Abstract
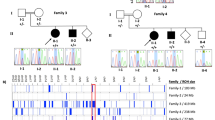
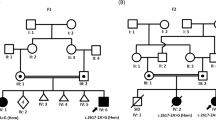
Lissencephaly is a rare brain malformation characterized by abnormal neuronal migration during cortical development. In this study, we performed a comprehensive genetic analysis using next-generation sequencing in 12 unsolved Japanese lissencephaly patients, in whom PAFAH1B1, DCX, TUBA1A, and ARX variants were excluded using the Sanger method. Exome sequencing (ES) was conducted on these 12 patients, identifying pathogenic variants in CEP85L, DYNC1H1, LAMC3, and DCX in four patients. Next, we performed genome sequencing (GS) on eight unsolved patients, and structural variants in PAFAH1B1, including an inversion and microdeletions involving several exons, were detected in three patients. Notably, these microdeletions in PAFAH1B1 could not to be detected by copy number variation (CNV) detection tools based on the depth of coverage methods using ES data. The density of repeat sequences, including Alu sequences or segmental duplications, which increase the susceptibility to structural variations, is very high in some lissencephaly spectrum genes (PAFAH1B1, TUBA1A, DYNC1H1). These missing CNVs were due to the limitations of detecting repeat sequences in ES-based CNV detection tools. Our study suggests that a combined approach integrating ES with GS can contribute to a higher diagnostic yield and a better understanding of the genetic landscape of the lissencephaly spectrum.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Oegema R, Barakat TS, Wilke M, Stouffs K, Amrom D, Aronica E, et al. International consensus recommendations on the diagnostic work-up for malformations of cortical development. Nat Rev Neurol. 2020;16:618–35.
Guerrini R, Filippi T. Neuronal migration disorders, genetics, and epileptogenesis. J Child Neurol. 2005;20:287–99.
Kato M, Dobyns WB. Lissencephaly and the molecular basis of neuronal migration. Hum Mol Genet. 2003;12:R89–96.
Barkovich AJ, Guerrini R, Kuzniecky RI, Jackson GD, Dobyns WB. A developmental and genetic classification for malformations of cortical development: update 2012. Brain. 2012;135:1348–69.
Di Donato N, Timms AE, Aldinger KA, Mirzaa GM, Bennett JT, Collins S, et al. Analysis of 17 genes detects mutations in 81% of 811 patients with lissencephaly. Genet Med. 2018;20:1354–64.
Kitamura K, Yanazawa M, Sugiyama N, Miura H, Iizuka-Kogo A, Kusaka M, et al. Mutation of ARX causes abnormal development of forebrain and testes in mice and X-linked lissencephaly with abnormal genitalia in humans. Nat Genet. 2002;32:359–69.
Gleeson JG, Allen KM, Fox JW, Lamperti ED, Berkovic S, Scheffer I, et al. Doublecortin, a brain-specific gene mutated in human X-linked lissencephaly and double cortex syndrome, encodes a putative signaling protein. Cell. 1998;92:63–72.
Vissers LE, de Ligt J, Gilissen C, Janssen I, Steehouwer M, de Vries P, et al. A de novo paradigm for mental retardation. Nat Genet. 2010;42:1109–12.
Reiner O, Carrozzo R, Shen Y, Wehnert M, Faustinella F, Dobyns WB, et al. Isolation of a Miller-Dieker lissencephaly gene containing G protein beta-subunit-like repeats. Nature. 1993;364:717–21.
Keays DA, Tian G, Poirier K, Huang GJ, Siebold C, Cleak J, et al. Mutations in alpha-tubulin cause abnormal neuronal migration in mice and lissencephaly in humans. Cell. 2007;128:45–57.
Watanabe K, Nakashima M, Kumada S, Mashimo H, Enokizono M, Yamada K, et al. Identification of two novel de novo TUBB variants in cases with brain malformations: case reports and literature review. J Hum Genet. 2021;66:1193–7.
Fromer M, Moran JL, Chambert K, Banks E, Bergen SE, Ruderfer DM, et al. Discovery and statistical genotyping of copy-number variation from whole-exome sequencing depth. Am J Hum Genet. 2012;91:597–607.
Uchiyama Y, Yamaguchi D, Iwama K, Miyatake S, Hamanaka K, Tsuchida N, et al. Efficient detection of copy-number variations using exome data: Batch- and sex-based analyses. Hum Mutat. 2021;42:50–65.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Hiraide T, Shimizu K, Miyamoto S, Aoto K, Nakashima M, Yamaguchi T, et al. Genome sequencing and RNA sequencing of urinary cells reveal an intronic FBN1 variant causing aberrant splicing. J Hum Genet. 2022;67:387–92.
Roller E, Ivakhno S, Lee S, Royce T, Tanner S. Canvas: versatile and scalable detection of copy number variants. Bioinformatics. 2016;32:2375–7.
Chen X, Schulz-Trieglaff O, Shaw R, Barnes B, Schlesinger F, Källberg M, et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics. 2016;32:1220–2.
Yu T, Huang X, Dou S, Tang X, Luo S, Theurkauf WE, et al. A benchmark and an algorithm for detecting germline transposon insertions and measuring de novo transposon insertion frequencies. Nucleic Acids Res. 2021;49:e44.
Li D, Liu CM, Luo R, Sadakane K, Lam TW. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics. 2015;31:1674–6.
Kanda Y. Investigation of the freely available easy-to-use software ‘EZR’ for medical statistics. Bone Marrow Transpl. 2013;48:452–8.
Dobyns WB, Truwit CL, Ross ME, Matsumoto N, Pilz DT, Ledbetter DH, et al. Differences in the gyral pattern distinguish chromosome 17-linked and X-linked lissencephaly. Neurology. 1999;53:270–7.
Abe K, Ando K, Kato M, Saitsu H, Nakashima M, Aoki S, et al. A new case with cortical malformation caused by biallelic variants in LAMC3. Neurol Genet. 2022;8:e680.
Pejaver V, Byrne AB, Feng BJ, Pagel KA, Mooney SD, Karchin R, et al. Calibration of computational tools for missense variant pathogenicity classification and ClinGen recommendations for PP3/BP4 criteria. Am J Hum Genet. 2022;109:2163–77.
Miyata Y, Kosuga M, Kato M, Watanabe K, Nakashima M, Saitsu H, et al. Microlissencephaly caused by a novel compound heterozygous variant in the WDR81 gene: A case report. No To Hattatsu. In press 2025.
Zou D, Wang L, Liao J, Xiao H, Duan J, Zhang T, et al. Genome sequencing of 320 Chinese children with epilepsy: a clinical and molecular study. Brain. 2021;144:3623–34.
Mei D, Lewis R, Parrini E, Lazarou LP, Marini C, Pilz DT, et al. High frequency of genomic deletions–and a duplication–in the LIS1 gene in lissencephaly: implications for molecular diagnosis. J Med Genet. 2008;45:355–61.
Arcot SS, Shaikh TH, Kim J, Bennett L, Alegria-Hartman M, Nelson DO, et al. Sequence diversity and chromosomal distribution of “young” Alu repeats. Gene. 1995;163:273–8.
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, et al. Initial sequencing and analysis of the human genome. Nature. 2001;409:860–921.
Morales ME, White TB, Streva VA, DeFreece CB, Hedges DJ, Deininger PL. The contribution of alu elements to mutagenic DNA double-strand break repair. PLoS Genet. 2015;11:e1005016.
Lobachev KS, Stenger JE, Kozyreva OG, Jurka J, Gordenin DA, Resnick MA. Inverted Alu repeats unstable in yeast are excluded from the human genome. Embo J. 2000;19:3822–30.
Deininger PL, Batzer MA, Hutchison CA 3rd, Edgell MH. Master genes in mammalian repetitive DNA amplification. Trends Genet. 1992;8:307–11.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–10.
Nord AS, Lee M, King MC, Walsh T. Accurate and exact CNV identification from targeted high-throughput sequence data. BMC Genomics. 2011;12:184.
Haverfield EV, Whited AJ, Petras KS, Dobyns WB, Das S. Intragenic deletions and duplications of the LIS1 and DCX genes: a major disease-causing mechanism in lissencephaly and subcortical band heterotopia. Eur J Hum Genet. 2009;17:911–8.
Acknowledgements
We thank the patient’s family for their participation in this study. We thank M. Tsujimura, K Shibasaki, M Tanaka, C. Ogawa and A. Kitamoto for their technical assistance. This work was supported in part by the Japan Society for the Promotion of Science, KAKENHI (Grant number JP 20H03641 and 23H02875 (HS), JP20K08236 (MK) and JP21K06819 (MN)), the Japan Agency for Medical Research and Development (AMED) (JP23ek0109549 and 23ek01099674, HS), the Takeda Science Foundation (MN and HS), and HUSM Grant-in-Aid from Hamamatsu University School of Medicine (MN and HS).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Furukawa, S., Kato, M., Ishiyama, A. et al. Exploring unsolved cases of lissencephaly spectrum: integrating exome and genome sequencing for higher diagnostic yield. J Hum Genet (2024). https://doi.org/10.1038/s10038-024-01283-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s10038-024-01283-0