Abstract
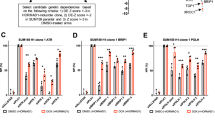
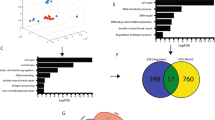
Oncogene-induced replication stress characterizes many aggressive cancers. Several treatments are being developed that target replication stress, however, identification of tumors with high levels of replication stress remains challenging. We describe a gene expression signature of oncogene-induced replication stress. A panel of triple-negative breast cancer (TNBC) and non-transformed cell lines were engineered to overexpress CDC25A, CCNE1 or MYC, which resulted in slower replication kinetics. RNA sequencing analysis revealed a set of 52 commonly upregulated genes. In parallel, mRNA expression analysis of patient-derived tumor samples (TCGA, n = 10,592) also revealed differential gene expression in tumors with amplification of oncogenes that trigger replication stress (CDC25A, CCNE1, MYC, CCND1, MYB, MOS, KRAS, ERBB2, and E2F1). Upon integration, we identified a six-gene signature of oncogene-induced replication stress (NAT10, DDX27, ZNF48, C8ORF33, MOCS3, and MPP6). Immunohistochemical analysis of NAT10 in breast cancer samples (n = 330) showed strong correlation with expression of phospho-RPA (R = 0.451, p = 1.82 × 10−20) and γH2AX (R = 0.304, p = 2.95 × 10−9). Finally, we applied our oncogene-induced replication stress signature to patient samples from TCGA (n = 8,862) and GEO (n = 13,912) to define the levels of replication stress across 27 tumor subtypes, identifying diffuse large B cell lymphoma, ovarian cancer, TNBC and colorectal carcinoma as cancer subtypes with high levels of oncogene-induced replication stress.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61–70.
Curtis C, Shah SP, Chin S-F, Turashvili G, Rueda OM, Dunning MJ, et al. The genomic and transcriptomic architecture of 2000 breast tumours reveals novel subgroups. Nature. 2012;486:346–52.
Gerlinger M, Swanton C. How Darwinian models inform therapeutic failure initiated by clonal heterogeneity in cancer medicine. Br J Cancer. 2010;103:1139–43.
Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature. 2011;474:609–15.
Zeman MK, Cimprich KA. Causes and consequences of replication stress. Nat Cell Biol. 2014;16:2–9.
Negrini S, Gorgoulis VG, Halazonetis TD. Genomic instability an evolving hallmark of cancer. Nat Rev Mol Cell Biol. 2010;11:220–8.
Bell SP, Dutta A. DNA replication in eukaryotic cells. Annu Rev Biochem. 2002;71:333–74.
Bester AC, Roniger M, Oren YS, Im MM, Sarni D, Chaoat M, et al. Nucleotide deficiency promotes genomic instability in early stages of cancer development. Cell. 2011;145:435–46.
Halazonetis TD, Gorgoulis VG, Bartek J. An oncogene-induced DNA damage model for cancer development. Science. 2008;319:1352–5.
Bartkova J, Horejsí Z, Koed K, Krämer A, Tort F, Zieger K, et al. DNA damage response as a candidate anti-cancer barrier in early human tumorigenesis. Nature. 2005;434:864–70.
Macheret M, Halazonetis TD. Intragenic origins due to short G1 phases underlie oncogene-induced DNA replication stress. Nature. 2018;555:112–6.
Neelsen KJ, Zanini IMY, Herrador R, Lopes M. Oncogenes induce genotoxic stress by mitotic processing of unusual replication intermediates. J Cell Biol. 2013;200:699–708.
Aziz K, Limzerwala JF, Sturmlechner I, Hurley E, Zhang C, Jeganathan KB, et al. Ccne1 overexpression causes chromosome instability in liver cells and liver tumor development in mice. Gastroenterology. 2019;157:210–226.e12.
López-Contreras AJ, Gutierrez-Martinez P, Specks J, Rodrigo-Perez S, Fernandez-Capetillo O. An extra allele of Chk1 limits oncogene-induced replicative stress and promotes transformation. J Exp Med. 2012;209:455–61.
Young LA, O’Connor LO, de Renty C, Veldman-Jones MH, Dorval T, Wilson Z, et al. Differential activity of ATR and WEE1 inhibitors in a highly sensitive subpopulation of DLBCL linked to replication stress. Cancer Res. 2019;79:3762–75.
Bartkova J, Rezaei N, Liontos M, Karakaidos P, Kletsas D, Issaeva N, et al. Oncogene-induced senescence is part of the tumorigenesis barrier imposed by DNA damage checkpoints. Nature. 2006;444:633–7.
Leijen S, van Geel RMJM, Sonke GS, de Jong D, Rosenberg EH, Marchetti S, et al. Phase II study of WEE1 inhibitor AZD1775 plus carboplatin in patients with TP53-mutated ovarian cancer refractory or resistant to first-line therapy within 3 months. J Clin Oncol. 2016;34:4354–61.
Chen X, Low K-H, Alexander A, Jiang Y, Karakas C, Hess KR, et al. Cyclin E overexpression sensitizes triple-negative breast cancer to Wee1 kinase inhibition. Clin Cancer Res. 2018;24:6594–610.
Kok YP, Guerrero Llobet S, Schoonen PM, Everts M, Bhattacharya A, Fehrmann RSN, et al. Overexpression of Cyclin E1 or Cdc25A leads to replication stress, mitotic aberrancies, and increased sensitivity to replication checkpoint inhibitors. Oncogenesis. 2020;9:88.
Heijink AM, Blomen VA, Bisteau X, Degener F, Matsushita FY, Kaldis P, et al. A haploid genetic screen identifies the G1/S regulatory machinery as a determinant of Wee1 inhibitor sensitivity. Proc Natl Acad Sci USA. 2015;112:15160–5.
Aarts M, Sharpe R, Garcia-Murillas I, Gevensleben H, Hurd MS, Shumway SD, et al. Forced mitotic entry of S-phase cells as a therapeutic strategy induced by inhibition of WEE1. Cancer Discov. 2012;2:524–39.
Wengner AM, Siemeister G, Lücking U, Lefranc J, Wortmann L, Lienau P, et al. The Novel ATR inhibitor BAY 1895344 is efficacious as monotherapy and combined with DNA damage-inducing or repair-compromising therapies in preclinical cancer models. Mol Cancer Ther. 2020;19:26–38.
Ferrao PT, Bukczynska EP, Johnstone RW, McArthur GA. Efficacy of CHK inhibitors as single agents in MYC-driven lymphoma cells. Oncogene. 2012;31:1661–72.
Shimura T, Ochiai Y, Noma N, Oikawa T, Sano Y, Fukumoto M. Cyclin D1 overexpression perturbs DNA replication and induces replication-associated DNA double-strand breaks in acquired radioresistant cells. Cell Cycle. 2013;12:773–82.
Aggarwal P, Lessie MD, Lin DI, Pontano L, Gladden AB, Nuskey B, et al. Nuclear accumulation of cyclin D1 during S phase inhibits Cul4-dependent Cdt1 proteolysis and triggers p53-dependent DNA rereplication. Genes Dev. 2007;21:2908–22.
Maya-Mendoza A, Ostrakova J, Kosar M, Hall A, Duskova P, Mistrik M, et al. Myc and Ras oncogenes engage different energy metabolism programs and evoke distinct patterns of oxidative and DNA replication stress. Mol Oncol. 2015;9:601–16.
Di Micco R, Fumagalli M, Cicalese A, Piccinin S, Gasparini P, Luise C, et al. Oncogene-induced senescence is a DNA damage response triggered by DNA hyper-replication. Nature. 2006;444:638–42.
Carlos AR, Escandell JM, Kotsantis P, Suwaki N, Bouwman P, Badie S, et al. ARF triggers senescence in Brca2-deficient cells by altering the spectrum of p53 transcriptional targets. Nat Commun. 2013;4:2697.
Kanu N, Cerone MA, Goh G, Zalmas L-P, Bartkova J, Dietzen M, et al. DNA replication stress mediates APOBEC3 family mutagenesis in breast cancer. Genome Biol. 2016;17:185.
Urzúa-Traslaviña CG, Leeuwenburgh VC, Bhattacharya A, Loipfinger S. van Vugt MATM, de Vries EGE et al. Improving gene function predictions using independent transcriptional components. Nat Commun. 2021;12:1464.
Liu H-Y, Liu Y-Y, Yang F, Zhang L, Zhang F-L, Hu X, et al. Acetylation of MORC2 by NAT10 regulates cell-cycle checkpoint control and resistance to DNA-damaging chemotherapy and radiotherapy in breast cancer. Nucleic Acids Res. 2020;48:3638–56.
Liu X, Tan Y, Zhang C, Zhang Y, Zhang L, Ren P, et al. NAT10 regulates p53 activation through acetylating p53 at K120 and ubiquitinating Mdm2. EMBO Rep. 2016;17:349–66.
Arango D, Sturgill D, Alhusaini N, Dillman AA, Sweet TJ, Hanson G, et al. Acetylation of Cytidine in mRNA promotes translation efficiency. Cell. 2018;175:1872–1886.e24.
Dominissini D, Rechavi G. N4-acetylation of Cytidine in mRNA by NAT10 regulates stability and translation. Cell. 2018;175:1725–7.
Shen Q, Zheng X, McNutt MA, Guang L, Sun Y, Wang J, et al. NAT10, a nucleolar protein, localizes to the midbody and regulates cytokinesis and acetylation of microtubules. Exp Cell Res. 2009;315:1653–67.
Guerrero Llobet S, van der Vegt B, Jongeneel E, Bense RD, Zwager MC, Schröder CP, et al. Cyclin E expression is associated with high levels of replication stress in triple-negative breast cancer. NPJ Breast Cancer. 2020;6:40.
Calzetta NL, González Besteiro MA, Gottifredi V. Mus81-Eme1-dependent aberrant processing of DNA replication intermediates in mitosis impairs genome integrity. Sci Adv. 2020;6:492–6.
Pillay N, Tighe A, Nelson L, Littler S, Coulson-Gilmer C, Bah N, et al. DNA replication vulnerabilities render ovarian cancer cells sensitive to Poly(ADP-Ribose) glycohydrolase inhibitors. Cancer Cell. 2019;35:519–533.e8.
Farshidfar F, Zheng S, Gingras M-C, Newton Y, Shih J, Robertson AG, et al. Integrative genomic analysis of cholangiocarcinoma identifies distinct IDH-mutant molecular profiles. Cell Rep. 2017;18:2780–94.
Gadhikar MA, Zhang J, Shen L, Rao X, Wang J, Zhao M, et al. CDKN2A/p16 deletion in head and neck cancer cells is associated with CDK2 activation, replication stress, and vulnerability to CHK1 inhibition. Cancer Res. 2018;78:781–97.
Williamson CT, Miller R, Pemberton HN, Jones SE, Campbell J, Konde A, et al. ATR inhibitors as a synthetic lethal therapy for tumours deficient in ARID1A. Nat Commun. 2016;7:13837.
Inoue S, Li WY, Tseng A, Beerman I, Elia AJ, Bendall SC, et al. Mutant IDH1 downregulates ATM and alters DNA repair and sensitivity to DNA damage independent of TET2. Cancer Cell. 2016;30:337–48.
Espana-Agusti J, Warren A, Chew SK, Adams DJ, Matakidou A. Loss of PBRM1 rescues VHL dependent replication stress to promote renal carcinogenesis. Nat Commun. 2017;8:2026.
Joseph NM, Tsokos CG, Umetsu SE, Shain AH, Kelley RK, Onodera C, et al. Genomic profiling of combined hepatocellular-cholangiocarcinoma reveals similar genetics to hepatocellular carcinoma. J Pathol. 2019;248:164–78.
de Jong MRW, Visser L, Huls G, Diepstra A, van Vugt M, Ammatuna E, et al. Identification of relevant drugable targets in diffuse large B-cell lymphoma using a genome-wide unbiased CD20 guilt-by association approach. PLoS One. 2018;13:e0193098.
Reddy A, Zhang J, Davis NS, Moffitt AB, Love CL, Waldrop A, et al. Genetic and functional drivers of diffuse Large B. Cell Lymphoma Cell. 2017;171:481–494.e15.
Tort F, Bartkova J, Sehested M, Orntoft T, Lukas J, Bartek J. Retinoblastoma pathway defects show differential ability to activate the constitutive DNA damage response in human tumorigenesis. Cancer Res. 2006;66:10258–63.
Konstantinopoulos PA, Cheng S-C, Wahner Hendrickson AE, Penson RT, Schumer ST, Doyle LA, et al. Berzosertib plus gemcitabine versus gemcitabine alone in platinum-resistant high-grade serous ovarian cancer: a multicentre, open-label, randomised, phase 2 trial. Lancet Oncol. 2020;21:957–68.
Italiano A, Infante JR, Shapiro GI, Moore KN, LoRusso PM, Hamilton E, et al. Phase I study of the checkpoint kinase 1 inhibitor GDC-0575 in combination with gemcitabine in patients with refractory solid tumors. Ann Oncol J Eur Soc Med Oncol. 2018;29:1304–11.
Rajeshkumar NV, De Oliveira E, Ottenhof N, Watters J, Brooks D, Demuth T, et al. MK-1775, a potent Wee1 inhibitor, synergizes with gemcitabine to achieve tumor regressions, selectively in p53-deficient pancreatic cancer xenografts. Clin Cancer Res. 2011;17:2799–806.
Acknowledgements
We thank members of the Medical Oncology laboratory for fruitful discussions. This work was financially supported by grant from the Netherlands Organization for Scientific Research (NWO-VIDI #917.13334 to M.A.T.M.v.V. and NWO-VENI #916-16025 to R.S.N.F.), the Dutch Cancer Society (RUG 2013-5960 to R.S.N.F and #11352 to M.A.T.M.v.V.) and from the European Research Council (ERC-Consolidator grant “TENSION” to M.A.T.M.v.V.).
Author information
Authors and Affiliations
Contributions
R.S.N.F. and M.A.T.M.v.V. conceived and supervised the study. S.G.L. performed in vitro studies. S.G.L., A.B., K.K. and R.S.N.F. generated and analyzed RNAseq data. A.B. and R.S.N.F. conducted computational analyses. M.E. and B.v.d.V. conducted immunohistochemical analysis of patient samples. S.G.L., A.B., R.S.N.F. and M.A.T.M.v.V. wrote the manuscript. All authors provided feedback and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
M.A.T.M.v.V. has acted on the Scientific Advisory Board of Repare Therapeutics, which is unrelated to this work. The other authors declare no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
shared senior authorship: Rudolf S.N. Fehrmann, Marcel A.T.M. van Vugt.
Rights and permissions
About this article
Cite this article
Guerrero Llobet, S., Bhattacharya, A., Everts, M. et al. An mRNA expression-based signature for oncogene-induced replication-stress. Oncogene 41, 1216–1224 (2022). https://doi.org/10.1038/s41388-021-02162-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-02162-0
This article is cited by
-
Dissecting the oncogenic properties of essential RNA-modifying enzymes: a focus on NAT10
Oncogene (2024)
-
Targeting ATR in patients with cancer
Nature Reviews Clinical Oncology (2024)
-
cGAS-STING pathway expression correlates with genomic instability and immune cell infiltration in breast cancer
npj Breast Cancer (2024)
-
Role of NAT10-mediated ac4C-modified HSP90AA1 RNA acetylation in ER stress-mediated metastasis and lenvatinib resistance in hepatocellular carcinoma
Cell Death Discovery (2023)
-
Targeting replication stress in cancer therapy
Nature Reviews Drug Discovery (2023)