Abstract
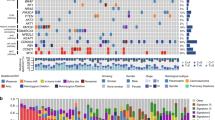
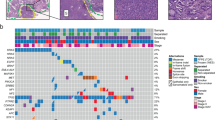
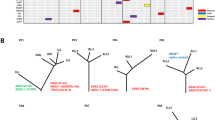
Pulmonary sarcomatoid carcinoma (PSC) contains carcinomatous component (CaC) and sarcomatous component (SaC). Herein, we explored the genomic origin and intratumor heterogeneity (ITH) of PSC. We collected 31 resected PSC tumors and obtained CaC and SaC by laser capture microdissection for next-generation sequencing. The majority of PSCs (97%) had component-shared alterations. Driver mutations in EGFR, KRAS, MET, PIK3CA, and EML4-ALK fusion were mostly component-shared. Twenty-seven (87%) PSCs had component-private alterations. Compared with pure lung adenocarcinoma (LUAD), adenocarcinoma component of PSC showed lower EGFR incidence. Compared with other typical sarcomas, numerous genes of SaC exhibited significant differences. CaC and SaC had equivalent and proportional tumor mutation burden (TMB), as well as PD-L1 level. Compared with LUAD, SaC had significant higher TMB and more patients with high PD-L1 expression (tumor proportion score ≥50%). PSC with lower proportion of component-shared alterations (trunk-ratio) had a prolonged disease-free survival (DFS), regardless of the influence of clinical factors. We conclude that most PSCs originate from a monoclone accompanied by genomic ITH which is a potential independent prognostic factor, and more proportion of PSCs may be beneficial from immune checkpoint inhibitors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The datasets supporting the conclusions of this article are available from the corresponding author on reasonable request.
References
Weissferdt A, Kalhor N, Correa AM, Moran CA. “Sarcomatoid” carcinomas of the lung: a clinicopathological study of 86 cases with a new perspective on tumor classification. Hum Pathol. 2017;63:14–26.
Yendamuri S, Caty L, Pine M, Adem S, Bogner P, Miller A, et al. Outcomes of sarcomatoid carcinoma of the lung: a Surveillance, Epidemiology, and End Results Database analysis. Surgery 2012;152:397–402.
Travis WD, Brambilla E, Nicholson AG, Yatabe Y, Austin JH, Beasley MB, et al. The 2015 World Health Organization classification of lung tumors: impact of genetic, clinical and radiologic advances since the 2004 classification. J Thorac Oncol. 2015;10:1243–60.
Martin LW, Correa AM, Ordonez NG, Roth JA, Swisher SG, Vaporciyan AA, et al. Sarcomatoid carcinoma of the lung: a predictor of poor prognosis. Ann Thorac Surg. 2007;84:973–80.
Vieira T, Girard N, Ung M, Monnet I, Cazes A, Bonnette P, et al. Efficacy of first-line chemotherapy in patients with advanced lung sarcomatoid carcinoma. J Thorac Oncol. 2013;8:1574–7.
Pelosi G, Sonzogni A, De Pas T, Galetta D, Veronesi G, Spaggiari L, et al. Review Article: Pulmonary sarcomatoid carcinomas: a practical overview. Int J Surgical Pathol. 2009;18:103–20.
Li Y, Gao L, Ma D, Qiu T, Li W, Li W, et al. Identification of MET exon14 skipping by targeted DNA-and RNA-based next-generation sequencing in pulmonary sarcomatoid carcinomas. Lung Cancer. 2018;122:113–9.
Liu X, Jia Y, Stoopler MB, Shen Y, Cheng H, Chen J, et al. Next-generation sequencing of pulmonary sarcomatoid carcinoma reveals high frequency of actionable MET gene mutations. J Clin Oncol. 2016;34:794–802.
Lococo F, Gandolfi G, Rossi G, Pinto C, Rapicetta C, Cavazza A, et al. Deep sequencing analysis reveals that KRAS mutation is a marker of poor prognosis in patients with pulmonary sarcomatoid carcinoma. J Thorac Oncol. 2016;11:1282–92.
Mehrad M, Roy S, LaFramboise WA, Petrosko P, Miller C, Incharoen P, et al. KRAS mutation is predictive of outcome in patients with pulmonary sarcomatoid carcinoma. Histopathology 2018;73:207–14.
Terra SB, Jang JS, Bi L, Kipp BR, Jen J, Eunhee SY, et al. Molecular characterization of pulmonary sarcomatoid carcinoma: analysis of 33 cases. Mod Pathol. 2016;29:824–31.
Schrock AB, Li SD, Frampton GM, Suh J, Braun E, Mehra R, et al. Pulmonary sarcomatoid carcinomas commonly harbor either potentially targetable genomic alterations or high tumor mutational burden as observed by comprehensive genomic profiling. J Thorac Oncol. 2017;12:932–42.
Liang X, Li Q, Xu B, Hu S, Wang Q, Li Y, et al. Mutation landscape and tumor mutation burden analysis of Chinese patients with pulmonary sarcomatoid carcinomas. Int J Clin Oncol. 2019;24:1061–8.
Kim S, Kim MY, Koh J, Go H, Lee DS, Jeon YK, et al. Programmed death-1 ligand 1 and 2 are highly expressed in pleomorphic carcinomas of the lung: comparison of sarcomatous and carcinomatous areas. Eur J Cancer. 2015;51:2698–707.
Goto T, Hirotsu Y, Mochizuki H, Nakagomi T, Oyama T, Amemiya K, et al. Stepwise addition of genetic changes correlated with histological change from “well-differentiated” to “sarcomatoid” phenotypes: a case report. BMC Cancer. 2017;17:65.
Manzotti G, Torricelli F, Benedetta D, Lococo F, Sancisi V, Rossi G, et al. An epithelial-to-mesenchymal transcriptional switch triggers evolution of pulmonary sarcomatoid carcinoma (PSC) and identifies dasatinib as new therapeutic option. Cancer Res. 2019;25:2348–60.
Nakagomi T, Goto T, Hirotsu Y, Shikata D, Yokoyama Y, Higuchi R, et al. New therapeutic targets for pulmonary sarcomatoid carcinomas based on their genomic and phylogenetic profiles. Oncotarget. 2018;9:10635–49.
Fallet V, Saffroy R, Girard N, Mazieres J, Lantuejoul S, Vieira T, et al. High-throughput somatic mutation profiling in pulmonary sarcomatoid carcinomas using the LungCarta™ Panel: exploring therapeutic targets. Ann Oncol. 2015;26:1748–53.
Sanchez-Vega F, Mina M, Armenia J, Chatila WK, Luna A, La KC, et al. Oncogenic signaling pathways in the cancer genome atlas. Cell 2018;173:321–37.
Blaukovitsch M, Halbwedl I, Kothmaier H, Gogg-Kammerer M, Popper HH. Sarcomatoid carcinomas of the lung—are these histogenetically heterogeneous tumors? Virchows Arch. 2006;449:455–61.
D’Angelo SP, Pietanza MC, Johnson ML, Riely GJ, Miller VA, Sima CS, et al. Incidence of EGFR exon 19 deletions and L858R in tumor specimens from men and cigarette smokers with lung adenocarcinomas. J Clin Oncol. 2011;29:2066.
Chang YL, Wu CT, Shih JY, Lee YC. EGFR and p53 status of pulmonary pleomorphic carcinoma: implications for EGFR tyrosine kinase inhibitors therapy of an aggressive lung malignancy. Ann Surgical Oncol. 2011;18:2952–60.
Italiano A, Cortot AB, Ilie M, Martel-Planche G, Fabas T, Pop D, et al. EGFR and KRAS status of primary sarcomatoid carcinomas of the lung: implications for anti-EGFR treatment of a rare lung malignancy. Int J Cancer. 2009;125:2479–82.
Kaira K, Horie Y, Ayabe E, Murakami H, Takahashi T, Tsuya A, et al. Pulmonary pleomorphic carcinoma: a clinicopathological study including EGFR mutation analysis. J Thorac Oncol. 2010;5:460–5.
Leone A, Graziano P, Gasbarra R, Paone G, Cardillo G, Mancuso A, et al. Identification of EGFR mutations in lung sarcomatoid carcinoma. Int J Cancer. 2011;128:732–5.
Pelosi G, Gasparini P, Cavazza A, Rossi G, Graziano P, Barbareschi M, et al. Multiparametric molecular characterization of pulmonary sarcomatoid carcinoma reveals a nonrandom amplification of anaplastic lymphoma kinase (ALK) gene. Lung Cancer. 2012;77:507–14.
Tong JH, Yeung SF, Chan AW, Chung LY, Chau SL, Lung RWM, et al. MET amplification and exon 14 splice site mutation define unique molecular subgroups of non–small cell lung carcinoma with poor prognosis. Clin Cancer Res. 2016;22:3048–56.
Yu Y, Zhang Q, Zhang J, Lu S. Prevalence of MET exon 14 skipping mutation in pulmonary sarcomatoid carcinoma patients without common targetable mutations: a single-institute study. J Cancer Res Therapeutics. 2019;15:909.
Lin G, Li C, Li PS, Fang WZ, Xu HP, Gong YH, et al. Genomic origin and EGFR-TKI treatments of pulmonary adenosquamous carcinoma. Ann Oncol. 2020;31:517–24.
McGranahan N, Favero F, de Bruin EC, Birkbak NJ, Szallasi Z, Swanton C. Clonal status of actionable driver events and the timing of mutational processes in cancer evolution. Sci Transl Med. 2015;7:283ra54–ra54.
Gibney GT, Weiner LM, Atkins MB. Predictive biomarkers for checkpoint inhibitor-based immunotherapy. Lancet Oncol. 2016;17:e542–e51.
Chan TA, Yarchoan M, Jaffee E, Swanton C, Quezada SA, Stenzinger A, et al. Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann Oncol. 2019;30:44–56.
Cimpeanu E, Ahmed J, Zafar W, DeMarinis A, Bardarov SS, Salman S, et al. Pembrolizumab-emerging treatment of pulmonary sarcomatoid carcinoma: a case report. World J Clin Cases 2020;8:97.
Babacan NA, Pina IB, Signorelli D, Prelaj A, Garassino MC, Tanvetyanon T. Relationship between programmed death receptor-ligand 1 expression and response to checkpoint inhibitor immunotherapy in pulmonary sarcomatoid carcinoma: a pooled analysis. Clinical Lung Cancer. 2020;21:e456–e463.
Lin Y, Yang H, Cai Q, Wang D, Rao H, Lin S, et al. Characteristics and prognostic analysis of 69 patients with pulmonary sarcomatoid carcinoma. Am J Clin Oncol. 2016;39:215–22.
Sim JK, Chung SM, Choi JH, Oh JY, Lee SH, Kim JH, et al. Clinical and molecular characteristics of pulmonary sarcomatoid carcinoma. Korean J Intern Med. 2018;33:737–44.
Chen F, Gu Q, Hu C, Cai X, Lei S. Poor prognosis of pulmonary sarcomatoid carcinoma with KRAS mutation and ALK fusion. Onco Targets Ther. 2019;12:3321–5.
McGranahan N, Swanton C. Clonal heterogeneity and tumor evolution: past, present, and the future. Cell 2017;168:613–28.
Bevilacqua C, Ducos B. Laser microdissection: a powerful tool for genomics at cell level. Mol Asp Med. 2018;59:5–27.
Nong J, Gong Y, Guan Y, Yi X, Yi Y, Chang L, et al. Circulating tumor DNA analysis depicts subclonal architecture and genomic evolution of small cell lung cancer. Nat Commun. 2018;9:3114.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C, et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol. 2013;31:213.
Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164.
Li J, Lupat R, Amarasinghe KC, Thompson ER, Doyle MA, Ryland GL, et al. CONTRA: copy number analysis for targeted resequencing. Bioinformatics. 2012;28:1307–13.
Chen K, Wallis JW, McLellan MD, Larson DE, Kalicki JM, Pohl CS, et al. BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nat Methods. 2009;6:677–81.
Roth A, Khattra J, Yap D, Wan A, Laks E, Biele J, et al. PyClone: statistical inference of clonal population structure in cancer. Nat Methods. 2014;11:396–8.
Lausen B, Schumacher M. Maximally selected rank statistics. Biometrics. 1992;48:73–85.
Acknowledgements
This study was supported by the National Natural Science Foundation of China (81572270), Nature Science Foundation of Hunan Province (2017JJ3188), Foundation of Hunan Health Committee Research Plan (A2017005) and Foundation of Wisdom Gathering and Talent Cultivating Program from the Third Xiangya Hospital.
Author information
Authors and Affiliations
Contributions
L Chen and CM conceived the study and designed the experiments. XL, FW, XC, XH, ZX, and YL prepared the research materials. QL, PL, and L Chang did the bioinformatic analysis and performed initial exploratory analysis. YG, XZ, LY, CX, HW, XY, JZ, and XX provided insight in methodological approaches and analysis. XL and L Chen supervised the study. XL, FW, CX, XC, XH, QL, and PL drafted the paper. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare the following conflict of interest: QL, PL, L Chang, YG, and XX are current employees of Geneplus-Beijing. XY and LY hold leadership positions and stocks of Geneplus-Beijing. JZ reports grants from Merck, Johnson and Johnson, consultant fees, advisory fees or honoraria from Bristol Myers Squibb, AstraZeneca, Geneplus, Innovent, OrigMed, Roche outside the submitted work. The remaining authors declare no conflict of interest.
Ethical approval
The present study was reviewed and approved by the Research Ethics Committee of the Sun Yat-sen University Cancer Center (B2020-139-01).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, X., Wang, F., Xu, C. et al. Genomic origin and intratumor heterogeneity revealed by sequencing on carcinomatous and sarcomatous components of pulmonary sarcomatoid carcinoma. Oncogene 40, 821–832 (2021). https://doi.org/10.1038/s41388-020-01573-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-01573-9