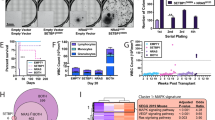
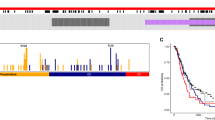
Abstract
The serine/threonine Protein Phosphatase 2A (PP2A) functions as a tumor suppressor by negatively regulating multiple oncogenic signaling pathways. The canonical PP2A holoenzyme comprises a scaffolding subunit (PP2A Aα/β), which serves as the platform for binding of both the catalytic C subunit and one regulatory B subunit. Somatic heterozygous missense mutations in PPP2R1A, the gene encoding the PP2A Aα scaffolding subunit, have been identified across multiple cancer types, but the effects of the most commonly mutated residue, Arg-183, on PP2A function have yet to be fully elucidated. In this study, we used a series of cellular and in vivo models and discovered that the most frequent Aα R183W mutation formed alternative holoenzymes by binding of different PP2A regulatory subunits compared with wild-type Aα, suggesting a rededication of PP2A functions. Unlike wild-type Aα, which suppressed tumorigenesis, the R183W mutant failed to suppress tumor growth in vivo through activation of the MAPK pathway in RAS-mutant transformed cells. Furthermore, cells expressing R183W were less sensitive to MEK inhibitors. Taken together, our results demonstrate that the R183W mutation in PP2A Aα scaffold abrogates the tumor suppressive actions of PP2A, thereby potentiating oncogenic signaling and reducing drug sensitivity of RAS-mutant cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kauko O, Laajala TD, Jumppanen M, Hintsanen P, Suni V, Haapaniemi P, et al. Label-free quantitative phosphoproteomics with novel pairwise abundance normalization reveals synergistic RAS and CIP2A signaling. Sci Rep. 2015;5:13099.
Rangarajan A, Hong SJ, Gifford A, Weinberg RA. Species- and cell type-specific requirements for cellular transformation. Cancer Cell. 2004;6:171–83.
Sablina AA, Hector M, Colpaert N, Hahn WC. Identification of PP2A complexes and pathways involved in cell transformation. Cancer Res. 2010;70:10474–84.
Sablina AA, Chen W, Arroyo JD, Corral L, Hector M, Bulmer SE, et al. The tumor suppressor PP2A Abeta regulates the RalA GTPase. Cell. 2007;129:969–82.
Chen W, Possemato R, Campbell KT, Plattner CA, Pallas DC, Hahn WC. Identification of specific PP2A complexes involved in human cell transformation. Cancer Cell. 2004;5:127–36.
Sangodkar J, Perl A, Tohme R, Kiselar J, Kastrinsky DB, Zaware N, et al. Activation of tumor suppressor protein PP2A inhibits KRAS-driven tumor growth. J Clin Investig. 2017;127:2081–90.
Kauko O, O’Connor CM, Kulesskiy E, Sangodkar J, Aakula A, Izadmehr S, et al. PP2A inhibition is a druggable MEK inhibitor resistance mechanism in KRAS-mutant lung cancer cells. Sci Transl Med. 2018;10:eaaq1093.
Seshacharyulu P, Pandey P, Datta K, Batra SK. Phosphatase: PP2A structural importance, regulation and its aberrant expression in cancer. Cancer Lett. 2013;335:9–18.
Shi Y. Serine/threonine phosphatases: mechanism through structure. Cell. 2009;139:468–84.
Sangodkar J, Farrington CC, McClinch K, Galsky MD, Kastrinsky DB, Narla G. All roads lead to PP2A: exploiting the therapeutic potential of this phosphatase. FEBS J. 2016;283:1004–24.
Zhou J, Pham HT, Ruediger R, Walter G. Characterization of the Aalpha and Abeta subunit isoforms of protein phosphatase 2A: differences in expression, subunit interaction, and evolution. Biochemical J. 2003;369(Pt 2):387–98.
Groves MR, Hanlon N, Turowski P, Hemmings BA, Barford D. The structure of the protein phosphatase 2A PR65/A subunit reveals the conformation of its 15 tandemly repeated HEAT motifs. Cell. 1999;96:99–110.
Calin GA, di Iasio MG, Caprini E, Vorechovsky I, Natali PG, Sozzi G, et al. Low frequency of alterations of the alpha (PPP2R1A) and beta (PPP2R1B) isoforms of the subunit A of the serine-threonine phosphatase 2A in human neoplasms. Oncogene. 2000;19:1191–5.
Shih IeM, Panuganti PK, Kuo KT, Mao TL, Kuhn E, Jones S, et al. Somatic mutations of PPP2R1A in ovarian and uterine carcinomas. Am J Pathol. 2011;178:1442–7.
Ruediger R, Pham HT, Walter G. Alterations in protein phosphatase 2A subunit interaction in human carcinomas of the lung and colon with mutations in the A beta subunit gene. Oncogene. 2001;20:1892–9.
Haesen D, Abbasi Asbagh L, Derua R, Hubert A, Schrauwen S, Hoorne Y, et al. Recurrent PPP2R1A mutations in uterine cancer act through a dominant-negative mechanism to promote malignant cell growth. Cancer Res. 2016;76:5719–31.
Chen W, Arroyo JD, Timmons JC, Possemato R, Hahn WC. Cancer-associated PP2A Aalpha subunits induce functional haploinsufficiency and tumorigenicity. Cancer Res. 2005;65:8183–92.
Walter G, Ruediger R. Mouse model for probing tumor suppressor activity of protein phosphatase 2A in diverse signaling pathways. Cell Cycle. 2012;11:451–9.
Ruediger R, Ruiz J, Walter G. Human cancer-associated mutations in the Aalpha subunit of protein phosphatase 2A increase lung cancer incidence in Aalpha knock-in and knockout mice. Mol Cell Biol. 2011;31:3832–44.
Jeong AL, Han S, Lee S, Su Park J, Lu Y, Yu S, et al. Patient derived mutation W257G of PPP2R1A enhances cancer cell migration through SRC-JNK-c-Jun pathway. Sci Rep. 2016;6:27391.
Kamburov A, Lawrence MS, Polak P, Leshchiner I, Lage K, Golub TR, et al. Comprehensive assessment of cancer missense mutation clustering in protein structures. Proc Natl Acad Sci USA. 2015;112:E5486–95.
Rahman M, Nakayama K, Rahman MT, Nakayama N, Katagiri H, Katagiri A, et al. PPP2R1A mutation is a rare event in ovarian carcinoma across histological subtypes. Anticancer Res. 2013;33:113–8.
Hoang LN, McConechy MK, Meng B, McIntyre JB, Ewanowich C, Gilks CB, et al. Targeted mutation analysis of endometrial clear cell carcinoma. Histopathology. 2015;66:664–74.
Jones S, Wang TL, Shih Ie M, Mao TL, Nakayama K, Roden R, et al. Frequent mutations of chromatin remodeling gene ARID1A in ovarian clear cell carcinoma. Science. 2010;330:228–31.
Junk DJ, Cipriano R, Bryson BL, Gilmore HL, Jackson MW. Tumor microenvironmental signaling elicits epithelial-mesenchymal plasticity through cooperation with transforming genetic events. Neoplasia. 2013;15:1100–9.
Pallas DC, Shahrik LK, Martin BL, Jaspers S, Miller TB, Brautigan DL, et al. Polyoma small and middle T antigens and SV40 small t antigen form stable complexes with protein phosphatase 2A. Cell. 1990;60:167–76.
O’Connor CM, Hoffa MT, Taylor SE, Avelar RA, Narla G. Protein phosphatase 2A Aalpha regulates Abeta protein expression and stability. J Biol Chem. 2019;294:5923–34.
Taylor SE, O’Connor CM, Wang Z, Shen G, Song H, Leonard D, et al. The highly recurrent PP2A Aalpha-subunit mutation P179R alters protein structure and impairs PP2A enzyme function to promote endometrial tumorigenesis. Cancer Res. 2019;79:4242–57.
Strack S, Cribbs JT, Gomez L. Critical role for protein phosphatase 2A heterotrimers in mammalian cell survival. J Biol Chem. 2004;279:47732–9.
Casado P, Rodriguez-Prados JC, Cosulich SC, Guichard S, Vanhaesebroeck B, Joel S, et al. Kinase-substrate enrichment analysis provides insights into the heterogeneity of signaling pathway activation in leukemia cells. Sci Signal. 2013;6:rs6.
Wiredja DD, Koyuturk M, Chance MR. The KSEA app: a web-based tool for kinase activity inference from quantitative phosphoproteomics. Bioinformatics. 2017.
Letourneux C, Rocher G, Porteu F. B56-containing PP2A dephosphorylate ERK and their activity is controlled by the early gene IEX-1 and ERK. EMBO J. 2006;33:3489–91.
Horn T, Sandmann T, Fischer B, Axelsson E, Huber W, Boutros M. Mapping of signaling networks through synthetic genetic interaction analysis by RNAi. Nat Methods. 2011;8:341–6.
Holt SV, Logie A, Davies BR, Alferez D, Runswick S, Fenton S, et al. Enhanced apoptosis and tumor growth suppression elicited by combination of MEK (selumetinib) and mTOR kinase inhibitors (AZD8055). Cancer Res. 2012;72:1804–13.
Roberts PJ, Usary JE, Darr DB, Dillon PM, Pfefferle AD, Whittle MC, et al. Combined PI3K/mTOR and MEK inhibition provides broad antitumor activity in faithful murine cancer models. Clin Cancer Res. 2012;18:5290–303.
Grinthal A, Adamovic I, Weiner B, Karplus M, Kleckner N. PR65, the HEAT-repeat scaffold of phosphatase PP2A, is an elastic connector that links force and catalysis. Proc Natl Acad Sci USA. 2010;107:2467–72.
Tsytlonok M, Craig PO, Sivertsson E, Serquera D, Perrett S, Best RB, et al. Complex energy landscape of a giant repeat protein. Structure. 2013;21:1954–65.
Caunt CJ, Sale MJ, Smith PD, Cook SJ. MEK1 and MEK2 inhibitors and cancer therapy: the long and winding road. Nat Rev Cancer. 2015;15:577–92.
Garbe JC, Bhattacharya S, Merchant B, Bassett E, Swisshelm K, Feiler HS, et al. Molecular distinctions between stasis and telomere attrition senescence barriers shown by long-term culture of normal human mammary epithelial cells. Cancer Res. 2009;69:7557–68.
Cipriano R, Kan CE, Graham J, Danielpour D, Stampfer M, Jackson MW. TGF-beta signaling engages an ATM-CHK2-p53-independent RAS-induced senescence and prevents malignant transformation in human mammary epithelial cells. Proc Natl Acad Sci USA. 2011;108:8668–73.
Wiemann SPC, Hu Y, Hunter P, Harbers M, Amiet A, Bethel G, et al. The ORFeome Collaboration: a genome-scale human ORF-clone resource. Nat Methods. 2016;13:191–2.
Schlatzer D, Haqqani AA, Li X, Dobrowolski C, Chance MR, Tilton JC. A targeted mass spectrometry assay for detection of HIV gag protein following induction of latent viral reservoirs. Anal Chem. 2017;89:5325–32.
Mellacheruvu D, Wright Z, Couzens AL, Lambert JP, St-Denis NA, Li T, et al. The CRAPome: a contaminant repository for affinity purification-mass spectrometry data. Nat Methods. 2013;10:730–6.
Morris JH, Knudsen GM, Verschueren E, Johnson JR, Cimermancic P, Greninger AL, et al. Affinity purification-mass spectrometry and network analysis to understand protein-protein interactions. Nat Protoc. 2014;9:2539–54.
Knight JDR, Choi H, Gupta GD, Pelletier L, Raught B, Nesvizhskii AI, et al. ProHits-viz: a suite of web tools for visualizing interaction proteomics data. Nat Methods. 2017;14:645–6.
Wisniewski JR, Zougman A, Nagaraj N, Mann M. Universal sample preparation method for proteome analysis. Nat Methods. 2009;6:359–62.
McClinch K, Avelar RA, Callejas D, Izadmehr S, Wiredja D, Perl A, et al. Small-molecule activators of protein phosphatase 2A for the treatment of castration-resistant prostate cancer. Cancer Res. 2018;78:2065–80.
Fabregat A, Sidiropoulos K, Garapati P, Gillespie M, Hausmann K, Haw R, et al. The reactome pathway knowledgebase. Nucleic acids Res. 2016;44(D1):D481–7.
Abagyan R, Totrov M, Kuznetsov D. ICM- A new method for protein modeling and design: applicataions to docking and structure prediction from the distorted native conformation. J Comput Chem. 1994;15:488–506.
Doerr S, Harvey MJ, Noe F, De Fabritiis G. HTMD: high-throughput molecular dynamics for molecular discovery. J Chem Theory Comput 2016;12:1845–52.
Case DA, Cheatham TE 3rd, Darden T, Gohlke H, Luo R, Merz KM Jr, et al. The amber biomolecular simulation programs. J Comput Chem 2005;26:1668–88.
Price DJ, Brooks CL 3rd. A modified TIP3P water potential for simulation with Ewald summation. J Chem Phys. 2004;121:10096–103.
Berendsen HJC, Postma JPM, Gunsteren WFv, DiNola A, Haak JR. Molecular dynamics with coupling to an external bath. J Chem Phy. 1984;81:3684–90.
Harvey MJ, Giupponi G, Fabritiis GD. ACEMD: accelerating biomolecular dynamics in the microsecond time scale. J Chem Theory Comput 2009;5:1632–9.
Bonomi M, Barducci A, Parrinello M. Reconstructing the equilibrium Boltzmann distribution from well-tempered metadynamics. J Comput Chem 2009;30:1615–21.
Skliros A, Zimmermann MT, Chakraborty D, Saraswathi S, Katebi AR, Leelananda SP, et al. The importance of slow motions for protein functional loops. Phys Biol. 2012;9:014001.
Hess B, Kutzner C, van der Spoel D, Lindahl E. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J Chem Theory Comput. 2008;4:435–47.
DeLano W PyMOL molecular viewer: Updates and refinements. Abstracts of Papers of the American Chemical Society. 2009;238.
Humphrey W, Dalke A, Schulten K. VMD: visual molecular dynamics. J Mol Graph. 1996;14:33–8, 27-8.
Acknowledgements
The authors wish to acknowledge the Young Scientist Foundation and the students who worked on this project through this program, including Aditya Upadhyay. This work was partially supported by the NIH/NCI R01CA181654 (to GN) and T32GM008803 (to CMO). This publication was made possible in part by the Clinical and Translational Science Collaborative of Cleveland, 4UL1TR000439 from the National Center for Advancing Translational Sciences (NCATS) component of the National Institutes of Health and NIH roadmap for Medical Research. This research was supported by the Athymic Animal and Preclinical Therapeutics and Cytometry and Imaging Microscopy cores of the Case Comprehensive Cancer Center at Case Western Reserve University (funded by NIH P30 CA043703). GN is supported by the Pardee-Gerstacker Professorship in Cancer Research.
Author information
Authors and Affiliations
Contributions
CMO and GN designed the research studies. CMO wrote the manuscript and GN edited the manuscript. CMO, DL, DW, RA, ZW, DS, BB, ET, AU, ST, and SH performed experiments and interpreted the data. JS, A-CG, JW, WX, AD, DB, SH, MJ, and GN provided observations and scientific interpretations. All authors discussed results and provided input on the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
O’Connor, C.M., Leonard, D., Wiredja, D. et al. Inactivation of PP2A by a recurrent mutation drives resistance to MEK inhibitors. Oncogene 39, 703–717 (2020). https://doi.org/10.1038/s41388-019-1012-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-1012-2
This article is cited by
-
The PPP2R1A cancer hotspot mutant p.R183W increases clofarabine resistance in uterine serous carcinoma cells by a gain-of-function mechanism
Cellular Oncology (2024)
-
PPP2R1A regulates migration persistence through the NHSL1-containing WAVE Shell Complex
Nature Communications (2023)
-
Genomic regulation of transcription and RNA processing by the multitasking Integrator complex
Nature Reviews Molecular Cell Biology (2023)
-
Mutant SETBP1 enhances NRAS-driven MAPK pathway activation to promote aggressive leukemia
Leukemia (2021)
-
Phosphatase PP2A enhances MCL-1 protein half-life in multiple myeloma cells
Cell Death & Disease (2021)