Abstract
Pathogens or danger signals trigger the immune response. Moderate immune response activation removes pathogens and avoids excessive inflammation and tissue damage. Histone demethylases (KDMs) regulate gene expression and play essential roles in numerous physiological processes by removing methyl groups from lysine residues on target proteins. Abnormal expression of KDMs is closely associated with the pathogenesis of various inflammatory diseases such as liver fibrosis, lung injury, and autoimmune diseases. Despite becoming exciting targets for diagnosing and treating these diseases, the role of these enzymes in the regulation of immune and inflammatory response is still unclear. Here, we review the underlying mechanisms through which KDMs regulate immune-related pathways and inflammatory responses. In addition, we also discuss the future applications of KDMs inhibitors in immune and inflammatory diseases.
Similar content being viewed by others
Facts
-
Abnormal expression of KDMs play a crucial role in the pathogenesis of inflammatory diseases.
-
KDMs regulate the development and function of immune cells and immune-related signaling pathways, and influence the expression of inflammatory cytokines.
-
Targeting specific KDM inhibitors is potential therapeutic approach in immune and inflammation-related diseases.
Open questions
-
What are the consequences of substrates that specific inhibition of KDMs on immune cells and inflammatory diseases?
-
Which KDMs in immune cells promote TLRs response?
-
What are the specific protein substrates of KDMs in different immune pathways? What non-histone substrates can be targeted by KDMs?
-
Which KDMs are good drug targets for the treatment of inflammatory diseases?
Introduction
The immune response is critical in regulating the body’s immune homeostasis by protecting it from infections and injury [1,2,3]. Epigenetics included the dynamic regulation of different gene expression processes, such as DNA modification, chromatin remodeling, non-coding RNA regulation and histone modification. These processes were relevant to fine tuning the immune response by establishing specific gene expression patterns, through gene expression regulation at the transcriptional and post-transcriptional levels [4,5,6]. Histones were the first proteins identified as lysine methylation substrates, while enzymes mediating methylation and demethylation were called histone methyltransferases (KMTs) and histone demethylases (KDMs) [7, 8]. Histone methylation refered to the process of transferring methyl from methionine to histone amino acids lysine (K) or arginine (R). Demethylation of different sites could lead to different effects, including the inhibition or activation of target genes [9]. Interestingly, abnormal histone demethylation levels are closely associated with various inflammatory diseases, including kidney injury, liver injury, and autoimmune diseases [10, 11]. Increasing evidence had revealed that this modification controls cell development and regulates immune pathways [12].
In this review, we summarized the role of KDMs in immune cells’ development and the regulation of immune-related pathways, including Toll-like receptor (TLR), cGAS-STING, and IFN signal pathways. This review focuses on the role of the KDM families of proteins in immune cells and their effect on inflammation to provide a theoretical basis for developing small-molecule KDM inhibitors in treating inflammatory diseases.
Overview of KDMs
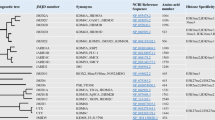
KDMs mediates the removal of methyl groups from histone lysine and arginine residues, a labile process that could regulate reversible chromatin marks [13, 14]. Histone lysine methylation had been determined as a reversible modification with the discovery of KDMs. More than 30 KDMs had been identified in humans and mice, with various motifs and essential functional domains for enzyme catalytic activity. KDMs were divided into two categories according to their catalytic mechanism (Fig. 1) :Lysine-specific demethylases (LSDs) and JmjC (Jumonji domain-containing protein) KDMs [7, 15,16,17]. LSDs were KDMs lacking the JmjC domain. The LSDs family were currently constituted by two members only, LSD1/KDM1A and LSD2/KDM1B [18, 19]. KDM1 had a C-terminal amine oxidase-like domain that contained the substrate and flavin adenine dinucleotide binding sites, which process the removal of monomethylated and dimethylated markers [20, 21]. KDM1 cannot remove trimethyl groups [22]. On the other hand, the KDM2-8 proteins contain a JmjC catalytic domain of ~170 amino acids with substrate specificity and demethylase activity that could remove mono-, di-, and trimethyl groups on specific sites of lysine residues [23]. This domain can remove methylation from histone H3 on lysine residues K4, K9, K27, and K36 and histone H4-K20 (Fig. 2). The JmjC domain had a similar structure to that of the Cupin protein family and contains oxygenase-catalyzed domains dependent on Fe(II) and α-KG [24]. This domain also contained a zinc finger (ZF), which brought the JmjC domain close to the C-terminus domain and was indispensable for the stability and function of KDMs. Besides the JmjC catalytic domain, all KDM proteins contain domains that interacted with the N-terminus of JmjN, providing structural integrity without catalytic function [25]. KDMs included Swi3p, Rsc8p, Moira domains, AT-rich interaction domains (ARID), plant homology domains (PHD), ZF, and Tudor domains. In addition, the F-box, Leucine Rich Repeat, and tetrapeptides domains play critical roles in their protein–protein interactions [24].
The key domains of each KDMs are represented in colored regions, and the number of amino acids is written in each functional domain. SWIRM Swi3p, Rsc8p, and Moira domain; JmjC Jumonji C domain, JmjN Jumonji N domain, ARID AT-rich interacting domain, PHD plant homeodomain domain, F-box F-box domain, LRR Leu-rich repeat domain, TPR tetratricopeptide domain, Tudor Tudor domain, CW CW-type zinc-finger domain, C6 C6 zinc-finger domain, CXXC CXXC zinc-finger domain, C5HC2 C5HC2 zinc-finger domain.
The first discovered histone demethylase containing a JmjC domain is the F-box and leucine-rich repeat protein 11 (also known as KDM2A). KDM2A had specific JmjC-dependent demethylation activity for H3K26me3 [26, 27]. Other domains of KDM2A include a ZF domain, a PHD domain, and three leucine-rich repeats. It was reported that KDM2A was recruited to CpG islands via the ZF domain, resulting in a unique chromatin state by removing the H3K36me2 modification [28]. The catalytically active ZF, ARID, and PHD domains in KDM5 separate their catalytic cores into N-terminus and C-terminus to improve substrate specificity [29, 30]. The N-terminal PHD domain binds to H3K4me, while the C-terminal PHD domain bound to H3K4me3/me2 [29]. In KDM5B, ARID was the primary DNA-binding interface through the L1 ring, which recognized the GCACA/C sequence, whereas PHD1 was involved in histone recognition and may be inhibited demethylase activity [29]. In contrast, the PHD of KDM7 had been reported to be non-essential for the catalytic activity, but was essential for substrates specificity [31]. For instance, the JmjC and PHD domains of KDM7B bind to H3K4me3, enhancing the demethylation rate on its homologous substrate, H3K4me2 [31, 32]. In addition, KDMs contain the Tudor domain, consisting of tubular structures with antiparallel beta chains present in many chromatin-associated with proteins [33]. KDM4A-C contained a conserved dual Tudor domain that determined the different binding preferences of these enzymes [34]. On the contrary, KDM4D and KDM4E do not contained a Tudor domain and used specific structures within the JmjC domain to recognize lysine [35].
Regulation of immune cell development by KDMs
Immune cells are critical components of the immune system, producing various kinds of cytokines, protecting against pathogens and clearing “nonself” substances to maintain the homeostasis of the immune system [36,37,38,39,40,41]. The development of hemopoietic stem cells (HSCs) into different immune cells, such as T lymphocytes, NK cells, B lymphocytes, and macrophages, involve selective gene expression patterns regulated by intricate mechanisms [42]. The development of many diseases is related to disequilibrium in the quantities or functions of immune cells, where epigenetic regulation plays an important role. KDMs modulate cellular differentiation by regulating the activity of cell-specific gene enhancers [43]. In addition, KDMs played a role in the lineage commitment of immune cells after stimulation by pathogens or other signals by regulating genes expression via histone demethylation [44]. Studies have shown that KDMs were essential players in innate and adaptive immunity and participate in antivirus and antitumor immunity [45,46,47,48]. KDMs control the maturation, differentiation, and function of immune cells, thus participating in the immune response and maintaining homeostasis (Fig. 3).
The KDM6 subfamily is important for the development of T lymphocytes. KDM6A and KDM6B promote the expression of S1pr1 and Klf2, crucial for the maturation of T cells, by targeting promoter H3K27me3. Subsequently, KDM6B modifies the expression of Th17-related genes including Il22, Il17f, Il17, Rorγt, and Th1-related genes including T-bet and Smad3, promoting the differentiation into Th17 and Th1 cells, respectively. During the differentiation of B cells, KDM1A takes part in repressing PU.1, IRF4 and Blimp-1 through decreasing H3K4me3, which is crucial for the formation of plasmablasts. KDM5A recruited by p50 is responsible for the downregulation of Socs1 through erasing H3K4me3 in NK cells, inducing IFN production. In macrophages, KDM3C modulates H3K9me of the miR-302a promoter and inhibits M1 macrophage differentiation via the miR-302a/METTL3 axis. Finally, the KDM6B/IFR4 axis promotes the differentiation of M2 macrophages.
Functions of KDMs in T lymphocytes development
T lymphocytes play a crucial role in regulating the immune system [49,50,51]. The activation of T cells must be maintained at an appropriate level to respond against pathogens and avoid overactivation and autoimmune diseases [52]. T cells were derived from lymphoid progenitors in the bone marrow, generating TCR in the thymus and developing into CD4+ or CD8+ single-positive T cells [53]. Upon stimulation by pathogens or cytokines, mature T cells in the peripheral circulation differentiate into various helper T cells (Th cells) or cytotoxic T lymphocytes (CTLs), a process widely believed to be regulated by KDMs [54]. Indeed, KDMs were shown to be associated with T cell maturation in the thymus. Histone demethylation promoted T cell maturation by suppressing marker genes of T cell precursors and inducing T cell-specific gene expression [48]. Knockdown of KDM1A inhibited the downregulation of Ctla4 and Prdm1, explicitly expressed in double negative T cells (CD4−CD8−), blocking the formation of double positive (CD4+CD8+) or CD4+ T cells [55]. It was also reported that KDM6A and KDM6B could induce the expression of S1pr1, a critical protein for T cell maturation, and the transcription factor Klf2 via H3K27me3 demethylation, inducing intrathymic T cell precursors to differentiate into terminal T cells [56].
Furthermore, KDMs are involved in the differentiation of CD4+ T cells into various subsets of Th cells that produce different cytokines [57, 58]. Wei et al. [59]. discovered that KDM1/5/6 targeted H3K4me3 and H3K27me3 and were associated with early T cell and Th cell differentiation by inducing the expression of signature genes in Th cells, such as Ifng, Il4, Il17, and Tbx21. Suzuki et al. [60]. reported that KDM1A increased the expression of transcription factors, including Tbx21, Eomes, and Runx2, which promote the development of Th1 cells and the production of IFN-γ. Similarly, Liu et al. [61]. indicated that the knockdown of KDM1A could inhibit the proliferation of CD4+ T cells, and decrease the secretion of IFN-γ and IL-17. Meanwhile, KDM5C was found to be downregulated in CD4+ T cells from multiple sclerosis (MS) patients, facilitating Th17-mediated IFN-γ secretion [62]. In addition, by interacting with lncRNA A112010, Yang et al. [63]. found that KDM5A enhanced the production of pro-inflammatory Th1 and Th17 cells by decreasing H3K4me3 and suppressing the transcription of IL-10. Furthermore, Ptaschinski et al. [64]. have shown that upon infection with the respiratory syncytial virus, ablation of KDM5B might enhance the production of IFN-β, IL-6 and TNF-α by increasing H3K4me3 in their promoters. Besides, the expression of Th2 cytokines was downregulated, resulting in chronic infection. Similarly, in the widely used autoimmune encephalomyelitis, MS mouse model, knockout of KDM6A limited the development of the autoimmune disease by suppressing CD44 expression on CD4+ T cells via H3K27me3 targeting Th2 cytokines expression and Th1 response blockade [65]. Using a colitis model, Qing et al. [57]. revealed that KDM6B could facilitate H3K27me3 demethylation on Th1 transcription factors, including Foxp3, Ifng and Cd44, while its deletion inhibited the transition of intestinal CD4+ T cells into Th1 cells. Furthermore, Liu et al. [66]. discovered that KDM6B was rapidly induced upon stimulation of TCR signaling. Overexpression of KDM6B decreased H3K27me3 in the Rorγt promoter, a crucial Th17 transcription factor, promoted Th17 differentiation, and induced the expression of cytokines such as Il17, Il17f, and Il22. However, depletion of KDM6B or treatment with GSK-J4, a KDM6A/6B specific inhibitor, reduced Rorγt expression, suppressing the generation of Th17 cells [66, 67]. Finally, KDM6B inhibited TGF-β-induced FOXP3 expression and the development of Treg cells and facilitated differentiation into pro-inflammatory phenotypes, including Th17 or Th1, thus aggravating inflammatory responses [68].
Besides CD4+ T cells, KDMs regulated the development of CD8+ and other subsets of T cells. After viral infection, KDM6B was rapidly upregulated in CD8 + T cells, decreasing H3K27me3 markers and enhancing CTL-mediated immune response and the formation of memory T cells [69, 70]. Similarly, depletion of KDM6A in mice treated with Listeria monocytogenes expressing OVA enhanced immune response after secondary infection by reducing CTLs formation through H3K27me3 regulation on Prdm1, which encoded Blimp1 and played essential roles in the development of CD8+ memory T cells [71]. In addition, KDM6B was shown to enhance the production of mucosal-associated invariant T cells (MAIT), a special type of T cells that produced Th1-like effectors. Indeed, the KDM6B co-factor α-KG was shown to enhance the effects of MAIT [72]. Intestinal intraepithelial lymphocytes (IELs) were another type of T -cells located in the intestinal epithelium. Knockout of KDM6B reduced the number of TCRαβ + CD8αα + IELs in mice, and impaired the lytic function of IELs through H3K27 demethylation on Bcl2 and FasL [44].
The role of KDMs in the development and function of B lymphocytes
B lymphocytes were essential effectors in humoral immunity, and epigenetic regulation is required for the proliferation and differentiation of B cells [73,74,75,76]. Upon activation by thymus-dependent antigen, naïve B cells (nB) transform into plasmablasts (PB) to form germinal centers (GC), from which antibody-secreting plasma cells and memory B cells were developed. Haines et al. [77]. found that the knockdown of KDM1A in mice treated with LPS led to a shift in H3K4me1 levels in cell cycle genes and transcription factors, including PU.1, IRF3 and Blimp-1, thus suppressing the differentiation of nB into PB.
The proliferation and differentiation of B cells in GC require the assistance of Th cells, mainly T follicular helper (Tfh) cells that play indispensable roles. Hung et al. [78]. indicated that Tfh-derived signaling upregulated KDM4A and KDM4C expression, blocking normal cell cycle and inhibiting the proliferation of B cells by interacting with WDR5. Besides, depletion of KDM6A in T cells was shown to inhibit the expression of interleukin-6 receptor-α, Icos, and other genes related to the development of Tfh cells [79]. Consequently, the decreased in Tfh cell population impaired ability of B cells to produce specific IgG antibodies in response to lymphocytic choriomeningitis virus (LCMV) infection. Kei et al. [80]. highlighted the role of KDM6A in GC B cell maturation, showing that IL-4 stimulation activates the STAT6 pathway, which induces KDM6A recruitment into Bcl6 enhancer, resulting in H3K27me3 demethylation and Bcl6 induction, thereby facilitating the formation of mature B cells in GC. Interestingly, the knockout of KDM1A restrained the Bcl6-derived proliferation of GC B cells. Interestingly, BCL6 could then form a suppressive complex with KDM1A and control the regulation of Bcl6-targeted genes. Similarly, KDM6B was also associated with B cell development [81], whereby KDM6B transcription was reported to be upregulated in GC B cells compared to undifferentiated B cells [82].
The role of KDMs in the polarization and function of macrophages
Macrophages are key components of innate immunity that regulate tissue homeostasis and participate in tissue repair by phagocytosis and clearing the debris of dead cells [83,84,85,86]. Macrophages could be polarized into functionally distinct phenotypes. Classically activated (M1) macrophages have pro-inflammatory effects, while alternatively activated (M2) macrophages manifest anti-inflammatory functions and promote tissue repair [87,88,89,90]. KDMs were involved in macrophage polarization. Indeed, KDM3C was shown to mediate M1 polarization and suppress glioma by stimulating the miR-302a/METTL3/SOCS2 axis [91]. KDM3C increased the expression of miR-302a by H3K9me1 demethylation in its promoter region, miR-302a targeted METTL3, which in turn inhibits SOCS2 via an m6A modification. While KDM6B did not influence M1 polarization, it was essential for M2 macrophage differentiation. Investigations on anti-helminth immunity revealed that KDM6B was a positive regulator of M2 activation by upregulating the transcription factor interferon regulatory factor 4 (IRF4) via H3K27me3 demethylation [92, 93]. Besides, the supplement of alpha-ketoglutaric acid (α-KG), in inflammatory diseases stimulated by the granulocyte-macrophage colony-stimulating factor, facilitated IL-4 induced M2 polarization via the KDM6B/IRF4 axis, while IFN-β reversed macrophage activation [94, 95]. In addition, stimulation of macrophages with LPS or cytokines activated of the NF-κB signaling pathway and enhanced the expression of KDM6B. The latter combined with PcG-target genes to modify downstream gene expression via H3K27me3 demethylation, fortifying macrophages’ response against inflammatory stimulation [96]. Interestingly, Huang et al. [97]. demonstrated that LPS induced the upregulation of KDM6B in bone marrow-derived macrophages, leading to decreased H3K27me3 on Nrf2 promoter, improving NLRP3 inflammasome activation in macrophages. Indeed, KDM6B-specific inhibitor GKS-J4 restrained this process and inhibited the development of colitis in sodium dextran sulfate-treated mice. Similarly, Zhuo et al. [98]. showed that the expression of KDM1A was induced, and the NLRP3 inflammasome was activated in RAW264.7 macrophages treated with pro-inflammatory Ox-LDL. Conversely, inhibition of KDM1A increased SESN2 and activated the PI3K/Akt/mTOR pathway, ultimately decreasing inflammation. In Leishmania donovani-infected macrophages, activated HIF-1α was shown to regulate macrophages polarization by inducing the expression of KDM5B and KDM6B, modifying H3K4 and K3K27 trimethylation and consequently suppressing M1 cytokines (TNF-α and IL-12) and promoting M2 factors such as Arg-1 [99].
The role of KDMs in the development and function of NK cells
Natural killer (NK) cells are cytotoxic lymphocytes that rapidly fight against pathogens to maintain homeostasis [100,101,102,103]. NK cells play important roles during early viral infection by releasing granules containing perforin and granzymes and secreting critical antiviral cytokines such as interferon-γ (IFN-γ) [104,105,106]. Bailey et al. [107]. showed that KDM1A regulated the production of reactive oxygen species and glutathione in NK cells, while SP-2577, an inhibitor of KDM1A, weakened the cytotoxic function of NK cells by impairing oxidative phosphorylation and glycolysis. In addition, Zhao et al. [108]. observed an upregulation of KDM5A in NK cells infected with monocytogenes (Lm) via p50 recruitment, leading to Socs1 transcription suppression by H3K4me3 demethylation on its promoter regions. Meanwhile KDM5A activated the JAK2-STAT4 signaling pathway to induce the secretion of IFN-γ. Studies by Adam et al. [109]. indicated that KDM6B from peripheral blood or tissues of rheumatoid arthritis (RA) patients was upregulated compared to healthy controls. Furthermore, GSK-J4, an inhibitor of the H3K27me3 demethylase KDM6B, inhibited the cytokine-stimulated production of IFN-γ, TNF-α, GM-CSF, and IL-10, thus suppressing the inflammatory response in NK cells.
The invariant natural killer T (iNKT) cells are the most well-studied type of NKT cells. They had the unique ability to recognize lipid antigens presented by CD1d on the surface of antigen-presenting cells [110, 111]. Interestingly, KDM6A was enriched in the promoter of PLZF in double-positive (CD4+CD8+) T cells, while knockout of KDM6A enhanced the repressive H3K27me3 and inhibited the transcription of PLZF, which controls the production of specific TCR of lipid antigens on NKT cells [112]. Similarly, Beyaz et al. [113]. showed that KDM6A inhibited the differentiation of glycosphingolipid α-galactosylceramide (α-GalCer)-induced mouse iNKT cells by decreasing H3K27me3 in the promoters of iNKT signature genes such as Tbx21, Il2rb and Klrd1. KDM6A has been also shown to interact with JunB to establish lineage commitment of iNKT cells. Finally, studies by Northrup et al. [114]. demonstrated that Kdm6a/Kdm6b depletion downregulated cell cycle-related genes in NKT cells, alleviating hepatic injury in ConA-induced mice liver injury model.
KDMs in the immune response and related signaling pathways
Immune responses depend on how immune cells react to stimuli. Pathogens or inflammatory signals recognized by pattern recognition receptors (PRRs) or cytokine receptors are transduced along various signaling pathways, activating nuclear transcription factors and inducing the production of effector molecules to stimulate the immune response. KDMs are involved in the regulation of gene transcription and thereby participate in the control of immunity (Fig. 4).
KDMs regulate TLR, cGAS-STING, and IFN signaling pathways to modulate inflammation and antiviral responses. (1) TLRs signaling pathway. KDM4D promotes TLR4 transcription by demethylating H3K9, thereby activating the TLR4/TIRAP/MyD88/NF-κB signaling pathway, which in turn promotes the expression of hepatic fibrosis factors COL1A1, ACTA2, VIM, and inflammatory cytokines TNF-α and IL-1β. Increasing the expression of KDM5A can inhibit Pten, activate PI3K/AKT/S6K1/PD-L1 and TLR7/8 signaling pathways, and promote the expression of cytokines and chemokines. (2) After viral infection of cells, KDM5B/5 C can inhibit STING expression by demethylation of H3K4me3, while KDM5 inhibitors increase IFN levels and improve antiviral immune response by enhancing the cGAS-STING-TBK1-IRF3 pathway. (3) Inhibition of KDM1A in cells induces the level of SESN2, which activates the autophagy-related PI3K/Akt/mTOR pathway, ultimately inhibiting the activation of the NLRP3 inflammasome. At the same time, it increases KDM6B and promotes ASC/NLRP3/Caspase, hence promoting the inflammatory response.
KDMs regulate the recognition and transduction of stimulatory signals
PRRs are important receptors on innate immune cells, which include TLRs, RNA helicases retinoic acid-inducible gene I that recognizes cellular RNA, and the cGAS-STING pathway recognizing DNA [115, 116]. Among these, TLR9 was shown to bind to the CpG motif of viral double-stranded DNA and plays critical roles in anti-viral immunity. Hasan et al. [117]. found that KDM5B was recruited by the E7 protein of HPV16 via NF-κB signaling pathway in infected human epithelial cells. KDM5B interacted with HDAC1 to downregulate TLR9 expression by modulating its promoter activity, thus suppressing viral infection. As for the regulation of the cGAS-STING pathway, Wu et al. [118]. reported that KDM5B/5C suppressed the expression of STING after viral infection by removing H3K4me3 modifications. An inhibitor of KDM5 activated the cGAS-STING-TBK1-IRF3 axis to boost IFN production and anti-viral immune response. Wang et al. [119]. indicated that KDM5A activated the TLR7/8 and PI3K-AKT-S6K1 pathways to promote PD-L1 expression. CCl4-induced liver fibrosis studies by Dong et al. [120]. showed that upregulation of KDM4D catalyzed H3K9 demethylation of TLR4 promoters, which in turn activated NF-κB signaling pathway to induce fibrosis. Taken together, these studies support that KDMs regulated stimuli recognition by PRRs.
The transduction of PRR signals relies on different cellular pathways, and KDMs play two roles in this process [121]. First, KDMs influence the activation of these pathways. For instance, in NK cells stimulated with Lm infection or IL12/18, KDM5A was recruited to the promoters of SOCS1 and inhibited its expression via H3K4me3 demethylation, which activated STAT4 signaling and led to IFN-γ production [108]. Similarly, TFN-α-induced endothelial inflammation was shown to be dependent on the activation of Jagged-1 and the Notch activator ADAM17 and by KDM6A/6B-induced H3K27me3 demethylation [122]. In addition, KDM7B inhibited IFN-γ-target genes by removing H4K20me1 and blocking the transduction of IFN-γ signals [123]. Second, KDMs regulated several transcription factors, and NF-κB was one of the most critical effectors mediating TLR or cytokine-target genes. Kim et al. [124]. indicated that in sepsis combined with lung injury, activated PKCα was transported into the nucleus and induced the interaction of KDM1A with p65, which led to p65 demethylation, suppressing inflammatory response via PKCα-KDM1A-NF-κB axis and ultimately reducing the death rate in mice. LPS could also induce the expression of FBX11 (KDM2A) via NF-κB pathway. However, KDM2A targeted the lysine K218 or K221 of p65 and inhibited p65 expression through demethylation on those sites, constituting negative feedback [125]. Through association with deubiquitinase USP38, KDM5B antagonized NF-κB signaling and inhibited LPS-stimulated activation of Il6 and Il23a promoters [126]. Connor et al. [127]. demonstrated that KDM6B was upregulated in epithelial cells infected with S. pneumoniae. Moreover, KDM6B removed H3K27me2 and recruited p65 to the Il11 promtoer, regulating the production of IL-11 and other cytokines including IL-1β and TNF-α. Studies of Higashijima et al. [128]. showed that KDM6A and KDM7A synergistically occupied the NF-κB binding site in TNF-α-induced endothelial cells, enhancing the expression of p65 target genes through the regulation of H3K9me2 and H3K27me3. Similarly, KDM7C was recruited by p65 to the promoters of Tnf, Ccl4 and Il11 upon LPS stimulation, which activated their expression by removing H4K30me3 [129]. Finally, KDM4A was also recruited by p65 after viral infection, inducing the production of INF-β through H3K9me3 demethylation [130].
KDMs mediate the expression and function of effector molecules
KDMs positively regulate inflammatory factors [131]. During the process of innate immunity, KDM6A/6B removed the repressive H3K27me3 and promoted IL-6 mediated inflammation [132,133,134]. Indeed, the KDM6 inhibitor GSK-J4 was shown to suppress TNF-α production upon LPS stimulation possibly via the regulation of TNFA transcription by KDM6A/6B [135]. During the development of dendritic cells (DCs), GSK-J4 significantly decreased the expression of the co-stimulatory molecules CD80/CD86, upregulated anti-inflammatory TGF-β1 and down-regulated pro-inflammatory IL-6, thus suppressing the activation of DCs [134, 136]. GSK-J4 also promoted IFN-γ secretion, since KDM6B get recruited by Cbx2 to IFN promoter to enhance its transcription through H3K27me3 demethylation [109, 137]. KDM6A was also shown to interact with the methyltransferase MLL4 and regulated the enhancers of Ifnb, thus promoting IFN-β expression [133]. In addition, KDM2B could interact with Brg1, which was the core component of the SWI/SNF complex, increasing chromatin accessibility of the Il6 promoter and inducing its expression in macrophages and DCs [131].
Conversely, KDM1 and KDM5 proteins that targeted the promoting H3K4 methylation often disrupt chromatin accessibility and inhibited gene expression [138, 139]. KDM1A could directly inhibit IL-1β, IL-6, IL-8 and the classic complement components, working synchronously with HDAC1 to restrain inflammation [140]. The endotoxic shock was shown to cause KDM1A suppression, which protected against inflammatory response over-activation, leading to excessive proliferation of myeloid progenitors, producing IL-1β and TNF-α and forming cytokine storms [138]. Furthermore, KDM5C could interact with Polycomb group factor 6 to regulate the H3K4me3 of Il11b and Ciita, suppressing the activation of DC and T cells [139]. By stimulating mouse embryonic fibroblasts with poly(I:C), Yu et al. [141]. showed that KDM5A was recruited to the IFN-β promoter, inhibiting its transcription via H3K4me3 demethylation.
KDMs and inflammatory diseases
There was growing evidence that abnormal expression of KDMs was associated with a variety of inflammatory diseases, including mastitis, liver fibrosis, inflammatory kidney injury, inflammatory pulmonary injury, colitis, autoimmune diseases, and inflammation-related cancers [96, 142,143,144]. The expression of KDMs was up-regulated or down-regulated in these diseases, and the transcription and secretion of inflammatory mediators were regulated through specific demethylation or activation of inflammation-related pathways, thus promoting the occurrence and development of inflammatory diseases [133, 145,146,147,148] (Table 1). Since there are so many inflammatory diseases that may involve the dysfunction of KDMs, this review we discuss six of the following inflammatory diseases.
Mastitis
Mastitis was a disease characterized by the inflammation of the breast parenchyma [149, 150]. In a mouse model of mastitis, LPS was shown to activate the NF-κB signaling pathway through TLRs recognition on mammary epithelial cells, promoting the production and release of inflammatory factors [151]. In a similar mouse model of mastitis induced by LPS and respective in vitro cell experiments, Wang et al. [152]. found that the expression of KDM6B in mammary epithelial cells was increased, and H3K27me3 demethylation in the promoter region induced the expression of TLR4 and activated downstream transduction of pro-inflammatory NF-κB signaling. Similarly, GSK-J1 suppressed KDM6B and therefore decreased the expression of inflammatory cytokines TNF-α, IL-1β and IL6. KDM1A was also a key pro-inflammatory regulator of mastitis, as demonstrated by Wang et al. [153] These authors had found that a KDM1A inhibitor (GSK-KDM1A) could up-regulate histone H3K4me2 and H3K9me2, which inhibited the NF-κB signaling pathway to reduce the expression of inflammatory factors including TNF-α, IL-6, and IL-8, thus reducing the inflammatory reaction in mammary epithelial cells.
Hepatic fibrosis
The central step of hepatic fibrosis development was the transformation of hepatic stellate cells (HSCs) into activated myofibroblasts, which played an important role in the regulation of hepatic fibrosis [154,155,156,157,158,159]. Studies have shown that KDMs control the activation of HSCs [160, 161]. Yan et al. [162]. proposed that KDM6B was a pivotal negative regulator of CCl4 and BDL-induced mouse liver fibrosis. On the one hand, KDM6B inhibited TGF-β/SMAD signaling by increasing BAMBI expression, thus reducing the transformation of HSCs. On the other hand, KDM6B promoted cell senescence by upregulating p21 and Gadd45, and suppressed the expression of extracellular matrix (ECM) proteins and α -smooth muscle actin. Ming et al. [163]. found that KDM4 was down-regulated in HSCs with liver fibrosis, and the overexpression of KDM4 could synergize with SREBP2 to activate miR-29 transcription and inhibit HSCs activation. Furthermore, Li et al. [164]. observed that Brg1 recruited KDM4 and interacted with β-catenin to enhance Wnt signaling in hepatocytes, thus promoting liver regeneration after injury. In contrast, Fang et al. [120]. found that the expression of KDM4D was up-regulated during HSC activation, which further activated the TLR4/NF-κB signaling pathway and promoted liver fibrosis. In addition, Yan et al. [165]. showed that in a CCl4-induced mouse liver fibrosis model, the expression of KDM3A in HSCs was down-regulated, whereas KDM3A overexpression induced the demethylation of H3K9me2 in the PPARγ promoter, leading to inhibition of HSC activation. A recent study supported that KDM5 proteins were key factors for sex differences in alcohol-related liver disease (ALD). Indeed, KDM5B and KDM5C promoted liver fibrosis through increasing transcription of fibrosis and inflammation related genes, such as Col3a1, Itgav, and Gabarapl1 and down-regulating the expression of AhR only in female mice and female ALD patients [166].
Inflammatory kidney injury
Kidney fibrosis was generally believed to be central to the progression from chronic kidney injury to end-stage renal disease [167], and it had been reported that the renal fibrosis signaling pathway was regulated by epigenetic mechanisms [168, 169]. Indeed, Yu et al. [170] showed that the expression of KDM6B was increased in a mouse model of renal fibrosis, inhibiting DNA methyltransferase 1 to promote Smad7 expression, and ultimately decreasing the TGF-β1/Smad3 pathway. On the other hand, KDM6B can upregulate FBXW7, a negative regulator of Notch, inhibiting the Notch signaling pathway and ultimately exerting anti-fibrotic effects. Likewise, He et al. [171] showed that KDM6B and miR-93-5p were up-regulated in mice with acute kidney injury (AKI) and that the expression of inflammatory factor TNF-α was reduced through demethylation of H3K27me3, thus alleviating kidney injury. Feng et al. [172] found that low expression of KDM6B could promote neointimal hyperplasia and inflammatory cell infiltration, which might cause vascular stenosis and access failure in renal dialysis patients. Chen et al. [173] found that KDM6A was a key pro-inflammatory factor of diabetic nephropathy (DKD) in db/db mice. KDM6A upregulation in the kidneys of diabetic mice promoted the expression of various inflammatory factors such as IL-1β, IL-8 and IL-6, leading to the deterioration of DKD. Liu et al. [174] demonstrated that dexmedetomidine (DEX) could alleviate LPS-induced AKI in mice by inhibiting the NF-κB signaling pathway and the expression of KDM5A. In addition, in HBV-infected HK-2 cells, enrichment of KDM1A reduced the level of H3K9me1/2 in the TLR4 promoter region, leading to the production of inflammatory cytokines through the TLR4-NF-κB/JNK axis and renal inflammatory response exacerbation [175].
Inflammatory pulmonary injury
Asthma was a chronic inflammatory disease of the airways, where exposure to allergen activate immune cells and triggered inflammation and immune responses [176,177,178]. Bajbouj et al. [179] showed that treating lung fibroblasts with IL-13 induced the expression of KDM4B and H3K36me3 demethylation, thus promoting nuclear translocation. Meanwhile, early inhibition of KDM4B activity could postpone or even prevent airway fibrosis in asthmatic patients. In a mouse asthma model established by Yu et al. [180], a KDM6B inhibitor (GSK-J4) was shown to reduce airway inflammation, hyperreactivity and remodeling by blocking Akt/JNK and TGF-β/Smad3 signaling. In addition, it was showed that DEX could promote the expression of keratinocyte growth factor (KGF-2) by down-regulating KDM6B and ameliorating endothelial barrier dysfunction in ischemia/reperfusion related lung injury [181]. He et al. [182] found that Cu, Zn-SOD could activate STAT6, which stimulated the expression of KDM6B and interacted with KDM6B to induce profibrotic M2 gene promoters, hence stimulating macrophage polarization to the M2 phenotype and exacerbating pulmonary fibrosis. In addition, Fraszczak et al. [183] found that mice with impaired binding of GFI1 to KDM1A, displayed increased expression levels of serum IL-6, TNF-α, and IL-1β, while their alveolar macrophages secreted a large amount of cytokines that ultimately led to increased mortality.
Colitis
A large amount of evidence suggests that histone demethylation played an important role in intestinal inflammatory responses and the transformation to colorectal cancer (CRC) [184]. Ma et al. [185] showed in necrotizing colitis mice that KDM6B was activated by STAT3 and participated in JAK2/STAT3 pathway to enhance the transcription of inflammatory genes. Besides, KDM6B might interact with NF-κB to activate the transcription of TNF-a/Il1b/Il6 and ultimately aggravated intestinal inflammatory damage. Moreover, Zhuo et al. [186] found that TNF-α induced high expression of KDM4D in colitis, which activated Hedgehog signaling to promote colon regeneration and CRC cell proliferation. Moreover, Sun et al. [187] reported that AMPK could recruit KDM1A to the Cdx2 promoters, upregulating Cdx2 expression and enhancing intestinal barrier function and epithelial differentiation. Finally, Parmar et al. [188] demonstrated that KDM1A was required to restore the intestinal barrier against pathogens invasion by promoting the maturation of intestinal epithelial goblet cells in mice with colitis induced by bacterial or parasitic infection.
Autoimmune diseases
The role of KDMs in gene expression regulation played an important role in the occurrence and development of autoimmune diseases [189]. Rheumatoid arthritis (RA) was an aggressive joint disease resulting from immune imbalance and excessive inflammatory responses. Wan et al. [190] showed that the up-regulated expression of KDM6B in RA promoted arthritis development by inducing the transcription of PCNA, which mediated the migration and proliferation of fibroblast-like synoviocytes (FLS). Subsequent studies showed that GSK-J4, a KDM6A/6B inhibitor, suppressed the expression of RANKL and the production of TNF-α and GM-CSF in NK cells, reducing osteoclast generation and inflammatory response [109]. Wu et al. [191] found that in synovioblasts of RA patients, cystathionine-gamma-lyase inhibited the progression of joint inflammation by reducing the expression of KDM6B, suppressing the transcription of TLR2 and some inflammatory factors including TNF-α and IL-6. Meanwhile, Zhao et al. [192] showed that in collagen-induced arthritis (CIA) mice, GSK-J4 inhibited IL-6 transcription in macrophages, thus alleviating RA symptoms. Likewise, Zhang et al. [193] found that KDM4B promoted the secretion of various pro-inflammatory factors and aggravated RA by activating STAT3 signaling pathway in FLS. Heng et al. [194] clarified that in systemic lupus erythematosus, the expression of KDM6B was increased, and its H3K27me3 demethylation activity enhanced the expression of CD11a, thus promoting T cell auto-reactivity and production of autoantibodies. MS is an autoimmune disease of the central nervous system, mediated by T cells. High expression of KDM6A on the X chromosome was found in an MS mouse model, which could explain the high prevalence of this disease in women. KDM6A could also downregulate Th1 and Th2 activation pathways in mouse CD4+ T cells, promoting neuroinflammatory signaling via TLRs and IL-17 [65].
KDMs inhibitors in the treatment of immune-related diseases
Many small-molecule inhibitors of KDMs have been developed in recent years with the understanding of the structures and mechanisms of KDMs [195]. According to existing research, some molecules have broad prospects for clinical use, such as the KDM1A inhibitor TCP, the KDM6-specific inhibitor GSK-J1/4 and the inhibitor of JmjC family JIB-04 (Table 1).
TCP was first approved for clinical use as a monoamine oxidase inhibitor to treat depression and was later shown to inhibit KDM1A as well [196]. Motivated by these discoveries, further studies developed dozens of inhibitors with increased specificity for KDM1A, some of which have entered clinical trials including ORY-1001 (Iadademstat), ORY-2001 (Vafidemstat), IMG-7289 (Bomedemstat), and CC-9001 [197, 198]. Studies on these drugs mainly focused on solid tumors such as small cell lung cancer and hematological diseases such as non-Hodnkin lymphoma and acute myeloid leukemia [199,200,201,202,203]. Despite the lack of clinical studies, KDM1A inhibitors have shown significant benefits in pre-clinical models of inflammatory diseases. KDM1A was shown to be upregulated and was associated with excessive inflammation in patients with hepatitis B virus (HBV)-associated glomerulonephritis. While in HBV transgenic mice, TCP treatment blocked the TLR4-NF-κB-JNK pathway and reduced inflammatory response [175]. ORY-2001 was shown to alleviate neuroinflammation by preventing the development of demyelination and inhibiting T cell infiltration in the spinal cord, hence could be beneficial in treating MS [204]. Importantly, ORY-2001 showed excellent safety and was proven to be able to penetrate the central nervous system [198].
GSK-J1/4, developed by Kruidenier et al. [135] in 2012, selectively inhibits KDM6 family members and is the most widely used inhibitor for JmjC demethylases. GSK-J1/4 showed anti-inflammatory effects in most in vivo and in vitro experiments, which was mainly achieved by inducing immune tolerance of NK cells and DCs. GSK-J4 was shown to suppress IFN-γ production by NK cells, thus reducing inflammatory injury of RA patients and impairing NK cell induced formation of osteoclasts and joint erosion [109]. GSK-J4 also reduced the inflammation of synovial fibroblasts and attenuated joint damage in CIA mice [191]. DCs produce cytokines and regulate the function of macrophages and Th cells to indirectly induce inflammatory response as well. However, GSK-J1/4 treatment transformed pro-inflammatory DCs into tolerogenic DCs that restrained inflammation and reduced the expression of the costimulatory molecules CD80/CD86 and pro-inflammatory IL-6, IFN-γ and TNF [134]. Additionally, tolerogenic DCs suppressed the formation of M1 macrophages and pro-inflammatory Th cells (Th1 and Th17), supporting the development of Treg cells [205]. In MS, injection of GSK-J4-treated DC cells into EAE mice reduced CD4+ T cell infiltration in the central nervous system and improved inflammatory responses [134]. Likewise, GSK-J4 promoted Treg differentiation and IL-10 secretion, which attenuated inflammatory responses and reduced IL-6 and IL-17 production in murine colitis [206]. Similarly, GSK-J1 treatment decreased the proportion of Th17 cells in mouse colon tissue and pro-inflammatory cytokines IL-17A, IL-22, IL-21, and transcription factors RORγt and STAT3 secreted by Th17 cells were suppressed, while Treg cell-related IL-10, TGF-β, and FOFP3 expression were increased [205]. Additionally, GSK-J4 could reduce the inflammatory response in diseases including osteoarthritis, asthma, and mastitis in animal models [152, 180, 207]. The clinical value of other JmjC family inhibitors such as the KDM4/5 family inhibitor JIB-04 and the KDM4C inhibitor SD70 warrants further exploration [208, 209].
Conclusions and perspectives
In complex and elaborate immune responses, epigenetic modulation of KDMs is an critical for maintaining inflammatory response progression. Different KDM family proteins fine-tune the switch of gene expression by manipulating activatory or inhibitory histone methylation markers, thus participating in various links of immune cells and inflammatory activity. During the initial phase of the immune response, KDMs regulate the activation of innate immune cells, such as macrophages, DCs and NK cells, and activate inflammation by regulating PRR and downstream pathways, or directly regulating the transcription of inflammatory factors at gene promoters. Furthermore, KDMs can induce the differentiation and function of adaptive immune cells, such as T and B cells by altering intracellular gene expression patterns. Finally, during the resolution phase, KDMs can regulate immune tolerance by macrophages, DCs, and Treg cells to prevent immune overreaction.
Dysfunction of KDMs can lead to insufficient or excessive immune response, which is associated with the development of various inflammatory diseases, autoimmune diseases, and tumors. Remarkably, the regulation of KDM6A/6B is the most extensively studied. Therefore, small molecule inhibitors targeting these KDMs may provide new strategies for the treatment of inflammatory diseases. However, there are still some obstacles to their clinical application. First, the mechanism of action of KDMs in different diseases remains to be further clarified. Second, there are no clinical trials on related diseases, and the safety and efficacy of GSK-J1/4 widely used in animal experiments remain to be discussed. Finally, the interaction between histone demethylation and other epigenetic regulatory systems is not clear, and only a clearer understanding of these mechanisms can help to avoid adverse drug reactions. In conclusion, we believe that the role of KDMs in inflammation and immune response and their clinical prospect warrant further investigations, and we expect that KDMs will improve the understanding and treatment of related diseases in the future.
References
Soares MP, Teixeira L, Moita LF. Disease tolerance and immunity in host protection against infection. Nat Rev Immunol. 2017;17:83–96.
Erttmann SF, Swacha P, Aung KM, Brindefalk B, Jiang H, Härtlova A, et al. The gut microbiota prime systemic antiviral immunity via the cGAS-STING-IFN-I axis. Immunity. 2022;55:847–861.e810.
Gray JI, Farber DL. Tissue-resident immune cells in humans. Annu Rev Immunol. 2022;40:195–220.
Zhang Q, Cao X. Epigenetic regulation of the innate immune response to infection. Nat Rev Immunol. 2019;19:417–32.
Wang K, Liu H, Hu Q, Wang L, Liu J, Zheng Z, et al. Epigenetic regulation of aging: implications for interventions of aging and diseases. Signal Transduct Target Ther. 2022;7:374.
Preissl S, Gaulton KJ, Ren B. Characterizing cis-regulatory elements using single-cell epigenomics. Nat Rev Genet. 2023;24:21–43.
Chang S, Yim S, Park H. The cancer driver genes IDH1/2, JARID1C/ KDM5C, and UTX/ KDM6A: crosstalk between histone demethylation and hypoxic reprogramming in cancer metabolism. Exp Mol Med. 2019;51:1–17.
Wu R, Li S, Hudlikar R, Wang L, Shannar A, Peter R, et al. Redox signaling, mitochondrial metabolism, epigenetics and redox active phytochemicals. Free Radic Biol Med. 2022;179:328–36.
Yang J, Hu Y, Zhang B, Liang X, Li X. The JMJD family histone demethylases in crosstalk between inflammation and cancer. Front Immunol. 2022;13:881396.
Ohkura N, Sakaguchi S. Transcriptional and epigenetic basis of Treg cell development and function: its genetic anomalies or variations in autoimmune diseases. Cell Res. 2020;30:465–74.
Wang X, Sun Y, Fu Y, Wu H, Chen Y, Ye Y, et al. Lysine specific demethylase 1 inhibitor alleviated lipopolysaccharide/D-galactosamine-induced acute liver injury. Eur J Pharm. 2022;932:175227.
Chen X, Lin X, Pang G, Deng J, Xie Q, Zhang Z. Significance of KDM6A mutation in bladder cancer immune escape. BMC Cancer. 2021;21:635.
Bilmez Y, Talibova G, Ozturk S. Dynamic changes of histone methylation in mammalian oocytes and early embryos. Histochem Cell Biol. 2022;157:7–25.
Zhang J, Fan X, Zhou Y, Chen L, Rao H. The PRMT5-LSD1 axis confers Slug dual transcriptional activities and promotes breast cancer progression. J Exp Clin Cancer Res. 2022;41:191.
Su Y, Wang X, Guo Z, Wang J. Aberrant JmjC domain-containing protein 8 (JMJD8) expression promotes activation of AKT and tumor epithelial-mesenchymal transition. Oncogene. 2020;39:6451–67.
Xiao RQ, Ran T, Huang QX, Hu GS, Fan DM, Yi J, et al. A specific JMJD6 inhibitor potently suppresses multiple types of cancers both in vitro and in vivo. Proc Natl Acad Sci USA. 2022;119:e2200753119.
Manni W, Jianxin X, Weiqi H, Siyuan C, Huashan S. JMJD family proteins in cancer and inflammation. Signal Transduct Target Ther. 2022;7:304.
Song Y, Zhang H, Yang X, Shi Y, Yu B. Annual review of lysine-specific demethylase 1 (LSD1/KDM1A) inhibitors in 2021. Eur J Med Chem. 2022;228:114042.
Zhang X, Wang X, Wu T, Yin W, Yan J, Sun Y, et al. Therapeutic potential of targeting LSD1/ KDM1A in cancers. Pharm Res. 2022;175:105958.
Zheng YC, Ma J, Wang Z, Li J, Jiang B, Zhou W, et al. A systematic review of histone lysine-specific demethylase 1 and its inhibitors. Med Res Rev. 2015;35:1032–71.
Liu X, Zhang Z, She N, Zhai J, Zhao Y, Wang C. Combination of multiple methods and views for recognition, transportation, and structure-guided modification of lysine-specific demethylase phenylcyclopropylamine inhibitor. Phys Chem Chem Phys. 2022;24:13806–23.
Maes T, Mascaró C, Ortega A, Lunardi S, Ciceri F, Somervaille TC, et al. KDM1 histone lysine demethylases as targets for treatments of oncological and neurodegenerative disease. Epigenomics. 2015;7:609–26.
Tsukada Y, Fang J, Erdjument-Bromage H, Warren ME, Borchers CH, Tempst P, et al. Histone demethylation by a family of JmjC domain-containing proteins. Nature. 2006;439:811–6.
Kooistra SM, Helin K. Molecular mechanisms and potential functions of histone demethylases. Nat Rev Mol Cell Biol. 2012;13:297–311.
Horton JR, Engstrom A, Zoeller EL, Liu X, Shanks JR, Zhang X, et al. Characterization of a Linked Jumonji Domain of the KDM5/JARID1 Family of Histone H3 Lysine 4 Demethylases. J Biol Chem. 2016;291:2631–46.
Chen L, Zhang J, Zou Y, Wang F, Li J, Sun F, et al. Kdm2a deficiency in macrophages enhances thermogenesis to protect mice against HFD-induced obesity by enhancing H3K36me2 at the Pparg locus. Cell Death Differ. 2021;28:1880–99.
Lu B, Wei J, Zhou H, Chen J, Li Y, Ye L, et al. Histone H3K36me2 demethylase KDM2A promotes bladder cancer progression through epigenetically silencing RARRES3. Cell Death Dis. 2022;13:547.
Blackledge NP, Zhou JC, Tolstorukov MY, Farcas AM, Park PJ, Klose RJ. CpG islands recruit a histone H3 lysine 36 demethylase. Mol Cell. 2010;38:179–90.
Liu X, Secombe J. The histone demethylase KDM5 activates gene expression by recognizing chromatin context through its PHD reader motif. Cell Rep. 2015;13:2219–31.
Ohguchi Y, Ohguchi H. Diverse functions of KDM5 in cancer: transcriptional repressor or activator? Cancers. 2022;14:3270.
Tsukada Y, Ishitani T, Nakayama KI. KDM7 is a dual demethylase for histone H3 Lys 9 and Lys 27 and functions in brain development. Genes Dev. 2010;24:432–7.
Feng W, Yonezawa M, Ye J, Jenuwein T, Grummt I. PHF8 activates transcription of rRNA genes through H3K4me3 binding and H3K9me1/2 demethylation. Nat Struct Mol Biol. 2010;17:445–50.
Cloos PA, Christensen J, Agger K, Helin K. Erasing the methyl mark: histone demethylases at the center of cellular differentiation and disease. Genes Dev. 2008;22:1115–40.
Su Z, Wang F, Lee JH, Stephens KE, Papazyan R, Voronina E, et al. Reader domain specificity and lysine demethylase-4 family function. Nat Commun. 2016;7:13387.
Hillringhaus L, Yue WW, Rose NR, Ng SS, Gileadi C, Loenarz C, et al. Structural and evolutionary basis for the dual substrate selectivity of human KDM4 histone demethylase family. J Biol Chem. 2011;286:41616–25.
Akar-Ghibril N. Defects of the innate immune system and related immune deficiencies. Clin Rev Allergy Immunol. 2022;63:36–54.
Kraus RF, Gruber MA. Neutrophils-from bone marrow to first-line defense of the innate immune system. Front Immunol. 2021;12:767175.
Borriello F, Poli V, Shrock E, Spreafico R, Liu X, Pishesha N, et al. An adjuvant strategy enabled by modulation of the physical properties of microbial ligands expands antigen immunogenicity. Cell. 2022;185:614–629.e621.
Christofides A, Strauss L, Yeo A, Cao C, Charest A, Boussiotis VA. The complex role of tumor-infiltrating macrophages. Nat Immunol. 2022;23:1148–56.
Chou WC, Rampanelli E, Li X, Ting JP. Impact of intracellular innate immune receptors on immunometabolism. Cell Mol Immunol. 2022;19:337–51.
Liu Q, Zhu F, Liu X, Lu Y, Yao K, Tian N, et al. Non-oxidative pentose phosphate pathway controls regulatory T cell function by integrating metabolism and epigenetics. Nat Metab. 2022;4:559–74.
Roy R, Ramamoorthy S, Shapiro BD, Kaileh M, Hernandez D, Sarantopoulou D, et al. DNA methylation signatures reveal that distinct combinations of transcription factors specify human immune cell epigenetic identity. Immunity. 2021;54:2465–2480.e2465.
Zhu Y, van Essen D, Saccani S. Cell-type-specific control of enhancer activity by H3K9 trimethylation. Mol Cell. 2012;46:408–23.
Zhang H, Hu Y, Liu D, Liu Z, Xie N, Liu S, et al. The histone demethylase Kdm6b regulates the maturation and cytotoxicity of TCRαβ(+)CD8αα(+) intestinal intraepithelial lymphocytes. Cell Death Differ. 2022;29:1349–63.
Qu LH, Fang Q, Yin T, Yi HM, Mei GB, Hong ZZ, et al. Comprehensive analyses of prognostic biomarkers and immune infiltrates among histone lysine demethylases (KDMs) in hepatocellular carcinoma. Cancer Immunol Immunother. 2022;71:2449–67.
Musella M, Guarracino A, Manduca N, Galassi C, Ruggiero E, Potenza A, et al. Type I IFNs promote cancer cell stemness by triggering the epigenetic regulator KDM1B. Nat Immunol. 2022;23:1379–92.
Ford BR, Vignali PDA, Rittenhouse NL, Scharping NE, Peralta R, Lontos K, et al. Tumor microenvironmental signals reshape chromatin landscapes to limit the functional potential of exhausted T cells. Sci Immunol. 2022;7:eabj9123.
Liu H, Lin J, Zhou W, Moses R, Dai Z, Kossenkov AV, et al. KDM5A inhibits antitumor immune responses through downregulation of the antigen-presentation pathway in ovarian cancer. Cancer Immunol Res. 2022;10:1028–38.
Chapman NM, Chi H. Metabolic adaptation of lymphocytes in immunity and disease. Immunity. 2022;55:14–30.
Belk JA, Daniel B, Satpathy AT. Epigenetic regulation of T cell exhaustion. Nat Immunol. 2022;23:848–60.
Carrasco E, Gómez de Las Heras MM, Gabandé-Rodríguez E, Desdín-Micó G, Aranda JF, Mittelbrunn M. The role of T cells in age-related diseases. Nat Rev Immunol. 2022;22:97–111.
Khan U, Ghazanfar HT. Lymphocytes and autoimmunity. Int Rev Cell Mol Biol. 2018;341:125–68.
James CA, Xu Y, Aguilar MS, Jing L, Layton ED, Gilleron M, et al. CD4 and CD8 co-receptors modulate functional avidity of CD1b-restricted T cells. Nat Commun. 2022;13:78.
Körte C, Burchardt P. Involvement of the epididymis in secondary syphilis. Hautarzt. 1969;20:369–70.
Stamos DB, Clubb LM, Mitra A, Chopp LB, Nie J, Ding Y, et al. The histone demethylase Lsd1 regulates multiple repressive gene programs during T cell development. J Exp Med. 2021;218:e20202012.
Manna S, Kim JK, Baugé C, Cam M, Zhao Y, Shetty J, et al. Histone H3 Lysine 27 demethylases Jmjd3 and Utx are required for T-cell differentiation. Nat Commun. 2015;6:8152.
Li Q, Zou J, Wang M, Ding X, Chepelev I, Zhou X, et al. Critical role of histone demethylase Jmjd3 in the regulation of CD4+ T-cell differentiation. Nat Commun. 2014;5:5780.
Jie X, Chen Y, Zhao Y, Yang X, Xu Y, Wang J, et al. Targeting KDM4C enhances CD8(+) T cell mediated antitumor immunity by activating chemokine CXCL10 transcription in lung cancer. J Immunother Cancer. 2022;10:e003716.
Wei G, Wei L, Zhu J, Zang C, Hu-Li J, Yao Z, et al. Global mapping of H3K4me3 and H3K27me3 reveals specificity and plasticity in lineage fate determination of differentiating CD4+ T cells. Immunity. 2009;30:155–67.
Suzuki J, Maruyama S, Tamauchi H, Kuwahara M, Horiuchi M, Mizuki M, et al. Gfi1, a transcriptional repressor, inhibits the induction of the T helper type 1 programme in activated CD4 T cells. Immunology. 2016;147:476–87.
Liu W, Fan JB, Xu DW, Zhu XH, Yi H, Cui SY, et al. Knockdown of LSD1 ameliorates the severity of rheumatoid arthritis and decreases the function of CD4 T cells in mouse models. Int J Clin Exp Pathol. 2018;11:333–41.
Doss P, Umair M, Baillargeon J, Fazazi R, Fudge N, Akbar I, et al. Male sex chromosomal complement exacerbates the pathogenicity of Th17 cells in a chronic model of central nervous system autoimmunity. Cell Rep. 2021;34:108833.
Yang X, Bam M, Becker W, Nagarkatti PS, Nagarkatti M. Long noncoding RNA AW112010 promotes the differentiation of inflammatory T cells by suppressing IL-10 expression through histone demethylation. J Immunol. 2020;205:987–93.
Ptaschinski C, Mukherjee S, Moore ML, Albert M, Helin K, Kunkel SL, et al. RSV-induced H3K4 demethylase KDM5B leads to regulation of dendritic cell-derived innate cytokines and exacerbates pathogenesis in vivo. PLoS Pathog. 2015;11:e1004978.
Itoh Y, Golden LC, Itoh N, Matsukawa MA, Ren E, Tse V, et al. The X-linked histone demethylase Kdm6a in CD4+ T lymphocytes modulates autoimmunity. J Clin Investig. 2019;129:3852–63.
Liu Z, Cao W, Xu L, Chen X, Zhan Y, Yang Q, et al. The histone H3 lysine-27 demethylase Jmjd3 plays a critical role in specific regulation of Th17 cell differentiation. J Mol Cell Biol. 2015;7:505–16.
Cribbs AP, Terlecki-Zaniewicz S, Philpott M, Baardman J, Ahern D, Lindow M, et al. Histone H3K27me3 demethylases regulate human Th17 cell development and effector functions by impacting on metabolism. Proc Natl Acad Sci USA. 2020;117:6056–66.
Browning LM, Miller C, Kuczma M, Pietrzak M, Jing Y, Rempala G, et al. Bone morphogenic proteins are immunoregulatory cytokines controlling FOXP3(+) T(reg) cells. Cell Rep. 2020;33:108219.
Osman IO, Melenotte C, Brouqui P, Million M, Lagier JC, Parola P, et al. Expression of ACE2, soluble ACE2, angiotensin I, angiotensin II and angiotensin-(1-7) is modulated in COVID-19 patients. Front Immunol. 2021;12:625732.
Xu T, Schutte A, Jimenez L, Gonçalves ANA, Keller A, Pipkin ME, et al. Kdm6b regulates the generation of effector CD8(+) T cells by inducing chromatin accessibility in effector-associated genes. J Immunol. 2021;206:2170–83.
Yamada T, Nabe S, Toriyama K, Suzuki J, Inoue K, Imai Y, et al. Histone H3K27 demethylase negatively controls the memory formation of antigen-stimulated CD8(+) T cells. J Immunol. 2019;202:1088–98.
Howson LJ, Li J, von Borstel A, Barugahare A, Mak JYW, Fairlie DP, et al. Mucosal-associated invariant T cell effector function is an intrinsic cell property that can be augmented by the metabolic cofactor α-ketoglutarate. J Immunol. 2021;206:1425–35.
Kong IY, Trezise S, Light A, Todorovski I, Arnau GM, Gadipally S, et al. Epigenetic modulators of B cell fate identified through coupled phenotype-transcriptome analysis. Cell Death Differ. 2022;29:2519–30.
Pratumchai I, Zak J, Huang Z, Min B, Oldstone MBA, Teijaro JR. B cell-derived IL-27 promotes control of persistent LCMV infection. Proc Natl Acad Sci USA. 2022;119:e2116741119.
Xiao F, Rui K, Shi X, Wu H, Cai X, Lui KO, et al. Epigenetic regulation of B cells and its role in autoimmune pathogenesis. Cell Mol Immunol. 2022;19:1215–34.
Chen HY, Almonte-Loya A, Lay FY, Hsu M, Johnson E, González-Avalos E, et al. Epigenetic remodeling by vitamin C potentiates plasma cell differentiation. Elife. 2022;11:e73754.
Haines RR, Barwick BG, Scharer CD, Majumder P, Randall TD, Boss JM. The histone demethylase LSD1 regulates B cell proliferation and plasmablast differentiation. J Immunol. 2018;201:2799–811.
Hung KH, Woo YH, Lin IY, Liu CH, Wang LC, Chen HY, et al. The KDM4A/KDM4C/NF-κB and WDR5 epigenetic cascade regulates the activation of B cells. Nucleic Acids Res. 2018;46:5547–60.
Cook KD, Shpargel KB, Starmer J, Whitfield-Larry F, Conley B, Allard DE, et al. T follicular helper cell-dependent clearance of a persistent virus infection requires T cell expression of the histone demethylase UTX. Immunity. 2015;43:703–14.
Haniuda K, Fukao S, Kitamura D. Metabolic reprogramming induces germinal center B cell differentiation through bcl6 locus remodeling. Cell Rep. 2020;33:108333.
Hatzi K, Geng H, Doane AS, Meydan C, LaRiviere R, Cardenas M, et al. Histone demethylase LSD1 is required for germinal center formation and BCL6-driven lymphomagenesis. Nat Immunol. 2019;20:86–96.
Anderton JA, Bose S, Vockerodt M, Vrzalikova K, Wei W, Kuo M, et al. The H3K27me3 demethylase, KDM6B, is induced by Epstein-Barr virus and over-expressed in Hodgkin’s Lymphoma. Oncogene. 2011;30:2037–43.
Paterson N, Lämmermann T. Macrophage network dynamics depend on haptokinesis for optimal local surveillance. Elife. 2022;11:e75354.
Mantovani A, Allavena P, Marchesi F, Garlanda C. Macrophages as tools and targets in cancer therapy. Nat Rev Drug Discov. 2022;21:799–820.
Aegerter H, Lambrecht BN, Jakubzick CV. Biology of lung macrophages in health and disease. Immunity. 2022;55:1564–80.
Hirao H, Nakamura K, Kupiec-Weglinski JW. Liver ischaemia-reperfusion injury: a new understanding of the role of innate immunity. Nat Rev Gastroenterol Hepatol. 2022;19:239–56.
Murray PJ. Macrophage polarization. Annu Rev Physiol. 2017;79:541–66.
Pan K, Farrukh H, Chittepu V, Xu H, Pan CX, Zhu Z. CAR race to cancer immunotherapy: from CAR T, CAR NK to CAR macrophage therapy. J Exp Clin Cancer Res. 2022;41:119.
Ge G, Bai J, Wang Q, Liang X, Tao H, Chen H, et al. Punicalagin ameliorates collagen-induced arthritis by downregulating M1 macrophage and pyroptosis via NF-κB signaling pathway. Sci China Life Sci. 2022;65:588–603.
Zhang L, Chen X, Cai P, Sun H, Shen S, Guo B, et al. Reprogramming mitochondrial metabolism in synovial macrophages of early osteoarthritis by a camouflaged meta-defensome. Adv Mater. 2022;34:e2202715.
Zhong C, Tao B, Yang F, Xia K, Yang X, Chen L, et al. Histone demethylase JMJD1C promotes the polarization of M1 macrophages to prevent glioma by upregulating miR-302a. Clin Transl Med. 2021;11:e424.
Ishii M, Wen H, Corsa CA, Liu T, Coelho AL, Allen RM, et al. Epigenetic regulation of the alternatively activated macrophage phenotype. Blood. 2009;114:3244–54.
Satoh T, Takeuchi O, Vandenbon A, Yasuda K, Tanaka Y, Kumagai Y, et al. The Jmjd3-Irf4 axis regulates M2 macrophage polarization and host responses against helminth infection. Nat Immunol. 2010;11:936–44.
Liu PS, Wang H, Li X, Chao T, Teav T, Christen S, et al. α-ketoglutarate orchestrates macrophage activation through metabolic and epigenetic reprogramming. Nat Immunol. 2017;18:985–94.
Ming-Chin Lee K, Achuthan AA, De Souza DP, Lupancu TJ, Binger KJ, Lee MKS, et al. Type I interferon antagonism of the JMJD3-IRF4 pathway modulates macrophage activation and polarization. Cell Rep. 2022;39:110719.
De Santa F, Totaro MG, Prosperini E, Notarbartolo S, Testa G, Natoli G. The histone H3 lysine-27 demethylase Jmjd3 links inflammation to inhibition of polycomb-mediated gene silencing. Cell. 2007;130:1083–94.
Huang M, Wang Q, Long F, Di Y, Wang J, Zhun Zhu Y, et al. Jmjd3 regulates inflammasome activation and aggravates DSS-induced colitis in mice. FASEB J. 2020;34:4107–19.
Zhuo X, Wu Y, Yang Y, Gao L, Qiao X, Chen T. Knockdown of LSD1 meliorates Ox-LDL-stimulated NLRP3 activation and inflammation by promoting autophagy via SESN2-mesiated PI3K/Akt/mTOR signaling pathway. Life Sci. 2019;233:116696.
Parmar N, Chandrakar P, Kar S. Leishmania donovani subverts host immune response by epigenetic reprogramming of macrophage M(Lipopolysaccharides + IFN-γ)/M(IL-10) polarization. J Immunol. 2020;204:2762–78.
Lau CM, Wiedemann GM, Sun JC. Epigenetic regulation of natural killer cell memory. Immunol Rev. 2022;305:90–110.
Westhoff Smith D, Chakravorty A, Hayes M, Hammerschmidt W, Sugden B. The Epstein-Barr virus oncogene EBNA1 suppresses natural killer cell responses and apoptosis early after infection of peripheral B cells. mBio. 2021;12:e0224321.
Laskowski TJ, Biederstädt A, Rezvani K. Natural killer cells in antitumour adoptive cell immunotherapy. Nat Rev Cancer. 2022;22:557–75.
Tong L, Jiménez-Cortegana C, Tay AHM, Wickström S, Galluzzi L, Lundqvist A. NK cells and solid tumors: therapeutic potential and persisting obstacles. Mol Cancer. 2022;21:206.
Sivori S, Vacca P, Del Zotto G, Munari E, Mingari MC, Moretta L. Human NK cells: surface receptors, inhibitory checkpoints, and translational applications. Cell Mol Immunol. 2019;16:430–41.
Feehan DD, Jamil K, Polyak MJ, Ogbomo H, Hasell M, Li SS, et al. Natural killer cells kill extracellular Pseudomonas aeruginosa using contact-dependent release of granzymes B and H. PLoS Pathog. 2022;18:e1010325.
Jewett A, Kos J, Fong Y, Ko MW, Safaei T, Perišić Nanut M, et al. NK cells shape pancreatic and oral tumor microenvironments; role in inhibition of tumor growth and metastasis. Semin Cancer Biol. 2018;53:178–88.
Bailey CP, Figueroa M, Gangadharan A, Lee DA, Chandra J. Scaffolding LSD1 inhibitors impair NK cell metabolism and cytotoxic function through depletion of glutathione. Front Immunol. 2020;11:2196.
Zhao D, Zhang Q, Liu Y, Li X, Zhao K, Ding Y, et al. H3K4me3 demethylase Kdm5a is required for NK cell activation by associating with p50 to suppress SOCS1. Cell Rep. 2016;15:288–99.
Cribbs A, Hookway ES, Wells G, Lindow M, Obad S, Oerum H, et al. Inhibition of histone H3K27 demethylases selectively modulates inflammatory phenotypes of natural killer cells. J Biol Chem. 2018;293:2422–37.
Bendelac A, Savage PB, Teyton L. The biology of NKT cells. Annu Rev Immunol. 2007;25:297–336.
Rossjohn J, Pellicci DG, Patel O, Gapin L, Godfrey DI. Recognition of CD1d-restricted antigens by natural killer T cells. Nat Rev Immunol. 2012;12:845–57.
Dobenecker MW, Kim JK, Marcello J, Fang TC, Prinjha R, Bosselut R, et al. Coupling of T cell receptor specificity to natural killer T cell development by bivalent histone H3 methylation. J Exp Med. 2015;212:297–306.
Beyaz S, Kim JH, Pinello L, Xifaras ME, Hu Y, Huang J, et al. The histone demethylase UTX regulates the lineage-specific epigenetic program of invariant natural killer T cells. Nat Immunol. 2017;18:184–95.
Northrup D, Yagi R, Cui K, Proctor WR, Wang C, Placek K, et al. Histone demethylases UTX and JMJD3 are required for NKT cell development in mice. Cell Biosci. 2017;7:25.
Kim CU, Jeong YJ, Lee P, Lee MS, Park JH, Kim YS, et al. Extracellular nucleoprotein exacerbates influenza virus pathogenesis by activating Toll-like receptor 4 and the NLRP3 inflammasome. Cell Mol Immunol. 2022;19:715–25.
Takeuchi O, Akira S. Pattern recognition receptors and inflammation. Cell. 2010;140:805–20.
Hasan UA, Zannetti C, Parroche P, Goutagny N, Malfroy M, Roblot G, et al. The human papillomavirus type 16 E7 oncoprotein induces a transcriptional repressor complex on the Toll-like receptor 9 promoter. J Exp Med. 2013;210:1369–87.
Wu L, Cao J, Cai WL, Lang SM, Horton JR, Jansen DJ, et al. KDM5 histone demethylases repress immune response via suppression of STING. PLoS Biol. 2018;16:e2006134.
Wang L, Gao Y, Zhang G, Li D, Wang Z, Zhang J, et al. Enhancing KDM5A and TLR activity improves the response to immune checkpoint blockade. Sci Transl Med. 2020;12:eaax2282.
Dong F, Jiang S, Li J, Wang Y, Zhu L, Huang Y, et al. The histone demethylase KDM4D promotes hepatic fibrogenesis by modulating Toll-like receptor 4 signaling pathway. EBioMedicine. 2019;39:472–83.
Jo YH, Lee JH, Patnaik BB, Keshavarz M, Lee YS, Han YS. Autophagy in tenebrio molitor immunity: conserved antimicrobial functions in insect defenses. Front Immunol. 2021;12:667664.
Katakia YT, Thakkar NP, Thakar S, Sakhuja A, Goyal R, Sharma H, et al. Dynamic alterations of H3K4me3 and H3K27me3 at ADAM17 and Jagged-1 gene promoters cause an inflammatory switch of endothelial cells. J Cell Physiol. 2022;237:992–1012.
Asensio-Juan E, Fueyo R, Pappa S, Iacobucci S, Badosa C, Lois S, et al. The histone demethylase PHF8 is a molecular safeguard of the IFNγ response. Nucleic Acids Res. 2017;45:3800–11.
Kim D, Nam HJ, Lee W, Yim HY, Ahn JY, Park SW, et al. PKCα-LSD1-NF-κB-signaling cascade is crucial for epigenetic control of the inflammatory response. Mol Cell. 2018;69:398–411.e396.
Lu T, Jackson MW, Wang B, Yang M, Chance MR, Miyagi M, et al. Regulation of NF-kappaB by NSD1/FBXL11-dependent reversible lysine methylation of p65. Proc Natl Acad Sci USA. 2010;107:46–51.
Zhao Z, Su Z, Liang P, Liu D, Yang S, Wu Y, et al. USP38 couples histone ubiquitination and methylation via KDM5B to resolve inflammation. Adv Sci. 2020;7:2002680.
Connor MG, Camarasa TMN, Patey E, Rasid O, Barrio L, Weight CM, et al. The histone demethylase KDM6B fine-tunes the host response to Streptococcus pneumoniae. Nat Microbiol. 2021;6:257–69.
Higashijima Y, Matsui Y, Shimamura T, Nakaki R, Nagai N, Tsutsumi S, et al. Coordinated demethylation of H3K9 and H3K27 is required for rapid inflammatory responses of endothelial cells. Embo J. 2020;39:e103949.
Stender JD, Pascual G, Liu W, Kaikkonen MU, Do K, Spann NJ, et al. Control of proinflammatory gene programs by regulated trimethylation and demethylation of histone H4K20. Mol Cell. 2012;48:28–38.
Jin J, Hu H, Li HS, Yu J, Xiao Y, Brittain GC, et al. Noncanonical NF-κB pathway controls the production of type I interferons in antiviral innate immunity. Immunity. 2014;40:342–54.
Zhou Q, Zhang Y, Wang B, Zhou W, Bi Y, Huai W, et al. KDM2B promotes IL-6 production and inflammatory responses through Brg1-mediated chromatin remodeling. Cell Mol Immunol. 2020;17:834–42.
Lee K, Na W, Lee JY, Na J, Cho H, Wu H, et al. Molecular mechanism of Jmjd3-mediated interleukin-6 gene regulation in endothelial cells underlying spinal cord injury. J Neurochem. 2012;122:272–82.
Li X, Zhang Q, Shi Q, Liu Y, Zhao K, Shen Q, et al. Demethylase Kdm6a epigenetically promotes IL-6 and IFN-β production in macrophages. J Autoimmun. 2017;80:85–94.
Doñas C, Carrasco M, Fritz M, Prado C, Tejón G, Osorio-Barrios F, et al. The histone demethylase inhibitor GSK-J4 limits inflammation through the induction of a tolerogenic phenotype on DCs. J Autoimmun. 2016;75:105–17.
Kruidenier L, Chung CW, Cheng Z, Liddle J, Che K, Joberty G, et al. A selective jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response. Nature. 2012;488:404–8.
Malinczak CA, Rasky AJ, Fonseca W, Schaller MA, Allen RM, Ptaschinski C, et al. Upregulation of H3K27 demethylase KDM6 during respiratory syncytial virus infection enhances proinflammatory responses and immunopathology. J Immunol. 2020;204:159–68.
Sun D, Cao X, Wang C. Polycomb chromobox Cbx2 enhances antiviral innate immunity by promoting Jmjd3-mediated demethylation of H3K27 at the Ifnb promoter. Protein Cell. 2019;10:285–94.
Wang J, Saijo K, Skola D, Jin C, Ma Q, Merkurjev D, et al. Histone demethylase LSD1 regulates hematopoietic stem cells homeostasis and protects from death by endotoxic shock. Proc Natl Acad Sci USA. 2018;115:E244–e252.
Boukhaled GM, Cordeiro B, Deblois G, Dimitrov V, Bailey SD, Holowka T, et al. The transcriptional repressor polycomb group factor 6, PCGF6, negatively regulates dendritic cell activation and promotes quiescence. Cell Rep. 2016;16:1829–37.
Janzer A, Lim S, Fronhoffs F, Niazy N, Buettner R, Kirfel J. Lysine-specific demethylase 1 (LSD1) and histone deacetylase 1 (HDAC1) synergistically repress proinflammatory cytokines and classical complement pathway components. Biochem Biophys Res Commun. 2012;421:665–70.
Yu X, Chen H, Zuo C, Jin X, Yin Y, Wang H, et al. Chromatin remodeling: demethylating H3K4me3 of type I IFNs gene by Rbp2 through interacting with Piasy for transcriptional attenuation. Faseb J. 2018;32:552–67.
Salminen A, Kaarniranta K, Hiltunen M, Kauppinen A. Histone demethylase Jumonji D3 (JMJD3/KDM6B) at the nexus of epigenetic regulation of inflammation and the aging process. J Mol Med. 2014;92:1035–43.
Liu D, Wang Y, Jia Z, Wang L, Wang J, Yang D, et al. Demethylation of IGFBP5 by histone demethylase KDM6B promotes mesenchymal stem cell-mediated periodontal tissue regeneration by enhancing osteogenic differentiation and anti-inflammation potentials. Stem Cells. 2015;33:2523–36.
Hung PH, Hsu YC, Chen TH, Ho C, Lin CL. The histone demethylase inhibitor GSK-J4 is a therapeutic target for the kidney fibrosis of diabetic kidney disease via DKK1 modulation. Int J Mol Sci. 2022;23:16.
Boulding T, McCuaig RD, Tan A, Hardy K, Wu F, Dunn J, et al. Author Correction: LSD1 activation promotes inducible EMT programs and modulates the tumour microenvironment in breast cancer. Sci Rep. 2019;9:18771.
Pan HC, Chen YH, Fang WC, Wu VC, Sun CY. Essential roles of the histone demethylase KDM4C in renal development and acute kidney injury. Int J Mol Sci. 2022;23:16.
Yan Q, Sun L, Zhu Z, Wang L, Li S, Ye RD. Jmjd3-mediated epigenetic regulation of inflammatory cytokine gene expression in serum amyloid A-stimulated macrophages. Cell Signal. 2014;26:1783–91.
Li Y, Reddy MA, Miao F, Shanmugam N, Yee JK, Hawkins D, et al. Role of the histone H3 lysine 4 methyltransferase, SET7/9, in the regulation of NF-kappaB-dependent inflammatory genes. Relevance to diabetes and inflammation. J Biol Chem. 2008;283:26771–81.
Valls-Bellés V, Abad C, Hernández-Aguilar MT, Nacher A, Guerrero C, Baliño P, et al. Human milk antioxidative modifications in mastitis: further beneficial effects of cranberry supplementation. Antioxidants. 2021;11:51.
Morales-Ferré C, Franch À, Castell M, Olivares M, Rodríguez-Lagunas MJ, Pérez-Cano FJ. Staphylococcus epidermidis’ overload during suckling impacts the immune development in rats. Front Nutr. 2022;9:916690.
Khan MZ, Khan A, Xiao J, Ma J, Ma Y, Chen T, et al. Overview of research development on the role of NF-κB signaling in mastitis. Animimal. 2020;10:1625.
Wang JJ, Wang X, Xian YE, Chen ZQ, Sun YP, Fu YW, et al. The JMJD3 histone demethylase inhibitor GSK-J1 ameliorates lipopolysaccharide-induced inflammation in a mastitis model. J Biol Chem. 2022;298:102017.
Jingjing W, Zhikai W, Xingyi Z, Peixuan L, Yiwu F, Xia W, et al. Lysine-specific demethylase 1 (LSD1) serves as an potential epigenetic determinant to regulate inflammatory responses in mastitis. Int Immunopharmacol. 2021;91:107324.
Higashi T, Friedman SL, Hoshida Y. Hepatic stellate cells as key target in liver fibrosis. Adv Drug Deliv Rev. 2017;121:27–42.
Kisseleva T, Brenner D. Molecular and cellular mechanisms of liver fibrosis and its regression. Nat Rev Gastroenterol Hepatol. 2021;18:151–66.
Fondevila MF, Fernandez U, Heras V, Parracho T, Gonzalez-Rellan MJ, Novoa E, et al. Inhibition of carnitine palmitoyltransferase 1A in hepatic stellate cells protects against fibrosis. J Hepatol. 2022;77:15–28.
Luo P, Liu D, Zhang Q, Yang F, Wong YK, Xia F, et al. Celastrol induces ferroptosis in activated HSCs to ameliorate hepatic fibrosis via targeting peroxiredoxins and HO-1. Acta Pharm Sin B. 2022;12:2300–14.
Zhang WS, Zhang R, Ge Y, Wang D, Hu Y, Qin X, et al. S100a16 deficiency prevents hepatic stellate cells activation and liver fibrosis via inhibiting CXCR4 expression. Metabolism. 2022;135:155271.
Li Y, Kang X, Zhou Z, Pan L, Chen H, Liang X, et al. The m(6)A methyltransferase Mettl3 deficiency attenuates hepatic stellate cell activation and liver fibrosis. Mol Ther. 2022;30:3714–28.
Tian T, Xie R, Ding K, Han B, Yang Q, Yang X. IOX1 protects from TGF-β induced fibrosis in LX-2 cells via the regulation of extracellular matrix protein expression. Exp Ther Med. 2021;21:180.
Wang Q, Wang LX, Zeng JP, Liu XJ, Liang XM, Zhou YB. Histone demethylase retinoblastoma binding protein 2 regulates the expression of α-smooth muscle actin and vimentin in cirrhotic livers. Braz J Med Biol Res. 2013;46:739–45.
Jiang Y, Xiang C, Zhong F, Zhang Y, Wang L, Zhao Y, et al. Histone H3K27 methyltransferase EZH2 and demethylase JMJD3 regulate hepatic stellate cells activation and liver fibrosis. Theranostics. 2021;11:361–78.
Kong M, Wu J, Fan Z, Chen B, Wu T, Xu Y. The histone demethylase Kdm4 suppresses activation of hepatic stellate cell by inducing MiR-29 transcription. Biochem Biophys Res Commun. 2019;514:16–23.
Li N, Kong M, Zeng S, Hao C, Li M, Li L, et al. Brahma related gene 1 (Brg1) contributes to liver regeneration by epigenetically activating the Wnt/β-catenin pathway in mice. FASEB J. 2019;33:327–38.
Jiang Y, Wang S, Zhao Y, Lin C, Zhong F, Jin L, et al. Histone H3K9 demethylase JMJD1A modulates hepatic stellate cells activation and liver fibrosis by epigenetically regulating peroxisome proliferator-activated receptor γ. FASEB J. 2015;29:1830–41.
Schonfeld M, Averilla J, Gunewardena S, Weinman SA, Tikhanovich I. Alcohol-associated fibrosis in females is mediated by female-specific activation of lysine demethylases KDM5B and KDM5C. Hepatol Commun. 2022;6:2042–57.
Yang Q, Ren GL, Wei B, Jin J, Huang XR, Shao W, et al. Conditional knockout of TGF-βRII /Smad2 signals protects against acute renal injury by alleviating cell necroptosis, apoptosis and inflammation. Theranostics. 2019;9:8277–93.
Mimura I, Tanaka T, Nangaku M. Novel therapeutic strategy with hypoxia-inducible factors via reversible epigenetic regulation mechanisms in progressive tubulointerstitial fibrosis. Semin Nephrol. 2013;33:375–82.
Zhang X, Li LX, Yu C, Nath KA, Zhuang S, Li X. Targeting lysine-specific demethylase 1A inhibits renal epithelial-mesenchymal transition and attenuates renal fibrosis. FASEB J. 2022;36:e22122.
Yu C, Xiong C, Tang J, Hou X, Liu N, Bayliss G, et al. Histone demethylase JMJD3 protects against renal fibrosis by suppressing TGFβ and Notch signaling and preserving PTEN expression. Theranostics. 2021;11:2706–21.
He Z, Wang H, Yue L. Endothelial progenitor cells-secreted extracellular vesicles containing microRNA-93-5p confer protection against sepsis-induced acute kidney injury via the KDM6B/H3K27me3/TNF-α axis. Exp Cell Res. 2020;395:112173.
Feng S, Peden EK, Guo Q, Lee TH, Li Q, Yuan Y, et al. Downregulation of the endothelial histone demethylase JMJD3 is associated with neointimal hyperplasia of arteriovenous fistulas in kidney failure. J Biol Chem. 2022;298:101816.
Chen H, Huang Y, Zhu X, Liu C, Yuan Y, Su H, et al. Histone demethylase UTX is a therapeutic target for diabetic kidney disease. J Physiol. 2019;597:1643–60.
Liu Y, Yu Y, Zhang J, Wang C. The therapeutic effect of dexmedetomidine on protection from renal failure via inhibiting KDM5A in lipopolysaccharide-induced sepsis of mice. Life Sci. 2019;239:116868.
Yang YT, Wang X, Zhang YY, Yuan WJ. The histone demethylase LSD1 promotes renal inflammation by mediating TLR4 signaling in hepatitis B virus-associated glomerulonephritis. Cell Death Dis. 2019;10:278.
Skevaki C, Karsonova A, Karaulov A, Xie M, Renz H. Asthma-associated risk for COVID-19 development. J Allergy Clin Immunol. 2020;146:1295–301.
Theofani E, Semitekolou M, Samitas K, Mais A, Galani IE, Triantafyllia V, et al. TFEB signaling attenuates NLRP3-driven inflammatory responses in severe asthma. Allergy. 2022;77:2131–46.
Georas SN, Wright RJ, Ivanova A, Israel E, LaVange LM, Akuthota P, et al. The Precision Interventions for Severe and/or Exacerbation-Prone (PrecISE) asthma network: an overview of Network organization, procedures, and interventions. J Allergy Clin Immunol. 2022;149:488–516.e489.
Bajbouj K, Hachim MY, Ramakrishnan RK, Fazel H, Mustafa J, Alzaghari S, et al. IL-13 augments histone demethylase JMJD2B/KDM4B expression levels, activity, and nuclear translocation in airway fibroblasts in asthma. J Immunol Res. 2021;2021:6629844.
Yu Q, Yu X, Zhao W, Zhu M, Wang Z, Zhang J, et al. Inhibition of H3K27me3 demethylases attenuates asthma by reversing the shift in airway smooth muscle phenotype. Clin Exp Allergy. 2018;48:1439–52.
Hong H, Huang Q, Cai Y, Lin T, Xia F, Jin Z. Dexmedetomidine preconditioning ameliorates lung injury induced by pulmonary ischemia/reperfusion by upregulating promoter histone H3K4me3 modification of KGF-2. Exp Cell Res. 2021;406:112762.
He C, Larson-Casey JL, Gu L, Ryan AJ, Murthy S, Carter AB. Cu,Zn-superoxide dismutase-mediated redox regulation of jumonji domain containing 3 modulates macrophage polarization and pulmonary fibrosis. Am J Respir Cell Mol Biol. 2016;55:58–71.
Fraszczak J, Arman KM, Lacroix M, Vadnais C, Gaboury L, Möröy T. Severe inflammatory reactions in mice expressing a GFI1(P2A) mutant defective in binding to the histone demethylase KDM1A (LSD1). J Immunol. 2021;207:1599–615.
Yang ZH, Dang YQ, Ji G. Role of epigenetics in transformation of inflammation into colorectal cancer. World J Gastroenterol. 2019;25:2863–77.
Ma S, Xu L, Chen L, Sun X, Hu F, Gong Y, et al. Novel pharmacological inhibition of JMJD3 improves necrotizing enterocolitis by attenuating the inflammatory response and ameliorating intestinal injury. Biochem Pharm. 2022;203:115165.
Zhuo M, Chen W, Shang S, Guo P, Peng K, Li M, et al. Inflammation-induced JMJD2D promotes colitis recovery and colon tumorigenesis by activating Hedgehog signaling. Oncogene. 2020;39:3336–53.
Sun X, Yang Q, Rogers CJ, Du M, Zhu MJ. AMPK improves gut epithelial differentiation and barrier function via regulating Cdx2 expression. Cell Death Differ. 2017;24:819–31.
Parmar N, Burrows K, Vornewald PM, Lindholm HT, Zwiggelaar RT, Díez-Sánchez A, et al. Intestinal-epithelial LSD1 controls goblet cell maturation and effector responses required for gut immunity to bacterial and helminth infection. PLoS Pathog. 2021;17:e1009476.
Szyf M. Epigenetic therapeutics in autoimmune disease. Clin Rev Allergy Immunol. 2010;39:62–77.
Jia W, Wu W, Yang D, Xiao C, Su Z, Huang Z, et al. Histone demethylase JMJD3 regulates fibroblast-like synoviocyte-mediated proliferation and joint destruction in rheumatoid arthritis. Faseb J. 2018;32:4031–42.
Wu W, Qin M, Jia W, Huang Z, Li Z, Yang D, et al. Cystathionine-γ-lyase ameliorates the histone demethylase JMJD3-mediated autoimmune response in rheumatoid arthritis. Cell Mol Immunol. 2019;16:694–705.
Zhao Z, Zhang Y, Gao D, Zhang Y, Han W, Xu X, et al. Inhibition of histone H3 lysine-27 demethylase activity relieves rheumatoid arthritis symptoms via repression of IL6 transcription in macrophages. Front Immunol. 2022;13:818070.
Zhang X, Nan H, Guo J, Liu J. KDM4B overexpression promotes the growth, migration, and invasion of rheumatoid arthritis fibroblast-like synoviocytes by activating STAT3 pathway. Biochem Genet. 2021;59:1427–40.
Yin H, Wu H, Zhao M, Zhang Q, Long H, Fu S, et al. Histone demethylase JMJD3 regulates CD11a expression through changes in histone H3K27 tri-methylation levels in CD4+ T cells of patients with systemic lupus erythematosus. Oncotarget. 2017;8:48938–47.
He X, Zhang H, Zhang Y, Ye Y, Wang S, Bai R, et al. Drug discovery of histone lysine demethylases (KDMs) inhibitors (progress from 2018 to present). Eur J Med Chem. 2022;231:114143.
Binda C, Valente S, Romanenghi M, Pilotto S, Cirilli R, Karytinos A, et al. Biochemical, structural, and biological evaluation of tranylcypromine derivatives as inhibitors of histone demethylases LSD1 and LSD2. J Am Chem Soc. 2010;132:6827–33.
Fang Y, Liao G, Yu B. LSD1/KDM1A inhibitors in clinical trials: advances and prospects. J Hematol Oncol. 2019;12:129.
Antonijoan RM, Ferrero-Cafiero JM, Coimbra J, Puntes M, Martínez-Colomer J, Arévalo MI, et al. First-in-human randomized trial to assess safety, tolerability, pharmacokinetics and pharmacodynamics of the KDM1A inhibitor vafidemstat. CNS Drugs. 2021;35:331–44.
Kanouni T, Severin C, Cho RW, Yuen NY, Xu J, Shi L, et al. Discovery of CC-90011: a potent and selective reversible inhibitor of lysine specific demethylase 1 (LSD1). J Med Chem. 2020;63:14522–9.
Hollebecque A, Salvagni S, Plummer R, Isambert N, Niccoli P, Capdevila J, et al. Phase I study of lysine-specific demethylase 1 inhibitor, CC-90011, in patients with advanced solid tumors and relapsed/refractory non-hodgkin lymphoma. Clin Cancer Res. 2021;27:438–46.
Salamero O, Montesinos P, Willekens C, Pérez-Simón JA, Pigneux A, Récher C, et al. First-in-human phase I study of Iadademstat (ORY-1001): a first-in-class lysine-specific histone demethylase 1A inhibitor, in relapsed or refractory acute myeloid leukemia. J Clin Oncol. 2020;38:4260–73.
Tremblay D, Mascarenhas J. Next generation therapeutics for the treatment of myelofibrosis. Cells. 2021;10:1034.
Gill H. Lysine-specific demethylase 1 (LSD1/KDM1A) inhibition as a target for disease modification in myelofibrosis. Cells. 2022;11:2107.
Cavalcanti F, Gonzalez-Rey E, Delgado M, Falo CP, Mestre L, Guaza C, et al. Efficacy of vafidemstat in experimental autoimmune encephalomyelitis highlights the KDM1A/RCOR1/HDAC epigenetic axis in multiple sclerosis. Pharmaceutics. 2022;14:1420.
Leng XY, Yang J, Fan H, Chen QY, Cheng BJ, He HX, et al. JMJD3/H3K27me3 epigenetic modification regulates Th17/Treg cell differentiation in ulcerative colitis. Int Immunopharmacol. 2022;110:109000.
Doñas C, Neira J, Osorio-Barrios F, Carrasco M, Fernández D, Prado C, et al. The demethylase inhibitor GSK-J4 limits inflammatory colitis by promoting de novo synthesis of retinoic acid in dendritic cells. Sci Rep. 2021;11:1342.
Jun Z, Xinmeng J, Yue L, Zhi W, Yan Z, Tieyi Y, et al. Jumonji domain containing-3 (JMJD3) inhibition attenuates IL-1β-induced chondrocytes damage in vitro and protects osteoarthritis cartilage in vivo. Inflamm Res. 2020;69:657–66.
Wang L, Chang J, Varghese D, Dellinger M, Kumar S, Best AM, et al. A small molecule modulates Jumonji histone demethylase activity and selectively inhibits cancer growth. Nat Commun. 2013;4:2035.
Jin C, Yang L, Xie M, Lin C, Merkurjev D, Yang JC, et al. Chem-seq permits identification of genomic targets of drugs against androgen receptor regulation selected by functional phenotypic screens. Proc Natl Acad Sci USA. 2014;111:9235–40.
Funding
This article was supported by the National Natural Science Foundation of China (grant no. 82172301, 81971966).
Author information
Authors and Affiliations
Contributions
HFD and XQL supervised the manuscript and edited it. LHQ, TY, and YJZ presented the concept of manuscript and wrote it. WTL, ZQL, CC, KJL, SGS, and RZ contributed to manuscript preparation and modification. All authors contributed to the manuscript and approved the final version of manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent for publication
All authors contributed to the manuscript and approved the submit the manuscript for publications.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Qu, L., Yin, T., Zhao, Y. et al. Histone demethylases in the regulation of immunity and inflammation. Cell Death Discov. 9, 188 (2023). https://doi.org/10.1038/s41420-023-01489-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-023-01489-9