Abstract
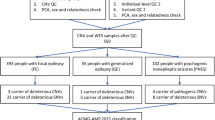
Variants in voltage-gated sodium channel (VGSC) genes are implicated in seizures, epilepsy, and neurodevelopmental disorders, constituting a significant aspect of hereditary epilepsy in the Chinese population. Through retrospective analysis utilizing next-generation sequencing (NGS), we examined the genotypes and phenotypes of VGSC-related epilepsy cases from a cohort of 691 epilepsy subjects. Our findings revealed that 5.1% of subjects harbored VGSC variants, specifically 22 with SCN1A, 9 with SCN2A, 1 with SCN8A, and 3 with SCN1B variants; no SCN3A variants were detected. Among these, 14 variants were previously reported, while 21 were newly identified. SCN1A variant carriers predominantly presented with Dravet Syndrome (DS) and Genetic Epilepsy with Febrile Seizures Plus (GEFS + ), featuring a heightened sensitivity to fever-induced seizures. Statistically significant disparities emerged between the SCN1A-DS and SCN1A-GEFS+ groups concerning seizure onset and genetic diagnosis age, incidence of status epilepticus, mental retardation, anti-seizure medication (ASM) responsiveness, and familial history. Notably, subjects with SCN1A variants affecting the protein’s pore region experienced more frequent cluster seizures. All SCN2A variants were of de novo origin, and 88.9% of individuals with SCN2A variations exhibited cluster seizures. This research reveals a significant association between variations in VGSC-related genes and the clinical phenotype diversity of epilepsy subjects in China, emphasizing the pivotal role of NGS screening in establishing accurate disease diagnoses and guiding the selection of ASM.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive (Genomics, Proteomics & Bioinformatics 2021) in National Genomics Data Center (Nucleic Acids Res 2022), China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences (GSA-Human: HRA004273) that are publicly accessible at https://ngdc.cncb.ac.cn/gsa-human.
References
Ademuwagun IA, Rotimi SO, Syrbe S, Ajamma YU, Adebiyi E. Voltage gated sodium channel genes in epilepsy: mutations, functional studies, and treatment dimensions. Front Neurol. 2021;12:600050.
Kaplan DI, Isom LL, Petrou S. Role of sodium channels in epilepsy. Cold Spring Harb Perspect Med. 2016;6:a022814.
Oliva M, Berkovic SF, Petrou S. Sodium channels and the neurobiology of epilepsy. Epilepsia. 2012;53:1849–59.
Lamar T, Vanoye CG, Calhoun J, Wong JC, Dutton SBB, Jorge BS, et al. SCN3A deficiency associated with increased seizure susceptibility. Neurobiol Dis. 2017;102:38–48.
Al-Zoubi T, Zhou Y, Yin X, Janicek B, Sun C, Schulz CE, et al. Preparation of nonprecious metal electrocatalysts for the reduction of oxygen using a low-temperature sacrificial metal. J Am Chem Soc. 2020;142:5477–81.
Du J, Simmons S, Brunklaus A, Adiconis X, Hession CC, Fu Z, et al. Differential excitatory vs inhibitory SCN expression at single cell level regulates brain sodium channel function in neurodevelopmental disorders. Eur J Paediatr Neurol. 2020;24:129–33.
Dang LT, Quinonez SC, Becka BR, Isom LL, Joshi SM. Dramatic improvement in seizures with phenytoin treatment in an individual with refractory epilepsy and a SCN1B variant. Pediatr Neurol. 2020;108:121–22.
Brunklaus A, Lal D. Sodium channel epilepsies and neurodevelopmental disorders: from disease mechanisms to clinical application. Developmental Med Child Neurol. 2020;62:784–92.
Lal D, May P, Perez-Palma E, Samocha KE, Kosmicki JA, Robinson EB, et al. Gene family information facilitates variant interpretation and identification of disease-associated genes in neurodevelopmental disorders. Genome Med. 2020;12:28.
Fang Z-X, Hong S-Q, Li T-S, Wang J, Xie L-L, Han W, et al. Genetic and phenotypic characteristics of SCN1A-related epilepsy in Chinese children. NeuroReport. 2019;30:671–80.
Cetica V, Chiari S, Mei D, Parrini E, Grisotto L, Marini C, et al. Clinical and genetic factors predicting Dravet syndrome in infants with SCN1A mutations. Neurology. 2017;88:1037–44.
Brunklaus A, Brünger T, Feng T, Fons C, Lehikoinen A, Panagiotakaki E, et al. The gain of function SCN1A disorder spectrum: novel epilepsy phenotypes and therapeutic implications. Brain. 2022;145:3816–31.
Wolff M, Johannesen KM, Hedrich UBS, Masnada S, Rubboli G, Gardella E, et al. Genetic and phenotypic heterogeneity suggest therapeutic implications in SCN2A-related disorders. Brain : a J Neurol. 2017;140:1316–36.
Reynolds C, King MD, Gorman KM. The phenotypic spectrum of SCN2A-related epilepsy. Eur J Paediatr Neurol. 2020;24:117–22.
Zeng Q, Yang Y, Duan J, Niu X, Chen Y, Wang D, et al. SCN2A-related epilepsy: the phenotypic spectrum, treatment and prognosis. Front Mol Neurosci. 2022;15:809951.
Zaman T, Helbig I, Božović IB, DeBrosse SD, Bergqvist AC, Wallis K, et al. Mutations in SCN3A cause early infantile epileptic encephalopathy. Ann Neurol. 2018;83:703–17.
Hu C, Luo T, Wang Y. Phenotypic and genetic spectrum in Chinese children with SCN8A-related disorders. Seizure. 2022;95:38–49.
Gardella E, Moller RS. Phenotypic and genetic spectrum of SCN8A-related disorders, treatment options, and outcomes. Epilepsia. 2019;60:S77–S85.
Gardella E, Marini C, Trivisano M, Fitzgerald MP, Alber M, Howell KB, et al. The phenotype of SCN8A developmental and epileptic encephalopathy. Neurology. 2018;91:e1112–e24.
Ramadan W, Patel N, Anazi S, Kentab AY, Bashiri FA, Hamad MH, et al. Confirming the recessive inheritance of SCN1B mutations in developmental epileptic encephalopathy. Clin Genet. 2017;92:327–31.
Darras N, Ha TK, Rego S, Martin PM, Barroso E, Slavotinek AM, et al. Developmental and epileptic encephalopathy in two siblings with a novel, homozygous missense variant in SCN1B. Am J Med Genet Part A. 2019;179:2190–95.
Steward CA, Roovers J, Suner M-M, Gonzalez JM, Uszczynska-Ratajczak B, Pervouchine D, et al. Re-annotation of 191 developmental and epileptic encephalopathy-associated genes unmasks de novo variants in SCN1A. NPJ Genom Med. 2019;4:31.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Claes LRF, Deprez L, Suls A, Baets J, Smets K, Van Dyck T, et al. TheSCN1Avariant database: a novel research and diagnostic tool. Hum Mutat. 2009;30:E904–E20.
Scheffer IE, Berkovic S, Capovilla G, Connolly MB, French J, Guilhoto L, et al. ILAE classification of the epilepsies: position paper of the ILAE Commission for Classification and Terminology. Epilepsia. 2017;58:512–21.
Ma H, Guo Y, Chen Z, Wang L, Tang Z, Zhang J, et al. Mutations in the sodium channel genes SCN1A, SCN3A, and SCN9A in children with epilepsy with febrile seizures plus(EFS+). Seizure. 2021;88:146–52.
Xie Y, Ng NN, Safrina OS, Ramos CM, Ess KC, Schwartz PH, et al. Comparisons of dual isogenic human iPSC pairs identify functional alterations directly caused by an epilepsy associated SCN1A mutation. Neurobiol Dis. 2020;134:104627.
Brunklaus A, Du J, Steckler F, Ghanty II, Johannesen KM, Fenger CD, et al. Biological concepts in human sodium channel epilepsies and their relevance in clinical practice. Epilepsia. 2020;61:387–99.
Barela AJ, Waddy SP, Lickfett JG, Hunter J, Anido A, Helmers SL, et al. An epilepsy mutation in the sodium ChannelSCN1AThat decreases channel excitability. J Neurosci. 2006;26:2714–23.
Marini C, Scheffer IE, Nabbout R, Suls A, De Jonghe P, Zara F, et al. The genetics of Dravet syndrome. Epilepsia. 2011;52:24–9.
Escayg A, Goldin AL. Sodium channel SCN1A and epilepsy: mutations and mechanisms. Epilepsia. 2010;51:1650–8.
Brunklaus A, Ellis R, Reavey E, Forbes GH, Zuberi SM. Prognostic, clinical and demographic features in SCN1A mutation-positive Dravet syndrome. Brain. 2012;135:2329–36.
Hedrich UBS, Lauxmann S, Lerche H. SCN2A channelopathies: mechanisms and models. Epilepsia. 2019;60:S68–S76.
Liao Y, Deprez L, Maljevic S, Pitsch J, Claes L, Hristova D, et al. Molecular correlates of age-dependent seizures in an inherited neonatal-infantile epilepsy. Brain. 2010;133:1403–14.
Xue Z, Liu K, Liu Q, Li Y, Li M, Su CY, et al. Missing-linker metal-organic frameworks for oxygen evolution reaction. Nat Commun. 2019;10:5048.
Nakamura K, Kato M, Osaka H, Yamashita S, Nakagawa E, Haginoya K, et al. Clinical spectrum of SCN2A mutations expanding to Ohtahara syndrome. Neurology. 2013;81:992–8.
Ben-Shalom R, Keeshen CM, Berrios KN, An JY, Sanders SJ, Bender KJ. Opposing effects on Na(V)1.2 function underlie differences between SCN2A variants observed in individuals with autism spectrum disorder or infantile seizures. Biol Psychiatry. 2017;82:224–32.
Flor-Hirsch H, Heyman E, Livneh A, Reish O, Watemberg N, Litmanovits I, et al. Lacosamide for SCN2A-related intractable neonatal and infantile seizures. Epileptic Disord. 2018;20:440–46.
Dilena R, Striano P, Gennaro E, Bassi L, Olivotto S, Tadini L, et al. Efficacy of sodium channel blockers in SCN2A early infantile epileptic encephalopathy. Brain Dev. 2017;39:345–48.
Bernardo S, Marchionni E, Prudente S, De Liso P, Spalice A, Giancotti A, et al. Unusual association of SCN2A epileptic encephalopathy with severe cortical dysplasia detected by prenatal MRI. Eur J Paediatr Neurol. 2017;21:587–90.
Sanders SJ, Campbell AJ, Cottrell JR, Moller RS, Wagner FF, Auldridge AL, et al. Progress in understanding and treating SCN2A-mediated disorders. Trends Neurosci. 2018;41:442–56.
Johannesen KM, Liu Y, Koko M, Gjerulfsen CE, Sonnenberg L, Schubert J, et al. Genotype-phenotype correlations in SCN8A-related disorders reveal prognostic and therapeutic implications. Brain. 2022;145:2991–3009.
Butler KM, da Silva C, Shafir Y, Weisfeld-Adams JD, Alexander JJ, Hegde M, et al. De novo and inherited SCN8A epilepsy mutations detected by gene panel analysis. Epilepsy Res. 2017;129:17–25.
Epifanio R, Zanotta N, Giorda R, Bardoni A, Zucca C. Novel epilepsy phenotype associated to a known SCN8A mutation. Seizure. 2019;67:15–17.
Ranza E, Z’Graggen W, Lidgren M, Beghetti M, Guipponi M, Antonarakis SE, et al. SCN8A heterozygous variants are associated with anoxic-epileptic seizures. Am J Med Genet A. 2020;182:1209–16.
Myers KA, Scheffer IE, Berkovic SF, Commission IG. Genetic literacy series: genetic epilepsy with febrile seizures plus. Epileptic Disord. 2018;20:232–38.
Wallace RH, Wang DW, Singh R, Scheffer IE, George AL, Phillips HA. Febrile seizures and generalized epilepsy associated with a mutation in the Na+-channel β1 subunit gene SCN1B. Nat Genet. 1998;19:366–70.
Fendri-Kriaa N, Kammoun F, Salem IH, Kifagi C, Mkaouar-Rebai E, Hsairi I, et al. New mutation c.374C>T and a putative disease-associated haplotype within SCN1B gene in Tunisian families with febrile seizures. Eur J Neurol. 2011;18:695–702.
Lindy AS, Stosser MB, Butler E, Downtain-Pickersgill C, Shanmugham A, Retterer K, et al. Diagnostic outcomes for genetic testing of 70 genes in 8565 patients with epilepsy and neurodevelopmental disorders. Epilepsia. 2018;59:1062–71.
Acknowledgements
We thank the subjects and their families for their participation and contribution to the work.
Funding
This work was supported by the National Natural Science Foundation of China (81671362 & 32000075), the financial funding from Department of Science and Technology of Shandong Province (ZR2020QC066 & ZR2020QH102), the financial funding from Jinan Science and Technology Bureau (201907007).
Author information
Authors and Affiliations
Contributions
XL, ZG and YL conceived and designed this study. Experiments were conducted by RD, HZ, KZ, and YL. Data were analyzed by RD, MX, and YL. Clinical diagnosis of the subjects was undertaken by RJ and HZ. The paper was written by RD and YL.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dong, R., Jin, R., Zhang, H. et al. Genotypic and phenotypic characteristics of sodium channel—associated epilepsy in Chinese population. J Hum Genet (2024). https://doi.org/10.1038/s10038-024-01257-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s10038-024-01257-2