B5 and B6 was near 1 μM against breast cancer MDA-MB-231, melanoma MDA-MB-435, and ovarian cancer OVCAR3 cell lines.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Newman DJ, Cragg GM. Natural products as sources of new drugs from 1981 to 2014. J Nat Prod. 2016;79:629–61.
Singh RK, Tiwari SP, Rai AK, Mohapatra TM. Cyanobacteria: an emerging source for drug discovery. J Antibiot. 2011;64:401–12.
Luesch H, Moore RE, Paul VJ, Mooberry SL, Corbett TH. Isolation of dolastatin 10 from the marine cyanobacterium symploca species VP642 and total stereochemistry and biological evaluation of its analogue symplostatin 1. J Nat Prod. 2001;64:907–10.
Senter PD, Sievers EL. The discovery and development of brentuximab vedotin for use in relapsed Hodgkin lymphoma and systemic anaplastic large cell lymphoma. Nat Biotechnol. 2012;30:631–7.
Luo S, et al. Trichormamides C and D, antiproliferative cyclic lipopeptides from the cultured freshwater cyanobacterium cf. Oscillatoria sp. UIC 10045 Bioorg Med Chem. 2015;23:3153–62.
Frankmölle WP, Knübel G, Moore RE, Patterson GM. Antifungal cyclic peptides from the terrestrial blue-green alga Anabaena laxa. II: Structures of laxaphycins A, B, D and E. J Antibiot. 1992;45:1458–66.
MacMillan JB, Ernst-Russell MA, De Ropp JS, Molinski TF. Lobocyclamides A-C, lipopeptides from a cryptic cyanobacterial mat containing Lyngbya confervoides. J Org Chem. 2002;67:8210–5.
Bonnard I, Rolland M, Salmon JM, Debiton E, Barthomeuf C, Banaigs B. Total structure and inhibition of tumor cell proliferation of laxaphycins. J Med Chem. 2007;50:1266–79.
Luo S, et al. Trichormamides A and B with antiproliferative activity from the cultured freshwater cyanobacterium Trichormus sp. UIC 10339. J Nat Prod. 2014;77:1871–80.
Cai W, Matthew S, Chen QY, Paul VJ, Luesch H. Discovery of new A- and B-type laxaphycins with synergistic anticancer activity. Bioorg Med Chem. 2018;26:2310–9.
Maru N, Ohno O, Uemura D. Lyngbyacyclamides A and B, novel cytotoxic peptides from marine cyanobacteria Lyngbya sp. Tetrahedron Lett. 2010;51:6384–7.
Harada K, et al. A Method Using LC/MS for Determination of Absolute Configuration of Constituent Amino Acids in Peptide–Advanced Marfey’s Method. Tetrahedron Lett. 1995;36:1515–8.
Fujii K, Harada KI. A nonempirical method using LC/MS for determination of the absolute configuration of constituent amino acids in a peptide: Combination of Marfey’s method with mass spectrometry and its practical application. Anal Chem. 1997;69:5146–51.
Fujii K, Ikai Y, Mayumi T, Oka H, Suzuki M, Harada KI. A nonempirical method using LC/MS for determination of the absolute configuration of constituent amino acids in a peptide: elucidation of limitations of Marfey’s method and of its separation mechanism. Anal Chem. 1997;69:3346–52.
Luzzatto-Knaan T, et al. Digitizing mass spectrometry data to explore the chemical diversity and distribution of marine cyanobacteria and algae. Elife. 2017;6:1–20.
Sharp K, et al. Phylogenetic and chemical diversity of three chemotypes of bloom-forming Lyngbya species (cyanobacteria: Oscillatoriales) from reefs of southeastern Florida. Appl Environ Microbiol. 2009;75:2879–88.
Murakami M, Suzuki S, Itou Y, Kodani S, Ishida K. New anabaenopeptins, carboxypeptidaze-A inhibitors from the cyanobacterium Aphanizomenon flos-aquae. J Nat Prod. 2000;63:1280–2.
Matthew S, Ross C, Paul VJ, Luesch H. Pompanopeptins A and B, new cyclic peptides from the marine cyanobacterium Lyngbya confervoides. Tetrahedron. 2008;64:4081–9.
Gerwick WH, Jiang ZD, Agarwal SK, Farmer BT. Total structure of hormothamnin A, A toxic cyclic undecapeptide from the tropical marine cyanobacterium Hormothamnion enteromorphoides. Tetrahedron. 1992;48:2313–24.
Bornancin L, et al. Structure and biological evaluation of new cyclic and acyclic laxaphycin-A type peptides. Bioorg Med Chem. 2019;27:1966–80.
Grewe CJ. Cyanopeptoline und scytocyclamide: zyklische peptide 487 aus scytonema hofmanni PCC 7110 struktur und biologische Q8488 aktivität. Freiburg im Breisgau, Germany: Albert Ludwig University of Freiburg; 2005.
Bornancin L, et al. Isolation and synthesis of laxaphycin b-type peptides: a case study and clues to their biosynthesis. Mar Drugs. 2015;13:7285–7300.
Gbanktoto A, Vigo J, Dramane K, Banaigs B, Aina E, Salmon JM. Cytotoxic effect of laxaphycins A and B on human lymphoblastic cells (CCRF-CEM) using digitised videomicrofluorometry. Vivo. 2005;19:577–82.
Fujii K, Sivonen K, Nakana T, Harada K. Structural elucidation of cyanobacterial peptides encoded by peptide synthetase gene in Anabaena species. Tetrahedron. 2002;58:6863–71.
Bister B, et al. Cyanopeptolin 963A, a chymotrypsin inhibitor of Microcystis PCC 7806. J Nat Prod. 2004;67:1755–7.
Itou Y, Ishida K, Shin HJ, Murakami M. Oscillapeptins A to F, Serine Protease Inhibitors from the Three Strains of Oscilltoria agardhii. Tetrahedron. 1999;55:6871–82.
Okumura HS, Philmus B, Portmann C, Hemscheidt TK. Homotyrosine-containing cyanopeptolins 880 and 960 and anabaenopeptins 908 and 915 from Planktothrix agardhii CYA 126/8. J Nat Prod. 2009;72:172–6.
Guljamow A, et al. High-density cultivation of terrestrial Nostoc strains leads to reprogramming of secondary metabolome. Appl Environ Microbiol. 2017;83:1–15.
Stein JR, ed. Handbook of phycological methods. Cambridge: Cambridge University Press; 1973.
Chlipala G, Mo S, Carcache De Blanco EJ, Ito A, Bazarek S, Orjala J. Investigation of antimicrobial and protease-inhibitory activity from cultured cyanobacteria. Pharm Biol. 2009;47:53–60.
May DS, et al. Merocyclophanes C and D from the cultured freshwater cyanobacterium nostoc sp. (UIC 10110). J Nat Prod. 2017;80:1073–80.
Nübel U, Garcia-Pichel F, Muyzer G. PCR primers to amplify 16S rRNA genes from cyanobacteria. Appl Environ Microbiol. 1997;63:3327–32.
Ramos V, Morais J, Vasconcelos VM. A curated database of cyanobacterial strains relevant for modern taxonomy and phylogenetic studies. Sci Data. 2017;4:1–8.
Garrity GeorgeM, Boone DavidR, Castenholz RW. Bergey’s manual of systematic bacteriology. 2nd ed. New York: Springer; 2001.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–10.
Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33:1870–4.
Ren Y, et al. Cardiac glycoside constituents of streblus asper with potential antineoplastic activity. J Nat Prod. 2017;80:648–58.
Kaiser K, Benner R. Hydrolysis-induced racemization of amino acids. Limnol Oceanogr Methods. 2005;3:318–25.
Davidson I. Hydrolysis of samples for amino acid analysis. In: Smith BJ, editors. Protein Sequencing Protocols. Methods in molecular biology. vol 221, 2nd ed. Totowa, New Jersey: Humana Press Inc.; 2003. p. 111–22.
Acknowledgements
This research was supported by the NCI/NIH P01 CA12506 grant and The Office of the Director, NIH National Center for Complementary and Integrative Health (NCCIH) T32AT007533-05 training grant (PS). We thank Dr Jonathan Bisson and Dr Charlotte Simmer for their assistance using the Q-TOF mass spectrometer, Rojin Ahadi and Angel Antunez for isolating and culturing strain UIC 10484, Dr Ben Ramirez for his guidance using the UIC Center for Structure Biology NMR instrumentation, Dr Chen of the Research Resources Center’s Mass Spectrometry Core for use of the Agilent 6454 LC/Q-TOF, and Dr Young Jeong for access to their HPLC system. We’d also like to acknowledge the James R Fuchs lab of Ohio State University for synthesizing the NMeIle and 3OHLeu standards.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Dedicated to Professor William Fenical in recognition of his contributions to marine derived secondary metabolites
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Sullivan, P., Krunic, A., Burdette, J.E. et al. Laxaphycins B5 and B6 from the cultured cyanobacterium UIC 10484. J Antibiot 73, 526–533 (2020). https://doi.org/10.1038/s41429-020-0301-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-020-0301-x